Abstract
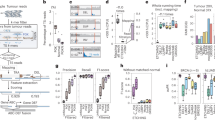
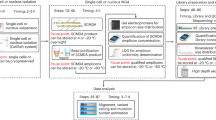
The detection and quantification of low-abundance somatic DNA mutations by high-throughput sequencing is challenging because of the difficulty of distinguishing errors from true mutations. There are several approaches available for analyzing somatic point mutations and small insertions or deletions, but an accurate genome-wide assessment of somatic structural variants (somSVs) in bulk DNA is still not possible. Here we present Structural Variant Search (SVS), a method to accurately detect rare somSVs by low-coverage sequencing. We demonstrate direct quantitative assessment of elevated somSV frequencies induced by known clastogenic compounds in human primary cells.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Tomasetti, C. & Vogelstein, B. Science 347, 78–81 (2015).
Campbell, C.D. & Eichler, E.E. Trends Genet. 29, 575–584 (2013).
Martincorena, I. & Campbell, P.J. Science 349, 1483–1489 (2015).
Gundry, M. & Vijg, J. Mutat. Res. 729, 1–15 (2012).
Maslov, A.Y., Quispe-Tintaya, W., Gorbacheva, T., White, R.R. & Vijg, J. Mutat. Res. 776, 136–143 (2015).
Gundry, M., Li, W., Maqbool, S.B. & Vijg, J. Nucleic Acids Res. 40, 2032–2040 (2012).
Kinde, I., Wu, J., Papadopoulos, N., Kinzler, K.W. & Vogelstein, B. Proc. Natl. Acad. Sci. USA 108, 9530–9535 (2011).
Lou, D.I. et al. Proc. Natl. Acad. Sci. USA 110, 19872–19877 (2013).
Schmitt, M.W. et al. Proc. Natl. Acad. Sci. USA 109, 14508–14513 (2012).
Wang, J. et al. Nat. Methods 8, 652–654 (2011).
Gorbacheva, T., Quispe-Tintaya, W., Popov, V.N., Vijg, J. & Maslov, A.Y. Biotechniques 58, 200–202 (2015).
Zhang, Z.D. et al. BMC Genomics 12, 375 (2011).
Akagi, K. et al. Genome Res. 24, 185–199 (2014).
Viguera, E., Canceill, D. & Ehrlich, S.D. J. Mol. Biol. 312, 323–333 (2001).
Scarpato, R. et al. Mutagenesis 28, 465–473 (2013).
Roy, B. & Hecht, S.M. J. Am. Chem. Soc. 136, 4382–4393 (2014).
Baranello, L. et al. Int. J. Mol. Sci. 15, 13111–13122 (2014).
Campbell, P.J. et al. Nat. Genet. 40, 722–729 (2008).
Rozowsky, J. et al. Nat. Biotechnol. 27, 66–75 (2009).
Acknowledgements
This research was supported by the NIH (grants AG017242, AG047200, and AG038072 to J.V.), the Glenn Foundation for Medical Research (J.V.), the Albert Einstein College of Medicine Human Genome Program (pilot grant to A.Y.M.) the Einstein-Nathan Shock Center of Excellence pilot and feasibility grant to A.Y.M., and the Ministry of Education and Science of the Russian Federation (grant 6.149.2014/K to V.N.P.).
Author information
Authors and Affiliations
Contributions
A.Y.M. and J.V. conceived the idea and wrote the manuscript; W.Q.-T., T.G., and M.L. performed the experiments; A.Y.M., J.V., and V.N.P. analyzed data; A.Y.M. wrote SVS; and S.M. advised on software design.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 PCR validation of novel HPV integration sites in CaSki cells.
(a) HPV integration site-specific primers. (b) Image shows gel-separated PCR products amplified with HPV integration site-specific primers from CaSki and IMR90 genomic DNA. Two out of three identified novel integration sites demonstrated presence of the predicted band in CaSki but not in IMR90 cells.
Supplementary Figure 2 Dose-dependent increase of inter- and intrachromosomal rearrangements in primary human cells after treatment with bleomycin (BLM) or etoposide (ETO).
(a) Frequencies of inter- and intrachromosomal rearrangements in IMR90 cells 72 hours after treatment with clastogens. (b) Frequencies of inter- and intrachromosomal rearrangements in IMR90 cells immediately after treatment with clastogens. All data points represent three biological replicates; data shown as average ± SD; asterisk (*) designates statistically significant difference with corresponding control (*P < 0.05; **P < 0.01; ***P < 0.001).
Supplementary Figure 3 PCR validation of discovered germline SVs specific for IMR90 cells.
Image shows gel-separated PCR products amplified with germline SVs specific primers. 94.4% of interchromosomal (17 out of all 18 detected) and 95.6% of intrachromosomal (66 out of all 69 detected) rearrangements demonstrated the presence of the predicted band in 100-150 bp range. The second higher molecular weight bands in some of the intrachromosomal germline SVs represent the wild-type alleles. First and/or last lane in each gel contains TrackIt 50 bp DNA Ladder (Life Technologies, USA).
Supplementary Figure 4 Microhomology sequences are enriched in the regions of breakpoints induced by bleomycin (BLM) and etoposide (ETO).
(a) Alignment of identified structural variant and human DNA fragments composing breakpoint (vertical line); 7nt-long microhomology highlighted in red. (b) Microhomology (5nt or larger) in germline and somatic (background and induced) SVs.
Supplementary Figure 6 Somatic SVs are enriched in open chromatin regions.
Fraction of SVs’ breakends localized in (a) exons, (b) transcription factor binding sites, and (c) DNAse sensitive sites. (d) Ratio between observed and expected frequencies of structural variants in different genomic regions. Expected frequencies were calculated based on the proportion of the genome occupied by each region considered.
Supplementary Figure 7 Spectra of intrachromosomal somatic SVs induced by different clastogens.
Inversions are defined as intrachromosomal SVs with breakpoints formed by fragments mapped in opposite directions.
Supplementary Figure 8 Frequency of identified germline SVs in each experimental group.
All data points represent three biological replicates; data shown as average ± SD.
Supplementary Figure 9 Possible source of artificial SVs.
(a) Schematic representing the mechanism whereby PCR amplification of transposon-tagged DNA fragments may produce chimeric sequences, as denoted by the presence of the transposon tag in the middle DNA fragment. (b) Structural variants identified by the SVS algorithm do not contain signature of chimeric sequences.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–9, Supplementary Tables 1–5 (PDF 3021 kb)
Supplementary Software
Archived SVS variant caller (ZIP 58 kb)
Source data
Rights and permissions
About this article
Cite this article
Quispe-Tintaya, W., Gorbacheva, T., Lee, M. et al. Quantitative detection of low-abundance somatic structural variants in normal cells by high-throughput sequencing. Nat Methods 13, 584–586 (2016). https://doi.org/10.1038/nmeth.3893
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.3893
This article is cited by
-
Chromosome instability and aneuploidy in the mammalian brain
Chromosome Research (2023)
-
Bleomycin-induced genome structural variations in normal, non-tumor cells
Scientific Reports (2018)