Abstract
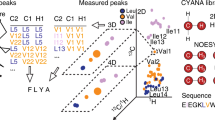
So far high-resolution structure determination by nuclear magnetic resonance (NMR) spectroscopy has been limited to proteins <30 kDa, although global fold determination is possible for substantially larger proteins. Here we present a strategy for assigning backbone and side-chain resonances of large proteins without deuteration, with which one can obtain high-resolution structures from 1H-1H distance restraints. The strategy uses information from through-bond correlation experiments to filter intraresidue and sequential correlations from through-space correlation experiments, and then matches the filtered correlations to obtain sequential assignment. We demonstrate this strategy on three proteins ranging from 24 to 65 kDa for resonance assignment and on maltose binding protein (42 kDa) and hemoglobin (65 kDa) for high-resolution structure determination. The strategy extends the size limit for structure determination by NMR spectroscopy to 42 kDa for monomeric proteins and to 65 kDa for differentially labeled multimeric proteins without the need for deuteration or selective labeling.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Wuthrich, K. NMR of Proteins and Nucleic Acids, (Wiley, New York, 1986).
Kainosho, M. et al. Optimal isotope labelling for NMR protein structure determinations. Nature 440, 52–57 (2006).
LeMaster, D.M. & Richards, F.M. NMR sequential assignment of Escherichia coli thioredoxin utilizing random fractional deuteriation. Biochemistry 27, 142–150 (1988).
Grzesiek, S., Anglister, J., Ren, H. & Bax, A. Carbon-13 line narrowing by deuterium decoupling in deuterium/carbon-13/nitrogen-15 enriched proteins. Application to triple resonance 4D J connectivity of sequential amides. J. Am. Chem. Soc. 115, 4369–4370 (1993).
Venters, R.A., Farmer, B.T., II, Fierke, C.A. & Spicer, L.D. Characterizing the use of perdeuteration in NMR studies of large proteins: 13C, 15N and 1H assignments of human carbonic anhydrase II. J. Mol. Biol. 264, 1101–1116 (1996).
Pervushin, K., Riek, R., Wider, G. & Wuthrich, K. Attenuated T2 relaxation by mutual cancellation of dipole-dipole coupling and chemical shift anisotropy indicates an avenue to NMR structures of very large biological macromolecules in solution. Proc. Natl. Acad. Sci. USA 94, 12366–12371 (1997).
Yang, D.W. & Kay, L.E. TROSY triple-resonance four-dimensional NMR spectroscopy of a 46 ns tumbling protein. J. Am. Chem. Soc. 121, 2571–2575 (1999).
Salzmann, M., Pervushin, K., Wider, G., Senn, H. & Wuthrich, K. NMR assignment and secondary structure determination of an octameric 110 kDa protein using TROSY in triple resonance experiments. J. Am. Chem. Soc. 122, 7543–7548 (2000).
Tugarinov, V., Muhandiram, R., Ayed, A. & Kay, L.E. Four-dimensional NMR spectroscopy of a 723-residue protein: chemical shift assignments and secondary structure of malate synthase G. J. Am. Chem. Soc. 124, 10025–10035 (2002).
Grzesiek, S., Wingfield, P., Stahl, S., Kaufman, J.D. & Bax, A. Four-dimensional 15N-separated NOESY of slowly tumbling perdeuterated 15N-enriched proteins. Application to HIV-1 Nef. J. Am. Chem. Soc. 117, 9594–9595 (1995).
Tjandra, N. & Bax, A. Direct measurement of distances and angles in biomolecules by NMR in a dilute liquid crystalline medium. Science 278, 1111–1114 (1997).
Delaglio, F., Kontaxis, G. & Bax, A. Protein structure determination using molecular fragment replacement and NMR dipolar couplings. J. Am. Chem. Soc. 122, 2142–2143 (2000).
Giesen, A.W., Homans, S.W. & Brown, J.M. Determination of protein global folds using backbone residual dipolar coupling and long-range NOE restraints. J. Biomol. NMR 25, 63–71 (2003).
Rosen, M.K. et al. Selective methyl group protonation of perdeuterated proteins. J. Mol. Biol. 263, 627–636 (1996).
Tugarinov, V., Choy, W.Y., Orekhov, V.Y. & Kay, L.E. Solution NMR-derived global fold of a monomeric 82-kDa enzyme. Proc. Natl. Acad. Sci. USA 102, 622–627 (2005).
Yang, D., Zheng, Y., Liu, D. & Wyss, D.F. Sequence-specific assignments of methyl groups in high-molecular weight proteins. J. Am. Chem. Soc. 126, 3710–3711 (2004).
Zheng, Y., Giovannelli, J.L., Ho, N.T., Ho, C. & Yang, D.W. Side-chain assignments of methyl-containing residues in a uniformly 13C-labeled hemoglobin in the carbonmonoxy form. J. Biomol. NMR 30, 423–429 (2004).
Xu, Y., Lin, Z., Ho, C. & Yang, D.W. A general strategy for the assignment of aliphatic side-chain resonances of uniformly 13C,15N-labeled large proteins. J. Am. Chem. Soc. 127, 11920–11921 (2005).
Ikura, M., Kay, L.E. & Bax, A. A novel approach for sequential assignment of proton, carbon-13, and nitrogen-15 spectra of larger proteins: heteronuclear triple-resonance three-dimensional NMR spectroscopy. Application to calmodulin. Biochemistry 29, 4659–4667 (1990).
Lin, Z., Huang, H., Siu, C-H. & Yang, D.W. 1H, 13C, and 15N resonance assignments of Ca2+-free DdCAD-1: A Ca2+-dependent cell-cell adhesion molecule. J. Biomol. NMR 30, 375–376 (2004).
Gardner, K.H., Zhang, X., Gehring, K. & Kay, L.E. Solution NMR studies of a 42 KDa Escherichia coli maltose binding protein/β-cyclodextrin complex: chemical shift assignments and analysis. J. Am. Chem. Soc. 120, 11738 (1998).
Lukin, J.A. et al. Backbone resonance assignments of human adult hemoglobin in the carbonmonoxy form. J. Biomol. NMR 28, 203–204 (2004).
Lin, Z., Xu, Y.Q., Yang, S. & Yang, D.W. Sequence-specific assignment of aromatic resonances of uniformly 13C,15N-labeled proteins by using 13C- and 15N-edited NOESY spectra. Angew. Chem. Int. Edn. Engl. 45, 1960–1963 (2006).
Skrynnikov, N.R. et al. Orienting domains in proteins using dipolar couplings measured by liquid-state NMR: differences in solution and crystal forms of maltodextrin binding protein loaded with beta-cyclodextrin. J. Mol. Biol. 295, 1265–1273 (2000).
Lukin, J.A. et al. Quaternary structure of hemoglobin in solution. Proc. Natl. Acad. Sci. USA 100, 517–520 (2003).
Wagner, G. & Wüthrich, K. Sequential resonance assignments in protein 1H NMR spectra: Basic pancreatic trypsin inhibitor. J. Mol. Biol. 155, 347–366 (1982).
Tugarinov, V., Kay, L.E., Ibraghimov, I. & Orekhov, V.Y. High-resolution four-dimensional 1H–13C NOE spectroscopy using methyl-TROSY, sparse data acquisition, and multidimensional decomposition. J. Am. Chem. Soc. 127, 2767–2775 (2005).
Coggins, B.E., Venters, R.A. & Zhou, P. Filtered back projection for the reconstruction of a high-resolution (4,2)D CH3-NH NOESY spectrum on a 29 kDa protein. J. Am. Chem. Soc. 127, 11562–11563 (2005).
Delaglio, F. et al. NMRPipe: a multidimensional spectral processing system based on UNIX pipes. J. Biomol. NMR 6, 277–293 (1995).
Acknowledgements
This research was supported by a grant from the Biomedical Research Council (BMRC), and Agency for Science, Technology and Research, A*Star of Singapore. D.Y. is the recipient of a BMRC Young Investigator Award. The authors thank L.E. Kay (University of Toronto) and C. Ho (Carnegie Mellon University) for providing the MBP and HbCO A samples, respectively.
Author information
Authors and Affiliations
Contributions
D.Y. designed the research study; Y.X., Y.Z., J.-S.F. and D.Y. performed the research; Y.X. analyzed the MBP data; Y.Z. assigned DdCAD-1 and HbCO A; J.-S.F. determined the structure of HbCO A; Y.X. and D.Y. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Fig. 1
2D TROSY-HSQC of non-deuterated MBP. (PDF 199 kb)
Supplementary Fig. 2
Distributions of peak signal-to-noise ratio for the 3D TROSY-HNCA experiments. (PDF 453 kb)
Supplementary Fig. 3
Detailed information on the backbone assignments. (PDF 188 kb)
Supplementary Fig. 4
Relative peak intensity as a function of overall correlation time for different types of correlations in a number of 3D and 4D spectra. (PDF 164 kb)
Supplementary Fig. 5
Pulse sequence for recording the 4D 13C,15N-edited NOESY. (PDF 91 kb)
Supplementary Table 1
NMR structural statistics for MBP and HbCO A. (PDF 121 kb)
Supplementary Table 2
Experimental parameters. (PDF 115 kb)
Rights and permissions
About this article
Cite this article
Xu, Y., Zheng, Y., Fan, JS. et al. A new strategy for structure determination of large proteins in solution without deuteration. Nat Methods 3, 931–937 (2006). https://doi.org/10.1038/nmeth938
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth938
This article is cited by
-
Spitzenkörper assembly mechanisms reveal conserved features of fungal and metazoan polarity scaffolds
Nature Communications (2020)
-
Automated NMR resonance assignments and structure determination using a minimal set of 4D spectra
Nature Communications (2018)
-
Silver nanoparticle-human hemoglobin interface: time evolution of the corona formation and interaction phenomenon
Nano Convergence (2017)
-
Structural Basis for pH-mediated Regulation of F-actin Severing by Gelsolin Domain 1
Scientific Reports (2017)
-
Integrating NOE and RDC using sum-of-squares relaxation for protein structure determination
Journal of Biomolecular NMR (2017)