Abstract
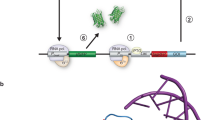
Maintenance of cellular protein homeostasis (proteostasis) depends on a complex network of molecular chaperones, proteases and other regulatory factors. Proteostasis deficiency develops during normal aging and predisposes individuals for many diseases, including neurodegenerative disorders. Here we describe sensor proteins for the comparative measurement of proteostasis capacity in different cell types and model organisms. These sensors are increasingly structurally destabilized versions of firefly luciferase. Imbalances in proteostasis manifest as changes in sensor solubility and luminescence activity. We used EGFP-tagged constructs to monitor the aggregation state of the sensors and the ability of cells to solubilize or degrade the aggregated proteins. A set of three sensor proteins serves as a convenient toolkit to assess the proteostasis status in a wide range of experimental systems, including cell and organism models of stress, neurodegenerative disease and aging.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Powers, E.T., Morimoto, R.I., Dillin, A., Kelly, J.W. & Balch, W.E. Biological and chemical approaches to diseases of proteostasis deficiency. Annu. Rev. Biochem. 78, 959–991 (2009).
Vabulas, R.M., Raychaudhuri, S., Hayer-Hartl, M. & Hartl, F.U. Protein folding in the cytoplasm and the heat shock response. Cold Spring Harb. Perspect. Biol. 2, a004390 (2010).
Frydman, J. Folding of newly translated proteins in vivo: the role of molecular chaperones. Annu. Rev. Biochem. 70, 603–647 (2001).
Rubinsztein, D.C. The roles of intracellular protein-degradation pathways in neurodegeneration. Nature 443, 780–786 (2006).
Ben-Zvi, A., Miller, E.A. & Morimoto, R.I. Collapse of proteostasis represents an early molecular event in Caenorhabditis elegans aging. Proc. Natl. Acad. Sci. USA 106, 14914–14919 (2009).
Balch, W.E., Morimoto, R.I., Dillin, A. & Kelly, J.W. Adapting proteostasis for disease intervention. Science 319, 916–919 (2008).
Mu, T.W. et al. Chemical and biological approaches synergize to ameliorate protein-folding diseases. Cell 134, 769–781 (2008).
Gidalevitz, T., Ben-Zvi, A., Ho, K.H., Brignull, H.R. & Morimoto, R.I. Progressive disruption of cellular protein folding in models of polyglutamine diseases. Science 311, 1471–1474 (2006).
Frydman, J., Nimmesgern, E., Ohtsuka, K. & Hartl, F.U. Folding of nascent polypeptide chains in a high molecular mass assembly with molecular chaperones. Nature 370, 111–117 (1994).
Thulasiraman, V. & Matts, R.L. Effect of geldanamycin on the kinetics of chaperone-mediated renaturation of firefly luciferase in rabbit reticulocyte lysate. Biochemistry 35, 13443–13450 (1996).
Nimmesgern, E. & Hartl, F.U. ATP-dependent protein refolding activity in reticulocyte lysate. Evidence for the participation of different chaperone components. FEBS Lett. 331, 25–30 (1993).
Schroder, H., Langer, T., Hartl, F.U. & Bukau, B. DnaK, DnaJ and GrpE form a cellular chaperone machinery capable of repairing heat-induced protein damage. EMBO J. 12, 4137–4144 (1993).
Conti, E., Franks, N.P. & Brick, P. Crystal structure of firefly luciferase throws light on a superfamily of adenylate-forming enzymes. Structure 4, 287–298 (1996).
Naylor, L.H. Reporter gene technology: the future looks bright. Biochem. Pharmacol. 58, 749–757 (1999).
Hageman, J., Vos, M.J., van Waarde, M.A. & Kampinga, H.H. Comparison of intra-organellar chaperone capacity for dealing with stress-induced protein unfolding. J. Biol. Chem. 282, 34334–34345 (2007).
Michels, A.A., Nguyen, V.T., Konings, A.W., Kampinga, H.H. & Bensaude, O. Thermostability of a nuclear-targeted luciferase expressed in mammalian cells. Destabilizing influence of the intranuclear microenvironment. Eur. J. Biochem. 234, 382–389 (1995).
Matsui, I. & Harata, K. Implication for buried polar contacts and ion pairs in hyperthermostable enzymes. FEBS J. 274, 4012–4022 (2007).
Schneider, C. et al. Pharmacologic shifting of a balance between protein refolding and degradation mediated by Hsp90. Proc. Natl. Acad. Sci. USA 93, 14536–14541 (1996).
Sharp, S. & Workman, P. Inhibitors of the HSP90 molecular chaperone: current status. Adv. Cancer Res. 95, 323–348 (2006).
Taipale, M., Jarosz, D.F. & Lindquist, S. HSP90 at the hub of protein homeostasis: emerging mechanistic insights. Nat. Rev. Mol. Cell Biol. 11, 515–528 (2010).
Muchowski, P.J. Protein misfolding, amyloid formation, and neurodegeneration: a critical role for molecular chaperones? Neuron 35, 9–12 (2002).
Broadley, S.A. & Hartl, F.U. The role of molecular chaperones in human misfolding diseases. FEBS Lett. 583, 2647–2653 (2009).
Morimoto, R.I. & Cuervo, A.M. Protein homeostasis and aging: taking care of proteins from the cradle to the grave. J. Gerontol. A Biol. Sci. Med. Sci. 64, 167–170 (2009).
Kern, A., Ackermann, B., Clement, A.M., Duerk, H. & Behl, C. HSF1-controlled and age-associated chaperone capacity in neurons and muscle cells of C. elegans. PLoS ONE 5, e8568 (2010).
Kaganovich, D., Kopito, R. & Frydman, J. Misfolded proteins partition between two distinct quality control compartments. Nature 454, 1088–1095 (2008).
Nakatsu, T. et al. Structural basis for the spectral difference in luciferase bioluminescence. Nature 440, 372–376 (2006).
Behrends, C. et al. Chaperonin TRiC promotes the assembly of polyQ expansion proteins into nontoxic oligomers. Mol. Cell 23, 887–897 (2006).
Acknowledgements
We thank M. Hipp, Y.E. Kim and M. Hayer-Hartl for discussion, F. Buchholz (Max Planck Institute for Molecular Cell Biology and Genetics) for providing Hsc70 endoribonuclease-prepared small interfering (esi)RNA, and H. Wagner (Institut für Med. Mikrobiologie, Immunologie und Hygiene, Technische Universität München) for the HSP70-Luc plasmid. We acknowledge the expert technical assistance of V. Marcus. This work has been supported by the European Commission within the 7th framework program Proteomics Specification in Time and Space.
Author information
Authors and Affiliations
Contributions
S.R. and F.U.H. conceived the idea and developed the method. A.B. designed Fluc mutants. R.G., C.L. and S.R. performed molecular biology and cell biology experiments. P.K. and M.Z. performed C. elegans experiments. A.V. and D.G. performed D. melanogaster S2 cell experiments. R.G. and S.R. analyzed results. S.R. and F.U.H. interpreted results and wrote the manuscript with assistance from D.G.
Corresponding authors
Ethics declarations
Competing interests
F.U.H is a paid consultant of Proteostasis Therapeutics Inc. A.V and D.G are employees of Proteostasis Therapeutics Inc.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8, Supplementary Tables 1–3, Supplementary Note 1 (PDF 5904 kb)
Rights and permissions
About this article
Cite this article
Gupta, R., Kasturi, P., Bracher, A. et al. Firefly luciferase mutants as sensors of proteome stress. Nat Methods 8, 879–884 (2011). https://doi.org/10.1038/nmeth.1697
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.1697
This article is cited by
-
The AAA+ chaperone VCP disaggregates Tau fibrils and generates aggregate seeds in a cellular system
Nature Communications (2023)
-
A fluorescent multi-domain protein reveals the unfolding mechanism of Hsp70
Nature Chemical Biology (2023)
-
Nuclear and cytoplasmic spatial protein quality control is coordinated by nuclear–vacuolar junctions and perinuclear ESCRT
Nature Cell Biology (2023)
-
LONRF2 is a protein quality control ubiquitin ligase whose deficiency causes late-onset neurological deficits
Nature Aging (2023)
-
NEMO reshapes the α-Synuclein aggregate interface and acts as an autophagy adapter by co-condensation with p62
Nature Communications (2023)