Abstract
Understanding genetic events that lead to cancer initiation and progression remains one of the biggest challenges in cancer biology. Traditionally, most algorithms for cancer-driver identification look for genes that have more mutations than expected from the average background mutation rate. However, there is now a wide variety of methods that look for nonrandom distribution of mutations within proteins as a signal for the driving role of mutations in cancer. Here we classify and review such subgene-resolution algorithms, compare their findings on four distinct cancer data sets from The Cancer Genome Atlas and discuss how predictions from these algorithms can be interpreted in the emerging paradigms that challenge the simple dichotomy between driver and passenger genes.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
References
Cancer Genome Atlas Research Network. The Cancer Genome Atlas Pan-Cancer analysis project. Nat. Genet. 45, 1113–1120 (2013).
Gerlinger, M. et al. Intratumor heterogeneity and branched evolution revealed by multiregion sequencing. N. Engl. J. Med. 366, 883–892 (2012).
Watson, I.R., Takahashi, K., Futreal, P.A. & Chin, L. Emerging patterns of somatic mutations in cancer. Nat. Rev. Genet. 14, 703–718 (2013).
Ortmann, C.A. et al. Effect of mutation order on myeloproliferative neoplasms. N. Engl. J. Med. 372, 601–612 (2015).
Lawrence, M.S. et al. Mutational heterogeneity in cancer and the search for new cancer-associated genes. Nature 499, 214–218 (2013).
Niu, B. et al. Protein-structure-guided discovery of functional mutations across 19 cancer types. Nat. Genet. 48, 827–837 (2016).
Leiserson, M.D. et al. Pan-cancer network analysis identifies combinations of rare somatic mutations across pathways and protein complexes. Nat. Genet. 47, 106–114 (2015).
Zhong, Q. et al. Edgetic perturbation models of human inherited disorders. Mol. Syst. Biol. 5, 321 (2009).
Ding, L., Wendl, M.C., McMichael, J.F. & Raphael, B.J. Expanding the computational toolbox for mining cancer genomes. Nat. Rev. Genet. 15, 556–570 (2014).
Gonzalez-Perez, A. et al. Computational approaches to identify functional genetic variants in cancer genomes. Nat. Methods 10, 723–729 (2013).
Leedham, S. & Tomlinson, I. The continuum model of selection in human tumors: general paradigm or niche product? Cancer Res. 72, 3131–3134 (2012).
Nussinov, R. & Tsai, C.J. 'Latent drivers' expand the cancer mutational landscape. Curr. Opin. Struct. Biol. 32, 25–32 (2015).
Castro-Giner, F., Ratcliffe, P. & Tomlinson, I. The mini-driver model of polygenic cancer evolution. Nat. Rev. Cancer 15, 680–685 (2015).
Tamborero, D., Gonzalez-Perez, A. & Lopez-Bigas, N. OncodriveCLUST: exploiting the positional clustering of somatic mutations to identify cancer genes. Bioinformatics 29, 2238–2244 (2013).
Ye, J., Pavlicek, A., Lunney, E.A., Rejto, P.A. & Teng, C.H. Statistical method on nonrandom clustering with application to somatic mutations in cancer. BMC Bioinformatics 11, 11 (2010).
Kamburov, A. et al. Comprehensive assessment of cancer missense mutation clustering in protein structures. Proc. Natl. Acad. Sci. USA 112, E5486–E5495 (2015).
Tokheim, C. et al. Exome-scale discovery of hotspot mutation regions in human cancer using 3D protein structure. Cancer Res. 76, 3719–3731 (2016).
Porta-Pardo, E. & Godzik, A. e-Driver: a novel method to identify protein regions driving cancer. Bioinformatics 30, 3109–3114 (2014).
Melloni, G.E. et al. LowMACA: exploiting protein family analysis for the identification of rare driver mutations in cancer. BMC Bioinformatics 17, 80 (2016).
Reimand, J. & Bader, G.D. Systematic analysis of somatic mutations in phosphorylation signaling predicts novel cancer drivers. Mol. Syst. Biol. 9, 637 (2013).
Porta-Pardo, E., Garcia-Alonso, L., Hrabe, T., Dopazo, J. & Godzik, A. A pan-cancer catalogue of cancer driver protein interaction interfaces. PLoS Comput. Biol. 11, e1004518 (2015).
Mészáros, B., Zeke, A., Reményi, A., Simon, I. & Dosztányi, Z. Systematic analysis of somatic mutations driving cancer: uncovering functional protein regions in disease development. Biol. Direct 11, 23 (2016).
Jia, P. et al. MSEA: detection and quantification of mutation hotspots through mutation set enrichment analysis. Genome Biol. 15, 489 (2014).
Van den Eynden, J., Fierro, A.C., Verbeke, L.P. & Marchal, K. SomInaClust: detection of cancer genes based on somatic mutation patterns of inactivation and clustering. BMC Bioinformatics 16, 125 (2015).
Araya, C.L. et al. Identification of significantly mutated regions across cancer types highlights a rich landscape of functional molecular alterations. Nat. Genet. 48, 117–125 (2016).
Lawrence, M.S. et al. Discovery and saturation analysis of cancer genes across 21 tumour types. Nature 505, 495–501 (2014).
Poole, W., Leinonen, K., Shmulevich, I., Knijnenburg, T.A. & Bernard, B. Multiscale mutation clustering algorithm identifies pan-cancer mutational clusters associated with pathway-level changes in gene expression. PLoS Comput. Biol. 13, e1005347 (2017).
Porta-Pardo, E., Hrabe, T. & Godzik, A. Cancer3D: understanding cancer mutations through protein structures. Nucleic Acids Res. 43, D968–D973 (2015).
Ryslik, G.A., Cheng, Y., Cheung, K.H., Modis, Y. & Zhao, H. Utilizing protein structure to identify nonrandom somatic mutations. BMC Bioinformatics 14, 190 (2013).
Ryslik, G.A., Cheng, Y., Cheung, K.H., Modis, Y. & Zhao, H. A graph theoretic approach to utilizing protein structure to identify nonrandom somatic mutations. BMC Bioinformatics 15, 86 (2014).
Gao, J. et al. 3D clusters of somatic mutations in cancer reveal numerous rare mutations as functional targets. Genome Med. 9, 4 (2017).
Ryslik, G.A. et al. A spatial simulation approach to account for protein structure when identifying nonrandom somatic mutations. BMC Bioinformatics 15, 231 (2014).
Miller, M.L. et al. Pan-cancer analysis of mutation hotspots in protein domains. Cell Syst. 1, 197–209 (2015).
Gonzalez-Perez, A. & Lopez-Bigas, N. Functional impact bias reveals cancer drivers. Nucleic Acids Res. 40, e169 (2012).
Chang, M.T. et al. Identifying recurrent mutations in cancer reveals widespread lineage diversity and mutational specificity. Nat. Biotechnol. 34, 155–163 (2016).
Futreal, P.A. et al. A census of human cancer genes. Nat. Rev. Cancer 4, 177–183 (2004).
Seldin, D.C. et al. CK2 as a positive regulator of Wnt signalling and tumourigenesis. Mol. Cell. Biochem. 274, 63–67 (2005).
Ahmad, K.A., Wang, G., Unger, G., Slaton, J. & Ahmed, K. Protein kinase CK2—a key suppressor of apoptosis. Adv. Enzyme Regul. 48, 179–187 (2008).
Ikeda, Y. et al. Germline PARP4 mutations in patients with primary thyroid and breast cancers. Endocr. Relat. Cancer 23, 171–179 (2016).
Brenan, L. et al. Phenotypic characterization of a comprehensive set of MAPK1/ERK2 missense mutants. Cell Rep. 17, 1171–1183 (2016).
Sim, N.L. et al. SIFT web server: predicting effects of amino acid substitutions on proteins. Nucleic Acids Res. 40, W452–W457 (2012).
Creixell, P. et al. Kinome-wide decoding of network-attacking mutations rewiring cancer signaling. Cell 163, 202–217 (2015).
Mosca, R. et al. dSysMap: exploring the edgetic role of disease mutations. Nat. Methods 12, 167–168 (2015).
Vázquez, M., Valencia, A. & Pons, T. Structure-PPi: a module for the annotation of cancer-related single-nucleotide variants at protein-protein interfaces. Bioinformatics 31, 2397–2399 (2015).
Puente, X.S. et al. Non-coding recurrent mutations in chronic lymphocytic leukaemia. Nature 526, 519–524 (2015).
Brennan, C.W. et al. The somatic genomic landscape of glioblastoma. Cell 155, 462–477 (2013).
Koboldt, D.C. et al. Cancer Genome Atlas Network. Comprehensive molecular portraits of human breast tumours. Nature 490, 61–70 (2012).
Weinstein, J.N. et al. Cancer Genome Atlas Research Network. Comprehensive molecular characterization of urothelial bladder carcinoma. Nature 507, 315–322 (2014).
Cancer Genome Atlas Research Network. Comprehensive molecular profiling of lung adenocarcinoma. Nature 511, 543–550 (2014).
Gonzalez-Perez, A. et al. IntOGen-mutations identifies cancer drivers across tumor types. Nat. Methods 10, 1081–1082 (2013).
Hrabe, T. et al. PDBFlex: exploring flexibility in protein structures. Nucleic Acids Res. 44 D1, D423–D428 (2016).
Tamborero, D. et al. Comprehensive identification of mutational cancer driver genes across 12 tumor types. Sci. Rep. 3, 2650 (2013).
Goldman, M. et al. The UCSC Cancer Genomics Browser: update 2015. Nucleic Acids Res. 43, D812–D817 (2015).
Acknowledgements
We would like to thank the people working at The Cancer Genome Atlas for their efforts and for making all the data publicly available. E.P.-P. and A.G. acknowledge the support from the Cancer Center grants P30 CA030199 (to our institute) and R35 GM118187 (A.G.). A.K. was supported by startup funds of G.G. and by a collaboration with Bayer AG. D.T. is supported by project SAF2015-74072-JIN, which is funded by the Agencia Estatal de Investigacion (AEI) and Fondo Europeo de Desarrollo Regional (FEDER). N.L.-B. acknowledges funding from the European Research Council (consolidator grant 682398). A.V. and T.P. acknowledge funding by the European Union Seventh Framework Programme (FP7/2007-2013) under grant agreement no. 305444 (RD-Connect).
Author information
Authors and Affiliations
Contributions
E.P.-P. and A.G. conceived the project. E.P.-P., D.T. and T.P. researched the data for the article. E.P.-P., A.K. and D.T. analyzed the data. All authors were involved in writing the article and reviewed and edited the manuscript before submission.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Coverage of the human proteome by different types of biological features.
Fraction of the proteome that is covered by linear regions (left), structures with over 95% sequence identity between the protein and the template (middle) or structures with a BLAST e-value between the protein and the template below 1e-9. The fraction is calculated for both, the absolute number of proteins (left columns) as well as the total number of residues (right columns). The distinction between the two is important because it is usually the case that we only know the structure for a fraction of the protein.
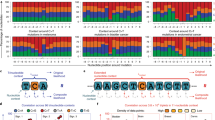
Supplementary Figure 2 Results of the different algorithms in the BLCA dataset.
Visualization is limited to genes detected by at least 4 methods or known drivers in BLCA detected by at least one algorithm.
Supplementary Figure 3 Results of the different algorithms in the BRCA dataset.
Visualization is limited to genes detected by at least 4 methods or known drivers in BRCA detected by at least one algorithm.
Supplementary Figure 4 Results of the different algorithms in the LUAD dataset.
Visualization is limited to genes detected by at least 4 methods or known drivers in LUAD detected by at least one algorithm.
Supplementary Figure 5 Sub-gene resolution algorithms detect more oncogenes than tumor-suppressors.
(a) Barplot showing the fraction of genes detected by each method that are oncogenes, tumor-suppressors, have a dual-role or whose mode of action is not yet known. (b) Fold-enrichment of each method in detected oncogenes or genes with dual-role when aggregating all four datasets.
Supplementary Figure 6 Description of the datasets.
(a) Mutation types in each patient of the different dataset. The majority of mutations are missense. (b) Number of patients (top) and violin plot showing the distribution of number of mutations (bottom) in each dataset. Each dot represents a sample.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6 (PDF 1200 kb)
Supplementary Table 1
Availability and statistical tests used by each method. (XLSX 13 kb)
Supplementary Table 2
Performance of the different methods in the BLCA dataset. (XLSX 10 kb)
Supplementary Table 3
Performance of the different methods on the BRCA dataset. (XLSX 10 kb)
Supplementary Table 4
Performance of the different methods on the GBM dataset. (XLSX 10 kb)
Supplementary Table 5
Performance of the different methods on the LUAD dataset. (XLSX 11 kb)
Supplementary Table 6
Driver genes not detected by whole-gene methods. (XLSX 26 kb)
Supplementary Table 7
Candidate novel driver genes detected only by sub-gene resolution algorithms. (XLSX 17 kb)
Rights and permissions
About this article
Cite this article
Porta-Pardo, E., Kamburov, A., Tamborero, D. et al. Comparison of algorithms for the detection of cancer drivers at subgene resolution. Nat Methods 14, 782–788 (2017). https://doi.org/10.1038/nmeth.4364
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.4364
This article is cited by
-
Comparative assessment of genes driving cancer and somatic evolution in non-cancer tissues: an update of the Network of Cancer Genes (NCG) resource
Genome Biology (2022)
-
Identification of important genes and drug repurposing based on clinical-centered analysis across human cancers
Acta Pharmacologica Sinica (2021)
-
Personalized oncology and BRAFK601N melanoma: model development, drug discovery, and clinical correlation
Journal of Cancer Research and Clinical Oncology (2021)
-
OncoOmics approaches to reveal essential genes in breast cancer: a panoramic view from pathogenesis to precision medicine
Scientific Reports (2020)
-
Interpreting pathways to discover cancer driver genes with Moonlight
Nature Communications (2020)