Abstract
Particle tracking is of key importance for quantitative analysis of intracellular dynamic processes from time-lapse microscopy image data. Because manually detecting and following large numbers of individual particles is not feasible, automated computational methods have been developed for these tasks by many groups. Aiming to perform an objective comparison of methods, we gathered the community and organized an open competition in which participating teams applied their own methods independently to a commonly defined data set including diverse scenarios. Performance was assessed using commonly defined measures. Although no single method performed best across all scenarios, the results revealed clear differences between the various approaches, leading to notable practical conclusions for users and developers.
Similar content being viewed by others
Main
Technological developments in the past two decades have greatly advanced the field of bioimaging and have enabled the investigation of dynamic processes in living cells at unprecedented spatial and temporal resolution. Examples include the study of cell membrane dynamics1, cytoskeletal filaments2, focal adhesions3, viral infection4, intracellular transport5, gene transcription6 and genome maintenance7. Apart from state-of-the-art light microscopy8,9 and fluorescent labeling10,11, a key technology in the quest for quantitative analysis of intracellular dynamic processes is particle tracking. Here, a 'particle' may be anything from a single molecule to a macromolecular complex, organelle, virus or microsphere12, and the task of detecting and following individual particles in a time series of images is often (somewhat confusingly) referred to as 'single-particle tracking'. As the number of particles may be very large (hundreds to thousands), requiring 'multiple-particle tracking'13,14,15, manual annotation of the image data is not feasible, and computer algorithms are needed to perform the task.
At present, dozens of software tools are available for particle tracking16. The image analysis methods on which they are based can generally be divided into two steps: (i) particle detection (the spatial aspect), in which spots that stand out from the background according to certain criteria are identified and their coordinates estimated in every frame of the image sequence, and (ii) particle linking (the temporal aspect), in which detected particles are connected from frame to frame using another set of criteria to form tracks. The two steps are commonly performed only once, but they may also be applied iteratively. For each of these steps, many methods have been devised over the years17,18,19,20,21,22, often originating from other areas of data analysis23,24. With so many methods currently known, the question arises as to what distinguishes them and how they perform relative to one another under different experimental conditions.
Several comparison studies have been published in recent years. Cheezum et al. compared four basic methods for localization of a single particle that are often used for particle tracking and concluded that Gaussian fitting performs best by several criteria25. A follow-up study, refining the conclusions by evaluating various practical aspects, was presented by Carter et al.26. A more extensive study, evaluating nine methods (including two machine learning methods) for multiple-particle detection, was conducted by Smal et al.27. They concluded that all methods perform well for sufficiently high signal-to-noise ratio (SNR ≥ 5); however, for low-quality images, learning-based methods are slightly superior, although other methods may yield comparable results and are easier to use. A similar study, published by Ruusuvuori et al.28, rightly added that “algorithms should be chosen with care.” Finally, Godinez et al. compared eight different methods for tracking virus particles and found probabilistic methods to be superior29.
Though interesting, the cited studies were limited to either one aspect of the task (detection rather than tracking) or one application (tracking of viruses rather than a broader set of particles). Moreover, the methods were implemented by the same group who performed the evaluation rather than by the original inventors. Obtaining a more complete picture of performance by combining the results of independent studies is usually hampered by their being based on different data sets and different evaluation criteria. Such fundamental problems have been recognized in the field of medical image analysis for more than 5 years and have resulted in the organization of international competitions (see http://www.grand-challenge.org/). The rationale behind such competitions is that the most objective evaluation of methods is achieved by having research groups apply their own methods independently, on a commonly defined data set and using commonly defined evaluation criteria. The first study in this spirit to be organized in the field of bioimage analysis was the digital reconstruction of axonal and dendritic morphology (DIADEM) challenge30. For particle tracking, the organization of a competition was first advocated by Saxton12 and in an editorial31.
Here we present an objective comparison of particle tracking methods based on an open competition that we organized in 2012 (see http://www.bioimageanalysis.org/track/). By announcements made through various media (at conferences, on the Web and via mailing lists and targeted emails) over 2 months, research groups worldwide were invited to participate. Next, registered teams were given 1 month to prepare their methods using representative training data and corresponding ground truth provided on the website. After release of the actual competition data, without ground truth, the teams were given 3 weeks to submit tracking results to an independent evaluator (one member of the organizing team who was not a contestant and the only one to have the ground truth). Preliminary results were presented and discussed at a workshop organized at the 2012 IEEE International Symposium on Biomedical Imaging. All participating teams sent their software to the independent evaluator who verified the results and performed an objective measurement of the computation times needed by the competing methods. A full analysis of the results and a discussion of the practical conclusions of our study are presented in this paper.
Results
Participating teams and methods
A total of 14 teams (Table 1) took up the challenge and submitted tracking results. Together they used many different methods32,33,34,35,36,37,38,39,40,41,42,43,44,45,46,47,48,49,50,51,52,53,54,55,56,57 (Table 1 and Supplementary Note 1) based on well-known as well as newly developed concepts. Approaches to particle detection ranged from simple thresholding or local-maxima finding to morphological processing, linear filtering (in particular, Gaussian, Laplacian of Gaussian and difference of Gaussian), linear and nonlinear model fitting, and centroid estimation schemes. Most detection methods were based on a combination of two or more of these. Approaches to linking of detected particles ranged from simple nearest-neighbor to multiframe association, including multiple hypothesis tracking, dynamic programming and combinatorial schemes, with or without explicit use of motion models and state estimation (Kalman filtering). Each tracking method consisted of a specific combination of detection and linking approaches as deemed appropriate by the corresponding team, who also determined suitable parameter settings for their method (Supplementary Table 1).
Data sets and ground truth
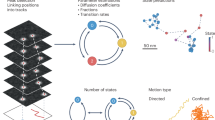
To allow an objective, quantitative comparison of the methods for a range of practical conditions, representative image data with exact ground truth was needed. Generally, ground truth is not available for real image data, and manual annotation by human observers is subjective, labor intensive for large numbers of particles and known14,19,58 to be potentially inferior to computational tracking in the first place, leading to inappropriate reference data. Therefore, we chose to simulate image data for this study (Fig. 1, Table 2 and Supplementary Videos 1,2,3,4,5,6,7,8,9,10). We identified three main factors affecting tracking performance in practice (Supplementary Note 2): dynamics (type of motion), density (number of particles within the fixed field of view) and signal (relative to noise). For particle dynamics we considered four types of motion representative of a variety of biological scenarios, namely Brownian (random-walk) motion similar to that of vesicles in the cytoplasm, directed (near constant–velocity) motion such as microtubule transport and random switching between these two motion models, with either random or constrained orientation for the directed component, as with membrane receptors or infecting viruses, respectively (Fig. 1a, Table 2 and Supplementary Videos 1,2,3,4 and 10). For particle density we considered three levels (Fig. 1b and Supplementary Videos 5, 1 and 6, respectively): low (∼100 particles), medium (∼500 particles) and high (∼1,000 particles), with random appearance and disappearance of particles. For particle signal relative to the noise, we considered four levels (Fig. 1c and Supplementary Videos 7, 8, 2 and 9, respectively): SNR = 1, 2, 4 and 7, where SNR = 4 was known from previous studies25,27 to be a critical level. Here, SNR = (Io − Ib)/√Io, with Io denoting the peak object (particle) intensity and Ib the mean background intensity. Together this resulted in 48 cases. In all cases we modeled particles as labeled with GFP and imaged with fluorescence microscopy in either wide-field or confocal mode. The exact number of particles in any frame of a simulated time series, and the initiation, termination and displacement of particles from frame to frame, was governed by realistic random processes. The resulting data contained ambiguities similar to those in real data, including noise, clutter, visual merging and splitting, and intersecting and parallel trajectories. In both the training and the competition phase of the study, participants were given only limited information about the data (Table 2).
Representative images of the three main factors (particle dynamics, density and signal) affecting tracking performance are shown. (a) Four biological scenarios were simulated, of which we show snapshot images (i–iv) and trajectories (v–viii) in arbitrary colors: particles showing random-walk motion imaged in two dimensions over time (2D+time) using wide-field microscopy (i,v); larger (elongated) particles represented by asymmetric Gaussians showing directed motion in 2D+time (ii,vi); particles switching between random-walk and randomly oriented directed motion imaged in 2D+time using confocal microscopy (iii,vii); and particles switching between random-walk and directed motion with restricted orientation imaged in 3D+time (only one slice is shown) using confocal microscopy (iv,viii). (b,c) Three density levels (b; low, medium and high) and four SNR levels (c; 1, 2, 4 and 7) were simulated.
Quantitative performance measures
A key problem when evaluating any method for tracking large numbers of particles is how to optimally pair the set of estimated tracks, Y, with the set of ground-truth tracks, X, which are likely to contain different numbers of elements (tracks and points within tracks). To solve this, we extended Y with dummy tracks and applied optimal subpattern assignment using the Munkres algorithm59, which yielded the globally best possible pairing (minimal total distance) of each ground-truth track, θkX, with either an estimated track (if available) or a dummy track (in the absence of a suitable estimated track), θkZ, where Z denotes the dummy-extended and ordered version of Y. In the pairing process, the distance d(θkX, θkZ) between two tracks was computed as the sum, over all time points t of the image sequence, of the gated Euclidean distance between the corresponding track points, d(θkX, θkZ) = ∑t|θkX(t) − θkZ(t)|2,ɛ, with θ(t) denoting the spatial position of a track at time t and |.|2,ɛ = min(|.|2,ɛ). If at any t a track point was missing, a dummy point was taken. The gate ɛ served both to determine whether the points of paired tracks were matching at any t and to apply a fixed penalty to nonmatching points. In this study, ɛ was set to 5 pixels, which was on the order of the Rayleigh distance in our data (Supplementary Note 2). The total distance d(X, Y) between track sets X and Y, minimized by the Munkres algorithm by optimizing Z, was simply the sum over all k of the distances d(θkX, θkZ) between paired tracks.
On this basis, we considered 14 different aspects of tracking accuracy, which we summarized into five performance measures. The five measures (Supplementary Note 3) were as follows.
1. α(X, Y) = 1−d(X, Y)/d(X, ∅). ∅ denotes a set of dummy tracks; hence, d(X, ∅) is the maximum possible total distance (error) from the ground truth. The measure ranges from 0 (worst) to 1 (best), indicating the overall degree of matching of ground-truth and estimated tracks without taking into account spurious (nonpaired estimated) tracks.
2. β(X, Y) = (d(X, ∅)−d(X, Y))/(d(X, ∅) + d(Ȳ, ∅)). Ȳ denotes the set of spurious tracks, and d(Ȳ, ∅) is the corresponding penalty term. The measure ranges from 0 (worst) to α (best) and is essentially α with a penalization of nonpaired estimated tracks.
3. JSC = TP/(TP + FN + FP). This is the Jaccard similarity coefficient for track points. It ranges from 0 (worst) to 1 (best) and characterizes overall particle detection performance. TP (true positives) denotes the number of matching points in the optimally paired tracks; FN (false negatives), the number of dummy points in the optimally paired tracks; and FP (false positives), the number of nonmatching points including those of the spurious tracks.
4. JSCθ = TPθ/(TPθ + FNθ + FPθ). This is the Jaccard similarity coefficient for entire tracks instead of single track points. Similarly to JSC, it ranges from 0 (worst) to 1 (best). TPθ denotes the number of estimated tracks paired with ground-truth tracks; FNθ, the number of dummy tracks paired with ground-truth tracks; and FPθ, the number of spurious tracks.
5. RMSE, the r.m.s. error, indicates the overall localization accuracy of matching points in the optimally paired tracks (the TP as in JSC).
Submission of tracking results
Not all teams submitted results for all 48 cases. Some of their methods were not designed to deal with severe noise or more complex shapes or dynamics. Some methods (Table 1) were developed for tracking in only two-dimensional (2D) time series and could not be applied to the 3D cases. And some teams reported insufficient time to complete the tracking of all cases within the 3-week competition phase. Nevertheless, of the 48 (data) × 14 (teams) = 672 possible tracking results, 505 (75%) were submitted to the independent evaluator, who computed the values of all performance measures (Supplementary Table 2), verified the results (Supplementary Table 3) and measured the computation times needed by the methods (Supplementary Table 4).
Performance of the methods
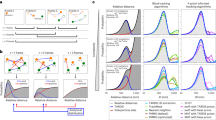
For each tracking method, the values of the performance measures were computed for each data case for which tracking results were submitted. Basing our analysis on the computed values, we studied the performance of the different methods as a function of particle dynamics (the different biological scenarios modeled), density and signal level (Fig. 2 and Supplementary Table 2) as well as in terms of their required computation times (Supplementary Table 4). We subsequently ranked the methods according to best performance per case (Supplementary Tables 2 and 4) as described in the Online Methods. From these rankings we considered the top 3 best-performing methods (Fig. 3) and studied the effects of decreasing the value of the gate parameter ɛ (Supplementary Table 5).
Values of three performance measures (α, β and RMSE) are plotted as a function of density (low, medium and high) and SNR for scenario 1. (a) α values (scoring the match between ground-truth and estimated tracks) for each density. (b) β values (α values with a penalty for nonmatching estimated tracks) for each density. (c) RMSE values (scoring localization accuracy) for each density. For some methods, the lines are incomplete, indicating missing (not submitted) tracking results.
The cells are color coded according to method number (Table 1).
The global observation from the results is that no one particle tracking method performed best for all data. Nevertheless, of the 14 competing methods, some populated the top ranks of the different performance measures considerably more than others (Fig. 3). Counting the number of top 3 occurrences leads to the conclusion that methods 5, 1 and 2 (in this order) were most accurate overall. However, this approach naturally disfavors methods for which only partial results were submitted, and a closer look reveals that some of these methods actually performed better for specific conditions. Examples include method 3, which performed best in terms of α, β, JSC and JSCθ for the higher-SNR data of scenario 3 and was among the top 3 best methods for many cases of scenario 1 (the only two scenarios for which results were submitted for this method); method 4, which performed best in terms of α and β for the higher-SNR data of scenario 2 (the only data for which results were submitted for this method) and in most cases was also the best in terms of RMSE for that data; method 7, which showed the best performance in terms of both JSC measures for the higher-SNR data of scenario 1; method 8, which, particularly in terms of α and β, performed best or second best for the higher-SNR data of scenario 1 but also for some of the lower-SNR data of other scenarios; method 11, which performed best or second best in terms of α, β and JSC for all cases of scenario 3 as well as many cases of scenario 2; method 12, which, in terms of RMSE, was a top 3 method in about half of the cases; and method 13, which was particularly strong for the lowest-SNR data. In terms of computation time, method 1 clearly performed best (fastest), followed by methods 13, 9 and 2 (in this order). Although decreasing ɛ affected the accuracy rankings to some extent, the same methods were found among the top 3 best-performing methods for the given cases (Supplementary Table 5).
Analyzing trends, we observed that within a given scenario, tracking performance depended on particle density and SNR. As expected, in terms of α, β, JSC and JSCθ, the performance of the methods generally decreased with increasing density (Fig. 2a,b and Supplementary Table 2). However, although the number of particles in the scene increased tenfold from lowest to highest density, performance did not drop by the same factor; the methods thus have a certain robustness with respect to increasing particle density. As anticipated, the performance generally did decrease very strongly with decreasing SNR, with the values of most measures dropping to nearly 0 at SNR = 1. Performance dropped especially rapidly below SNR = 4, in line with and confirming earlier findings25,27. In terms of RMSE (Fig. 2c and Supplementary Table 2), the methods showed a similar dependence on SNR (though not as strongly) but virtually no dependence on particle density. This can be explained from the fact that RMSE calculations were limited to matched track points only (Supplementary Note 3). However, localization performance did depend on the scenario. In scenarios 1 and 3, which had relatively simple particle shapes (rotationally symmetric 2D point-spread functions (PSFs)), most methods were able to achieve subpixel localization accuracy for SNR = 4 and SNR = 7, and some even for SNR = 2. By contrast, in scenarios 2 and 4, which had more complex particle shapes (asymmetric Gaussians or 3D PSFs), most methods were considerably less accurate. This can be attributed to the theoretically higher uncertainty in localization of asymmetric objects and to the fact that most methods in this study were not specifically designed for such data and used suboptimal approaches.
The question arises as to what distinguishes the best-performing methods from the other methods in terms of underlying algorithms. Regarding particle detection, all methods used a series of image processing steps, with many commonalities between them (Table 1 and Supplementary Note 1). The general approach to detection is to first preprocess the images to reduce noise and selectively enhance objects (using median, wavelet-based, Gaussian, Laplacian-of-Gaussian or other filters), then to identify prominent spots (often using local-maxima finding or thresholding) and, finally, to estimate the center coordinates of these spots (using Gaussian fitting or intensity-based centroid calculation, or by simply taking the coordinates of the local maxima). The best-performing methods each had slightly different execution without being conceptually very different from some of the lower-performing methods. This suggests that careful numerical implementation and parameter tuning of the algorithms were key factors to success. Some of the methods (1, 8 and 12) made extra efforts in the localization step (iterative centroid calculation or parabolic interpolation), which may explain their superior performance.
As for linking of detected particles, the best methods used multiframe and/or multitrack optimization, going beyond straightforward nearest-neighbor linking (Table 1 and Supplementary Note 1). In particular, Kalman filtering (method 5), multiple hypothesis tracking (methods 2 and 3) and other optimization approaches (methods 1, 4 and 11) were used. If a two-frame approach was used (methods 12 and 13), it was in combination with a gap-closing scheme, essentially combining results from multiple frames to build more consistent tracks. However, similar schemes were used also by many of the lower-performing methods. Rather, the key factor distinguishing the best methods appears to be that they made explicit use of available (or measured) knowledge about the particle motion in each scenario, whereas many of the other methods did so to a lesser extent or even used (implicitly or explicitly) an inappropriate model altogether. It may be argued that this was not fair and that the best methods were perhaps overtrained. However, in biological experiments, where nature does not provide us with a ground-truth training set, it is advisable to use the same approach: assess (theoretically or by initial measurement on the real data) the main parameters of the imaging process and object properties (such as the ones considered in this study), use this prior knowledge to generate synthetic training data (with ground truth) mimicking the real data, use an appropriate image analysis method and fine-tune its parameters on the synthetic data and, finally, apply the fine-tuned method to the real data. This study provides experimentalists with tools to do just that. In addition, the presented results (Fig. 2 and Supplementary Table 2) can be used either to anticipate the success rate of automated particle tracking given the image quality or to determine the image quality required to assure a desired performance level according to the different criteria.
Analysis of biophysical measures
Although we used a comprehensive set of quantitative measures based on concepts also used in other fields, other, more specific measures might be desirable for specific biophysical analyses. Such measures can be easily applied retrospectively, as all results from our study are publicly available. To illustrate this, we performed additional analyses on the tracking results of the methods included in this study. Specifically, for each method and for each case for which results were submitted for that method, we computed the mean-squared displacement (MSD) for a representative range of time intervals (Supplementary Table 6). The resulting MSD curves (Supplementary Figs. 1, 2, 3, 4) represent the estimated dynamic behavior of the particles. Generally, these results confirmed our finding that accuracy increases with increasing SNR and decreasing particle density. Furthermore, we observed that if particle motion is more purely diffusive (as in the vesicle scenarios of our study) rather than directed (as in the microtubule scenarios), most methods are less sensitive to SNR in estimating the MSD and yield good estimates also for SNR as low as 2 or even 1. This is to be expected, as in that case the displacements from one time point to the next are uncorrelated, and track switching errors have much less impact if all particles are subject to the same diffusion process. We also observed that in the case of a directed motion component (all considered scenarios except the vesicle scenarios), there is a general tendency by many methods to underestimate the MSD. This may be explained by the fact that longer particle jumps are more likely to be missed (the tracking methods may be too restrictive) and that track switching errors bias the results toward diffusive motion over longer time scales (if we assume track directions to be random and uncorrelated). We found that, by and large, the top-performing methods (Fig. 3) also performed best in terms of MSD estimation for the indicated cases, reconfirming the suitability of the measures we used for the competition. Similar observations followed from analyzing the results of instantaneous velocity estimation (Supplementary Table 7 and Supplementary Figs. 5, 6, 7, 8). Finally, our retrospective analysis of the distribution of localization errors (Supplementary Table 8 and Supplementary Figs. 9, 10, 11, 12) support and enhance our conclusions above regarding the top-performing methods in terms of RMSE.
Discussion
We acknowledge that our study was one possible comparison of particle tracking methods, and future studies may extend ours in any of its three main aspects: methods, data or measures. Regarding the first, by design our study was limited to those methods developed by teams willing to participate in the competition at the time it was held. Fortunately, many traditional as well as more sophisticated tracking methods were included, and we believe our study was representative of the present state of the art. Regarding data, our study was limited to computer simulations, as this allowed for a controlled analysis (based on absolute ground truth) of tracking performance as a function of different factors. Although we believe that we considered the most important factors (dynamics, density and signal), additional factors could be modeled, such as nonuniform background (more cell like), particle shape and size (varying within images and over time), frame rate (relative to particle velocity) and photobleaching (effectively allowing a time-dependent SNR). However, not only would a full analysis of all these factors be hampered by the 'curse of dimensionality'—that is, an increased difficulty and resource requirement—but we can also expect tracking methods to perform only worse in such (more complex) data; even with most of our data, no method performed anywhere near perfectly. The ultimate challenge remains to obtain real experimental image data with as-accurate-as-possible ground truth. It has been suggested, for example, to use piezo stage–controlled particle motion60.
Notwithstanding inevitable practical constraints, we believe that the present study is a major step toward more objective comparison of particle tracking methods, yielding important results and lessons for future development and experimentation. We identified important factors affecting particle tracking in practice and developed software for computer simulation of challenging image data to analyze tracking performance as a function of these factors. We also identified important measures to quantitatively score estimated tracks with respect to ground-truth reference tracks and developed software to automatically compute them. The software tools are publicly available as part of this article and can be used or further extended by any of those who are interested in benchmarking their particle tracking methods. We mobilized the field and stimulated groups worldwide to compare their methods in an open competition to improve transparency for potential users of the methods. Finally, we used the competition framework to compare current state-of-the-art particle tracking methods, and we performed additional analyses to illustrate the possibility to retrospectively study the impact on specific biophysical parameters beyond those considered in the competition itself.
In closing this article we summarize the main lessons learned for users and developers. Our results indicate that, at present, there exists no universally best method for particle tracking. Users should be aware that a method reported to work for certain experiments may not be the right choice for their application. As we pointed out, it is advisable to use synthetic image data mimicking the real data at hand, both to find the best parameter settings of a given method and to assess its potential performance. To this end, the tools developed as part of our study will prove useful for a wide range of biological scenarios, and the presented results already enable users to anticipate the performance of the tested methods for their applications. Users should be especially cautious when the SNR of their images is considerably lower than 4 (with our definition of SNR), although in the case of more diffusive (rather than directed) particle motion, most methods are able to yield accurate estimations of dynamics even for lower SNR. In selecting a method, users should also bear in mind that methods based on multiframe and/or multitrack optimization schemes in the linking stage, as well as well-tuned motion models, are likely to perform better than methods using simple per-frame and per-particle nearest-neighbor approaches. Thus, although more sophisticated methods may be more difficult to comprehend and control, they may be worth the time investment. For developers, the importance of parameter tuning and making the best possible use of prior knowledge about the data emphasizes the need for domain modeling in computational image analysis and suggests the use of learning-based tracking methods. Because none of the tested methods performed perfectly on any of the data, and real biological data can be even more complex, the quest for better particle tracking methods remains. The results of the present study will serve as a useful baseline for testing the performance of future methods.
Methods
Software implementations.
The software for fully automated generation of the simulated image data used in this study and the software for computation of the performance measures (Supplementary Note 4) were written in the Java programming language as plug-ins for the open bioimage informatics platform Icy61 (Supplementary Software). Software implementations of the particle tracking methods of the participating teams (Supplementary Note 1) were written using various programming languages and platforms, including Java (stand-alone modules or plug-ins for ImageJ/Fiji62 or Icy), C++ (provided as source code or executable) and Matlab (MathWorks).
Analysis of performance results.
For each tracking method and each performance measure, 48 values could in principle be computed, corresponding to the 48 data cases (different combinations of particle dynamics, densities and signal levels). However, not all teams submitted tracking results for all cases, which ruled out the possibility to perform an overall comparison and ranking of the different methods based on all cases. We observed that teams who did not apply their method to all 48 cases generally focused on one or more of the four dynamics scenarios representing different biological applications, but even per scenario not all teams applied their method to all pertaining cases. Therefore, we decided to rank the methods according to best performance per measure and per data case (Fig. 3 and Supplementary Table 2).
Verification of tracking results.
Minor differences between the originally submitted tracking results and the verified results were to be expected because some of the software tools were converted to another platform to allow execution on the single evaluation system, and some methods were probabilistic in nature. Therefore, for each method, differences were considered acceptable (reproducible) if their means for each of α, β, JSC and JSCθ were within 3% and the RMSE was within 0.5 pixel. In the vast majority of cases, the differences were acceptable, and the larger differences in some cases could be traced back to bug fixes and minor improvements in the software or parameter settings used for verification as compared to the original versions. In very few instances the results could not be verified owing to hardware or software limitations (Supplementary Table 3). For the analysis, the performance values computed from the originally submitted tracking results, not the verified results, were used.
Scoring of computation times.
Computation times of all methods were measured on a single workstation (64-bit Intel Xeon X5550 2.67 GHz processor with 24 GB of RAM and running Microsoft Windows 7 Professional or Linux Fedora 16) to allow a fair comparison. We timed only those cases for which tracking results were submitted and verified. Similarly to the analysis of the accuracy performance measures, we ranked the methods according to best timing per data case (Fig. 3 and Supplementary Table 4).
References
Saxton, M.J. & Jacobson, K. Single-particle tracking: applications to membrane dynamics. Annu. Rev. Biophys. Biomol. Struct. 26, 373–399 (1997).
Akhmanova, A. & Steinmetz, M.O. Tracking the ends: a dynamic protein network controls the fate of microtubule tips. Nat. Rev. Mol. Cell Biol. 9, 309–322 (2008).
Berginski, M.E., Vitriol, E.A., Hahn, K.M. & Gomez, S.M. High-resolution quantification of focal adhesion spatiotemporal dynamics in living cells. PLoS ONE 6, e22025 (2011).
Brandenburg, B. & Zhuang, X. Virus trafficking–learning from single-virus tracking. Nat. Rev. Microbiol. 5, 197–208 (2007).
Jandt, U. & Zeng, A.-P. Modeling of intracellular transport and compartmentation. Adv. Biochem. Eng. Biotechnol. 127, 221–249 (2012).
Sinha, B. et al. Dynamic organization of chromatin assembly and transcription factories in living cells. Methods Cell Biol. 98, 57–78 (2010).
Agarwal, S. et al. ATP-dependent and independent functions of Rad54 in genome maintenance. J. Cell Biol. 192, 735–750 (2011).
Stephens, D.J. & Allan, V.J. Light microscopy techniques for live cell imaging. Science 300, 82–86 (2003).
Ji, N., Shroff, H., Zhong, H. & Betzig, E. Advances in the speed and resolution of light microscopy. Curr. Opin. Neurobiol. 18, 605–616 (2008).
Shaner, N.C., Steinbach, P.A. & Tsien, R.Y. A guide to choosing fluorescent proteins. Nat. Methods 2, 905–909 (2005).
Giepmans, B.N.G., Adams, S.R., Ellisman, M.H. & Tsien, R.Y. The fluorescent toolbox for assessing protein location and function. Science 312, 217–224 (2006).
Saxton, M.J. Single-particle tracking: connecting the dots. Nat. Methods 5, 671–672 (2008).
Genovesio, A. et al. Multiple particle tracking in 3-D+t microscopy: method and application to the tracking of endocytosed quantum dots. IEEE Trans. Image Process. 15, 1062–1070 (2006).
Smal, I., Draegestein, K., Galjart, N., Niessen, W. & Meijering, E. Particle filtering for multiple object tracking in dynamic fluorescence microscopy images: application to microtubule growth analysis. IEEE Trans. Med. Imaging 27, 789–804 (2008).
Kalaidzidis, Y. Multiple objects tracking in fluorescence microscopy. J. Math. Biol. 58, 57–80 (2009).
Meijering, E., Dzyubachyk, O. & Smal, I. Methods for cell and particle tracking. Methods Enzymol. 504, 183–200 (2012).
Meijering, E., Smal, I. & Danuser, G. Tracking in molecular bioimaging. IEEE Signal Process. Mag. 23, 46–53 (2006).
Kalaidzidis, Y. Intracellular objects tracking. Eur. J. Cell Biol. 86, 569–578 (2007).
Dorn, J.F., Danuser, G. & Yang, G. Computational processing and analysis of dynamic fluorescence image data. Methods Cell Biol. 85, 497–538 (2008).
Meijering, E., Dzyubachyk, O., Smal, I. & van Cappellen, W.A. Tracking in cell and developmental biology. Semin. Cell Dev. Biol. 20, 894–902 (2009).
Jaqaman, K. & Danuser, G. Computational image analysis of cellular dynamics: a case study based on particle tracking. Cold Spring Harb. Protoc. 2009, pdb.top65 (2009).
Rohr, K. et al. Tracking and quantitative analysis of dynamic movements of cells and particles. Cold Spring Harb. Protoc. 2010, pdb.top80 (2010).
Blackman, S. & Popoli, R. Design and Analysis of Modern Tracking Systems (Artech House, Norwood, Massachusetts, USA, 1999).
Sonka, M., Hlavac, V. & Boyle, R. Image Processing, Analysis, and Machine Vision 3rd edn. (Cengage Learning, Florence, Kentucky, USA, 2007).
Cheezum, M.K., Walker, W.F. & Guilford, W.H. Quantitative comparison of algorithms for tracking single fluorescent particles. Biophys. J. 81, 2378–2388 (2001).
Carter, B.C., Shubeita, G.T. & Gross, S.P. Tracking single particles: a user-friendly quantitative evaluation. Phys. Biol. 2, 60–72 (2005).
Smal, I., Loog, M., Niessen, W. & Meijering, E. Quantitative comparison of spot detection methods in fluorescence microscopy. IEEE Trans. Med. Imaging 29, 282–301 (2010).
Ruusuvuori, P. et al. Evaluation of methods for detection of fluorescence labeled subcellular objects in microscope images. BMC Bioinformatics 11, 248 (2010).
Godinez, W.J. et al. Deterministic and probabilistic approaches for tracking virus particles in time-lapse fluorescence microscopy image sequences. Med. Image Anal. 13, 325–342 (2009).
Gillette, T.A., Brown, K.M., Svoboda, K., Liu, Y. & Ascoli, G.A. DIADEMchallenge.org: a compendium of resources fostering the continuous development of automated neuronal reconstruction. Neuroinformatics 9, 303–304 (2011).
Anonymous. Going for algorithm gold. Nat. Methods 5, 659 (2008).
Sbalzarini, I.F. & Koumoutsakos, P. Feature point tracking and trajectory analysis for video imaging in cell biology. J. Struct. Biol. 151, 182–195 (2005).
Coraluppi, S. & Carthel, C. Recursive track fusion for multi-sensor surveillance. Inf. Fusion 5, 23–33 (2004).
Coraluppi, S. & Carthel, C. Multi-stage multiple-hypothesis tracking. J. Adv. Inf. Fusion 6, 57–67 (2011).
Olivo-Marin, J.-C. Extraction of spots in biological images using multiscale products. Pattern Recognit. 35, 1989–1996 (2002).
Chenouard, N., Bloch, I. & Olivo-Marin, J.-C. Multiple hypothesis tracking in cluttered condition. in Proc. Int. Conf. Image Proc. 3621–3624 (IEEE, 2009).
Chenouard, N., Bloch, I. & Olivo-Marin, J.-C. Multiple hypothesis tracking in microscopy images. in Proc. IEEE Int. Symp. Biomed. Imaging 1346–1349 (IEEE, 2009).
Winter, M.R., Fang, C., Banker, G., Roysam, B. & Cohen, A.R. Axonal transport analysis using Multitemporal Association Tracking. Int. J. Comput. Biol. Drug Des. 5, 35–48 (2012).
Winter, M. et al. Vertebrate neural stem cell segmentation, tracking and lineaging with validation and editing. Nat. Protoc. 6, 1942–1952 (2011).
Godinez, W.J., Lampe, M., Eils, R., Müller, B. & Rohr, K. Tracking multiple particles in fluorescence microscopy images via probabilistic data association. in Proc. IEEE Int. Symp. Biomed. Imaging 1925–1928 (IEEE, 2011).
Rink, J., Ghigo, E., Kalaidzidis, Y. & Zerial, M. Rab conversion as a mechanism of progression from early to late endosomes. Cell 122, 735–749 (2005).
Liang, L., Shen, H., De Camilli, P. & Duncan, J.S. Tracking clathrin coated pits with a multiple hypothesis based method. Med. Image Comput. Comput. Assist. Interv. 6362, 315–322 (2010).
Yin, Z., Kanade, T. & Chen, M. Understanding the phase contrast optics to restore artifact-free microscopy images for segmentation. Med. Image Anal. 16, 1047–1062 (2012).
Magnusson, K.E.G. & Jaldén, J. A batch algorithm using iterative application of the Viterbi algorithm to track cells and construct cell lineages. in Proc. IEEE Int. Symp. Biomed. Imaging 382–385 (IEEE, 2012).
Husain, M., Boudier, T., Paul-Gilloteaux, P., Casuso, I. & Scheuring, S. Software for drift compensation, particle tracking and particle analysis of high-speed atomic force microscopy image series. J. Mol. Recognit. 25, 292–298 (2012).
Casuso, I. et al. Characterization of the motion of membrane proteins using high-speed atomic force microscopy. Nat. Nanotechnol. 7, 525–529 (2012).
Rao, A.R. & Schunck, B.G. Computing oriented texture fields. CVGIP: Graph. Models Image Process. 53, 157–185 (1991).
Hager, G.D., Dewan, M. & Stewart, C.V. Multiple kernel tracking with SSD. in Proc. IEEE Comput. Soc. Conf. Comput. Vis. Pattern Recognit. 790–797 (IEEE, 2004).
Rousseeuw, P.J. & Leroy, A.M. Robust Regression and Outlier Detection (Wiley, Hoboken, New Jersey, USA, 2003).
Thompson, R.E., Larson, D.R. & Webb, W.W. Precise nanometer localization analysis for individual fluorescent probes. Biophys. J. 82, 2775–2783 (2002).
Shafique, K. & Shah, M. A noniterative greedy algorithm for multiframe point correspondence. IEEE Trans. Pattern Anal. Mach. Intell. 27, 51–65 (2005).
Jaqaman, K. et al. Robust single-particle tracking in live-cell time-lapse sequences. Nat. Methods 5, 695–702 (2008).
Lowe, D.G. Distinctive image features from scale-invariant keypoints. Int. J. Comput. Vis. 60, 91–110 (2004).
Crocker, J.C. & Grier, D.G. Methods of digital video microscopy for colloidal studies. J. Colloid Interface Sci. 179, 298–310 (1996).
Celler, K., van Wezel, G.P. & Willemse, J. Single particle tracking of dynamically localizing TatA complexes in Streptomyces coelicolor. Biochem. Biophys. Res. Commun. 438, 38–42 (2013).
Ku, T.-C. et al. An automated tracking system to measure the dynamic properties of vesicles in living cells. Microsc. Res. Tech. 70, 119–134 (2007).
Ku, T.-C., Kao, L.-S., Lin, C.-C. & Tsai, Y.-S. Morphological filter improve the efficiency of automated tracking of secretory vesicles with various dynamic properties. Microsc. Res. Tech. 72, 639–649 (2009).
Huth, J. et al. Significantly improved precision of cell migration analysis in time-lapse video microscopy through use of a fully automated tracking system. BMC Cell Biol. 11, 24 (2010).
Munkres, J. Algorithms for the assignment and transportation problems. J. Soc. Ind. Appl. Math. 5, 32–38 (1957).
Saxton, M.J. Wanted: a positive control for anomalous subdiffusion. Biophys. J. 103, 2411–2422 (2012).
de Chaumont, F. et al. Icy: an open bioimage informatics platform for extended reproducible research. Nat. Methods 9, 690–696 (2012).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Acknowledgements
The authors are grateful to the organizing committee of the 2012 IEEE International Symposium on Biomedical Imaging (ISBI) for hosting the workshop held as part of the study reported in this article. They also acknowledge their collaborators and financial support from their funding agencies. E.M. and I.S.: Dutch Technology Foundation (STW grant 10443); J.-C.O.-M., N.C. and F.d.C.: Agence Nationale de la Recherche (grants ANR-10-INBS-04-06, ANR-10-LABX-62-IBEID, FranceBioImaging) and Programme C'Nano Région IDF (France); N.C.: US National Institutes of Health (NIH grants MH064070 and MH071739); M.M. and C.O.d.S.: Spanish Ministry of Economy and Competitiveness (grant DPI2012-38090-C03-02); M.M.: Czech Ministry of Education (grant 1.07/2.3.00/30.0009); I.F.S. and Y.G.: Swiss National Science Foundation (SNF grant CRSII3-132396/1); I.F.S. and J.C.: Swiss Federal Commission for Technology and Innovation (CTI grant 9325.2-PFLS-LS); A.R.C. and M.W.: US National Institute of Neurological Disorders and Stroke (grant R01NS076709); W.J.G. and K.R.: German Federal Ministry of Education and Research (FORSYS project ViroQuant) and the European Commission (FP7 Project SysPatho); Y.X.: Fundamental Research Funds for the Central Universities, Excellent Young Faculty Award (Zijin Plan) at Zhejiang University, and Ministry of Education (MOE) Key Laboratory of Biomedical Engineering; K.E.G.M. and J.J.: Swedish Research Council (VR grant 621-2011-5884); P.R., C.K. and F.W.: Inria, PICT-IBiSA Institut Curie-CNRS, and French Life Imaging Microscopy Network (GDR 2588-CNRS); J.-Y.T. and S.L.S.: European Commission (FP7 ICT Project MEMI) and N. Perry, R. Henriques and J. Schindelin for their help; K.C., J.W. and G.P.v.W.: Dutch Technology Foundation (STW VICI grant 10379).
Author information
Authors and Affiliations
Contributions
E.M. and J.-C.O.-M. conceived of the study and organized the corresponding workshop at the 2012 IEEE International Symposium on Biomedical Imaging. F.d.C., I.S. and N.C. designed and implemented the software for image generation and comparison of tracking results. F.d.C. generated the image data and ground truth used in the training phase. M.M. generated the image data and ground truth used in the competition phase. M.M. and C.O.d.S. verified the files submitted by the participants and performed the scoring and assessed the computational speed of all methods. M.M., N.C. and E.M. analyzed the results. The methods were designed, implemented, run, and described by the participating teams: method 1: I.F.S., Y.G. and J.C.; method 2: C.C. and S.C.; method 3: N.C., F.d.C. and J.-C.O.-M.; method 4: M.W. and A.R.C.; method 5: W.J.G. and K.R.; method 6: Y.K.; method 7: L.L., J.D., H.S. and Y.X.; method 8: K.E.G.M., J.J. and H.M.B.; method 9: P.P.-G.; method 10: P.R., C.K. and F.W.; method 11: I.S. and E.M.; method 12: J.-Y.T. and S.L.S.; method 13: J.W., K.C. and G.P.v.W.; and method 14: H.-W.D. and Y.-S.T. The article was written by E.M. with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–12, Supplementary Table 5 and Supplementary Notes 1–4 (PDF 25933 kb)
Supplementary Table 1
Parameter values of the particle tracking methods (XLSX 751 kb)
Supplementary Table 2
Performance values of the particle tracking methods (XLSX 491 kb)
Supplementary Table 3
Verification of the performance values (XLSX 366 kb)
Supplementary Table 4
Timings of the particle tracking methods (XLSX 33 kb)
Supplementary Table 6
Mean squared displacement values (XLSX 1077 kb)
Supplementary Table 7
Velocity histogram bin values (XLSX 112 kb)
Supplementary Table 8
Localization histogram bin values (XLSX 99 kb)
Supplementary Video 1
Illustration of the simulated image data for Scenario 1 at medium particle density and a signal-to-noise ratio of 4. (AVI 25606 kb)
Supplementary Video 2
Illustration of the simulated image data for Scenario 2 at medium particle density and a signal-to-noise ratio of 4. (AVI 25606 kb)
Supplementary Video 3
Illustration of the simulated image data for Scenario 3 at medium particle density and a signal-to-noise ratio of 4. (AVI 25606 kb)
Supplementary Video 4
Illustration of the simulated image data for Scenario 4 at medium particle density and a signal-to-noise ratio of 4. Only slice number 5 of each 3D stack in the time series is shown. See Supplementary Video 10 for a volume rendering of the same data. (AVI 25606 kb)
Supplementary Video 5
Illustration of the simulated image data for Scenario 1 at low particle density and a signal-to-noise ratio of 4. (AVI 25606 kb)
Supplementary Video 6
Illustration of the simulated image data for Scenario 1 at high particle density and a signal-to-noise ratio of 4. (AVI 25606 kb)
Supplementary Video 7
Illustration of the simulated image data for Scenario 2 at medium particle density and a signal-to-noise ratio of 1. (AVI 25606 kb)
Supplementary Video 8
Illustration of the simulated image data for Scenario 2 at medium particle density and a signal-to-noise ratio of 2. (AVI 25606 kb)
Supplementary Video 9
Illustration of the simulated image data for Scenario 2 at medium particle density and a signal-to-noise ratio of 7. (AVI 25606 kb)
Supplementary Video 10
Illustration of the simulated image data for Scenario 4 at medium particle density and a signal-to-noise ratio of 4. A volume rendering of the time series is shown. See Supplementary Video 4 for a single slice of each 3D stack in the same data. (AVI 30981 kb)
Supplementary Software
Simulation and evaluation software (ZIP 161123 kb)
Source data
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/.
About this article
Cite this article
Chenouard, N., Smal, I., de Chaumont, F. et al. Objective comparison of particle tracking methods. Nat Methods 11, 281–289 (2014). https://doi.org/10.1038/nmeth.2808
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.2808
This article is cited by
-
Temporal analysis of relative distances (TARDIS) is a robust, parameter-free alternative to single-particle tracking
Nature Methods (2024)
-
Metrics reloaded: recommendations for image analysis validation
Nature Methods (2024)
-
Machine-learning-powered extraction of molecular diffusivity from single-molecule images for super-resolution mapping
Communications Biology (2023)
-
Force transmission by retrograde actin flow-induced dynamic molecular stretching of Talin
Nature Communications (2023)
-
Quantifying postsynaptic receptor dynamics: insights into synaptic function
Nature Reviews Neuroscience (2023)