Abstract
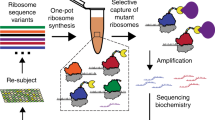
Ribosome display is an in vitro selection and evolution technology for proteins and peptides from large libraries1. As it is performed entirely in vitro, there are two main advantages over other selection technologies2,3. First, the diversity of the library is not limited by the transformation efficiency of bacterial cells, but only by the number of ribosomes and different mRNA molecules present in the test tube. Second, random mutations can be introduced easily after each selection round, as no library must be transformed after any diversification step. This allows facile directed evolution of binding proteins over several generations (Box 1). A prerequisite for the selection of proteins from libraries is the coupling of genotype (RNA, DNA) and phenotype (protein). In ribosome display, this link is accomplished during in vitro translation by stabilizing the complex consisting of the ribosome, the mRNA and the nascent, correctly folded polypeptide (Fig. 1). The DNA library coding for a particular library of binding proteins is genetically fused to a spacer sequence lacking a stop codon. This spacer sequence, when translated, is still attached to the peptidyl tRNA and occupies the ribosomal tunnel, and thus allows the protein of interest to protrude out of the ribosome and fold. The ribosomal complexes are allowed to bind to surface-immobilized target. Whereas non-bound complexes are washed away, mRNA of the complexes displaying a binding polypeptide can be recovered, and thus, the genetic information of the binding polypeptides is available for analysis. Here we describe a step-by-step procedure to perform ribosome display selection using an Escherichia coli S30 extract for in vitro translation, based on the work originally described and further refined in our laboratory1. A protocol that makes use of eukaryotic in vitro translation systems for ribosome display4,6,7 is also included in this issue8.

A DNA library in the form of a PCR product ('library'; see Fig. 2 for construct details), coding for binding proteins, is ligated into the ribosome display vector pRDV, thereby genetically fusing it to a tolA spacer sequence in-frame, and providing a promoter and translation initiation region at the 5′ end. The final ribosome display construct is obtained by PCR amplification of both flanking regions and the library insert from the ligated vector. In vitro transcription of this PCR product yields mRNA that is used for in vitro translation. The ribosome stalls at the end of the mRNA and does not release the encoded and properly folded protein because of the absence of a stop codon. The mRNA-ribosome-protein ternary complexes are used for affinity selection on an immobilized target. mRNA of bound complexes is recovered after washing, reverse transcribed and amplified by PCR. Thereby the selected pools of binders can be used directly for the next cycle of ribosome display or analysis of single clones after cloning into expression vectors.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

Similar content being viewed by others
References
Hanes, J. & Plückthun, A. In vitro selection and evolution of functional proteins by using ribosome display. Proc. Natl. Acad. Sci. USA 94, 4937–4942 (1997).
Levin, A.M. & Weiss, G.A. Optimizing the affinity and specificity of proteins with molecular display. Mol. Biosystems 2, 49–57 (2006).
Hawkins, R.E., Russell, S.J. & Winter, G. Selection of phage antibodies by binding affinity. Mimicking affinity maturation. J. Mol. Biol. 226, 889–896 (1992).
Gersuk, G.M. et al. High-affinity peptide ligands to prostate-specific antigen identified by polysome selection. Biochem. Biophys. Res. Commun. 232, 578–582 (1997).
Hanes, J., Jermutus, L., Schaffitzel, C. & Plückthun, A. Comparison of Escherichia coli and rabbit reticulocyte ribosome display systems. FEBS Lett. 450, 105–110 (1999).
He, M. et al. Selection of a human anti-progesterone antibody fragment from a transgenic mouse library by ARM ribosome display. J. Immunol. Methods 231, 105–117 (1999).
He, M. & Taussig, M.J. Antibody-ribosome-mRNA (ARM) complexes as efficient selection particles for in vitro display and evolution of antibody combining sites. Nucleic Acids Res. 25, 5132–5134 (1997).
He, M. & Taussig, M.J. Eukaryotic ribosome display with in situ DNA recovery. Nat. Methods 4, 281–288 (2006).
Amstutz, P., Binz, H.K., Zahnd, C. & Plückthun, A. Ribosome display: in vitro selection of protein-protein interactions. in Cell Biology, A Laboratory Handbook (ed. Celis, J.) 497–509 (Elsevier Academic Press, 2006).
Pokrovskaya, I.D. & Gurevich, V.V. In vitro transcription: preparative RNA yields in analytical scale reactions. Anal. Biochem. 220, 420–423 (1994).
Hanes, J., Jermutus, L. & Plückthun, A. Selecting and evolving functional proteins in vitro by ribosome display. Methods Enzymol. 328, 404–430 (2000).
Mattheakis, L.C., Bhatt, R.R. & Dower, W.J. An in vitro polysome display system for identifying ligands from very large peptide libraries. Proc. Natl. Acad. Sci. USA 91, 9022–9026 (1994).
Lamla, T. & Erdmann, V.A. Searching sequence space for high-affinity binding peptides using ribosome display. J. Mol. Biol. 329, 381–388 (2003).
Weichhart, T. et al. Functional selection of vaccine candidate peptides from Staphylococcus aureus whole-genome expression libraries in vitro. Infect. Immun. 71, 4633–4641 (2003).
Yau, K.Y., Groves, M.A., Li, S. & Sheedy, C. Selection of hapten-specific single-domain antibodies from a non-immunized llama ribosome display library. J. Immunol. Methods 281, 161–175 (2003).
Hanes, J., Jermutus, L., Weber-Bornhauser, S., Bosshard, H.R. & Plückthun, A. Ribosome display efficiently selects and evolves high-affinity antibodies in vitro from immune libraries. Proc. Natl. Acad. Sci. USA 95, 14130–14135 (1998).
Hanes, J., Schaffitzel, C., Knappik, A. & Plückthun, A. Picomolar affinity antibodies from a fully synthetic naive library selected and evolved by ribosome display. Nat. Biotechnol. 18, 1287–1292 (2000).
Knappik, A. et al. Fully synthetic human combinatorial antibody libraries (HuCAL) based on modular consensus frameworks and CDRs randomized with trinucleotides. J. Mol. Biol. 296, 57–86 (2000).
Lee, M.S. et al. Selection of scFvs specific for HBV DNA polymerase using ribosome display. J. Immunol. Methods 284, 147–157 (2004).
Binz, H.K. et al. High-affinity binders selected from designed ankyrin repeat protein libraries. Nat. Biotechnol. 22, 575–582 (2004).
Schaffitzel, C. et al. In vitro generated antibodies specific for telomeric guanine-quadruplex DNA react with Stylonychia lemnae macronuclei. Proc. Natl. Acad. Sci. USA 98, 8572–8577 (2001).
Zahnd, C. et al. Directed in vitro evolution and crystallographic analysis of a peptide-binding single chain antibody fragment (scFv) with low picomolar affinity. J. Biol. Chem. 279, 18870–18877 (2004).
Jermutus, L., Honegger, A., Schwesinger, F., Hanes, J. & Plückthun, A. Tailoring in vitro evolution for protein affinity or stability. Proc. Natl. Acad. Sci. USA 98, 75–80 (2001).
Matsuura, T. & Plückthun, A. Selection based on the folding properties of proteins with ribosome display. FEBS Lett. 539, 24–28 (2003).
Amstutz, P. et al. In vitro selection for catalytic activity with ribosome display. J. Am. Chem. Soc. 124, 9396–9403 (2002).
Takahashi, F. et al. Ribosome display for selection of active dihydrofolate reductase mutants using immobilized methotrexate on agarose beads. FEBS Lett. 514, 106–110 (2002).
Hanes, J. & Plückthun, A. In vitro selection methods for screening of peptide and protein libraries. in Combinatorial Chemistry in Biology, Vol. 243 (eds. Famulok, M., Winnacker, E.-L. & Wong, C.-H.) 107–122 (Springer Verlag, Berlin Heidelberg, 1999).
Schaffitzel, C., Zahnd, C., Amstutz, P., Luginbühl, B. & Plückthun, A. In vitro selection and evolution of protein-ligand interactions by ribosome display. in Protein-Protein Interactions A Molecular Cloning Manual (eds. Golemis, E. & Adams, P.) 517–548 (Cold Spring Harbor Laboratory Press, Cold Spring Harbor, New York, 2005).
Cadwell, R.C. & Joyce, G.F. Randomization of genes by PCR mutagenesis. PCR Methods Appl. 2, 28–33 (1992).
Stemmer, W.P. Rapid evolution of a protein in vitro by DNA shuffling. Nature 370, 389–391 (1994).
Zaccolo, M., Williams, D.M., Brown, D.M. & Gherardi, E. An approach to random mutagenesis of DNA using mixtures of triphosphate derivatives of nucleoside analogues. J. Mol. Biol. 255, 589–603 (1996).
Schwesinger, F. et al. Unbinding forces of single antibody-antigen complexes correlate with their thermal dissociation rates. Proc. Natl. Acad. Sci. USA 97, 9972–9977 (2000).
Northrup, S.H. & Erickson, H.P. Kinetics of protein-protein association explained by Brownian dynamics computer simulation. Proc. Natl. Acad. Sci. USA 89, 3338–3342 (1992).
Boder, E.T. & Wittrup, K.D. Yeast surface display for screening combinatorial polypeptide libraries. Nat. Biotechnol. 15, 553–557 (1997).
Yang, W.P. et al. CDR walking mutagenesis for the affinity maturation of a potent human anti-HIV-1 antibody into the picomolar range. J. Mol. Biol. 254, 392–403 (1995).
Acknowledgements
We thank R. Skirgaila for originally suggesting and testing Phusion polymerase in ribosome display, as well as A. Batyuk, D. Ferrari, T. Huber, P. Martin Killias, P. Parizek, N. Sainz-Pastor and S.R. Wyss-Stoeckle for experimentally checking the protocol in detail and for many helpful suggestions and discussions, as well as former members of the Plückthun laboratory for establishing the protocol.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
A.P. is an inventor on a patent on ribosome display. Molecular Partners AG is using ribosome display commercially.
Supplementary information
Rights and permissions
About this article
Cite this article
Zahnd, C., Amstutz, P. & Plückthun, A. Ribosome display: selecting and evolving proteins in vitro that specifically bind to a target. Nat Methods 4, 269–279 (2007). https://doi.org/10.1038/nmeth1003
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth1003
This article is cited by
-
The lyophilized chloroplasts store synthetic DARPin G3 as bioactive encapsulated organelles
Journal of Biological Engineering (2023)
-
Assessing immunogenicity barriers of the HIV-1 envelope trimer
npj Vaccines (2023)
-
Trapping the HIV-1 V3 loop in a helical conformation enables broad neutralization
Nature Structural & Molecular Biology (2023)
-
The production of the first functional antibody mimetic in higher plants: the chloroplast makes the DARPin G3 for HER2 imaging in oncology
Biological Research (2022)
-
A cell-free approach to identify binding hotspots in plant immune receptors
Scientific Reports (2022)