Abstract
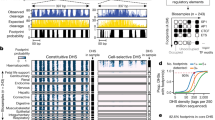
Localized accessibility of critical DNA sequences to the regulatory machinery is a key requirement for regulation of human genes. Here we describe a high-resolution, genome-scale approach for quantifying chromatin accessibility by measuring DNase I sensitivity as a continuous function of genome position using tiling DNA microarrays (DNase-array). We demonstrate this approach across 1% (∼30 Mb) of the human genome, wherein we localized 2,690 classical DNase I hypersensitive sites with high sensitivity and specificity, and also mapped larger-scale patterns of chromatin architecture. DNase I hypersensitive sites exhibit marked aggregation around transcriptional start sites (TSSs), though the majority mark nonpromoter functional elements. We also developed a computational approach for visualizing higher-order features of chromatin structure. This revealed that human chromatin organization is dominated by large (100–500 kb) 'superclusters' of DNase I hypersensitive sites, which encompass both gene-rich and gene-poor regions. DNase-array is a powerful and straightforward approach for systematic exposition of the cis-regulatory architecture of complex genomes.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
References
Felsenfeld, G. Chromatin as an essential part of the transcriptional mechanism. Nature 355, 219–224 (1992).
Felsenfeld, G. & Groudine, M. Controlling the double helix. Nature 421, 448–453 (2003).
Wu, C. The 5′ ends of Drosophila heat shock genes in chromatin are hypersensitive to DNase. Nature 286, 854–860 (1980).
Keene, M.A., Corces, V., Lowenhaupt, K. & Elgin, S.C. DNase I hypersensitive sites in Drosophila chromatin occur at the 5′ ends of regions of transcription. Proc. Natl. Acad. Sci. USA 78, 143–146 (1981).
McGhee, J.D., Wood, W.I., Dolan, M., Engel, J.D. & Felsenfeld, G. A 200 base pair region at the 5′ end of the chicken adult β-globin gene is accessible to nuclease digestion. Cell 27, 45–55 (1981).
Gross, D.S. & Garrard, W.T. Nuclease hypersensitive sites in chromatin. Annu. Rev. Biochem. 57, 159–197 (1988).
Li, Q., Peterson, K.R., Fang, X. & Stamatoyannopoulos, G. Locus control regions. Blood 100, 3077–3086 (2002).
Weintraub, H. & Groudine, M. Chromosomal subunits in active genes have an altered conformation. Science 193, 848–856 (1976).
Felsenfeld, G. et al. Chromatin boundaries and chromatin domains. Cold Spring Harb. Symp. Quant. Biol. 69, 245–250 (2004).
Sproul, D., Gilbert, N. & Bickmore, W.A. The role of chromatin structure in regulating the expression of clustered genes. Nat. Rev. Genet. 6, 775–781 (2005).
The Encode Consortium. The ENCODE (ENCyclopedia Of DNA Elements) Project. Science 306, 636–640 (2004).
Elgin, S.C. The formation and function of DNase I hypersensitive sites in the process of gene activation. J. Biol. Chem. 263, 19259–19262 (1988).
Dorschner, M.O. et al. High-throughput localization of functional elements by quantitative chromatin profiling. Nat. Methods 1, 219–225 (2004).
Lee, G.R., Fields, P.E., Griffin, T.J. & Flavell, R.A. Regulation of the Th2 cytokine locus by a locus control region. Immunity 19, 145–153 (2003).
Lee, G.R., Spilianakis, C.G. & Flavell, R.A. Hypersensitive site 7 of the TH2 locus control region is essential for expressing TH2 cytokine genes and for long-range intrachromosomal interactions. Nat. Immunol. 6, 42–48 (2005).
Loots, G.G. et al. Identification of a coordinate regulator of interleukins 4, 13, and 5 by cross-species sequence comparisons. Science 288, 136–140 (2000).
Ansel, K.M. et al. Deletion of a conserved Il4 silencer impairs T helper type 1-mediated immunity. Nat. Immunol. 5, 1251–1259 (2004).
Smale, S.T. & Kadonaga, J.T. The RNA polymerase II core promoter. Annu. Rev. Biochem. 72, 449–479 (2003).
West, A.G. & Fraser, P. Remote control of gene transcription. Hum. Mol. Genet. 14 (Spec. No. 1), R101–111 (2005).
Yelin, R. et al. Widespread occurrence of antisense transcription in the human genome. Nat. Biotechnol. 21, 379–386 (2003).
Cawley, S. et al. Unbiased mapping of transcription factor binding sites along human chromosomes 21 and 22 points to widespread regulation of noncoding RNAs. Cell 116, 499–509 (2004).
Katayama, S. et al. Antisense transcription in the mammalian transcriptome. Science 309, 1564–1566 (2005).
Hinrichs, A.S. et al. The UCSC Genome Browser Database: update 2006. Nucleic Acids Res. 34, D590–D598 (2006).
Percival, D.B. Wavelet methods for time series analysis (Cambridge University Press, Cambridge, 2000).
Korenberg, J.R. et al. Down syndrome phenotypes: the consequences of chromosomal imbalance. Proc. Natl. Acad. Sci. USA 91, 4997–5001 (1994).
Ren, B. et al. Genome-wide location and function of DNA binding proteins. Science 290, 2306–2309 (2000).
Sabo, P.J. et al. Discovery of functional noncoding elements by digital analysis of chromatin structure. Proc. Natl. Acad. Sci. USA 101, 16837–16842 (2004).
Sabo, P.J. et al. Genome-wide identification of DNaseI hypersensitive sites using active chromatin sequence libraries. Proc. Natl. Acad. Sci. USA 101, 4537–4542 (2004).
McArthur, M., Gerum, S. & Stamatoyannopoulos, G. Quantification of DNaseI-sensitivity by real-time PCR: quantitative analysis of DNaseI-hypersensitivity of the mouse beta-globin LCR. J. Mol. Biol. 313, 27–34 (2001).
Acknowledgements
This work was supported by grants from the US National Institute of General Medical Sciences and the National Human Genome Research Institute to J.A.S. and W.S.N.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
R.D.G. is an employee of NimbleGen Systems, a manufacturer of microarrays, which potentially stands to benefit from the results published in this article.
Supplementary information
Supplementary Fig. 1
Reproducibility of DNase-array. (PDF 31 kb)
Supplementary Fig. 2
Conventional validation of DNase-array. (PDF 3110 kb)
Supplementary Table 1
Shown are location (chromosome, position) of 2,690 DHSs and distance of each to 5′ and 3′ ends of the nearest known gene, mRNA, and spliced EST from the UCSC database. (PDF 226 kb)
Supplementary Table 2
Comparison of DNase-array and conventional Southern validation assays. (PDF 116 kb)
Supplementary Table 3
Primer sequences referred to in Methods and Supplementary Methods. (PDF 53 kb)
Rights and permissions
About this article
Cite this article
Sabo, P., Kuehn, M., Thurman, R. et al. Genome-scale mapping of DNase I sensitivity in vivo using tiling DNA microarrays. Nat Methods 3, 511–518 (2006). https://doi.org/10.1038/nmeth890
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth890
This article is cited by
-
Long-range single-molecule mapping of chromatin accessibility in eukaryotes
Nature Methods (2020)
-
Towards a comprehensive catalogue of validated and target-linked human enhancers
Nature Reviews Genetics (2020)
-
Chromatin accessibility and the regulatory epigenome
Nature Reviews Genetics (2019)
-
Hypomethylated domain-enriched DNA motifs prepattern the accessible nucleosome organization in teleosts
Epigenetics & Chromatin (2017)
-
Genome-wide analysis of p53-regulated transcription in Myc-driven lymphomas
Oncogene (2017)