Abstract
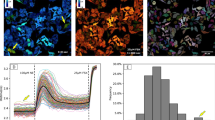
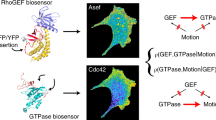
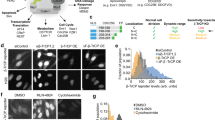
Extracellular stimuli are transduced inside the cell by posttranslational modifications (PTMs), such as phosphorylation, of proteins in signaling networks. Insight into the structure of these networks requires quantification of PTM levels in individual cells. Fluorescence resonance energy transfer (FRET) measured by fluorescence lifetime imaging microscopy (FLIM) is a powerful tool to image PTM levels in situ. FLIM on cell arrays that express fluorescent protein fusions can quantify tyrosine phosphorylation patterns in large networks in individual cells. We identified tyrosine kinase substrates by imaging their phosphorylation levels after inhibition of protein tyrosine phosphatases. Analysis of the correlation between protein phosphorylation and expression levels at single cell resolution allowed us to identify positive feedback motifs. Using FLIM on cell arrays (CA-FLIM), we uncovered components that transduce signals from epidermal growth factor receptor.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Santos, S.D., Verveer, P.J. & Bastiaens, P.I. Growth factor-induced MAPK network topology shapes Erk response determining PC-12 cell fate. Nat. Cell Biol. 9, 324–330 (2007).
Zamir, E. & Bastiaens, P.I. Reverse engineering intracellular biochemical networks. Nat. Chem. Biol. 4, 643–647 (2008).
Stiffler, M.A. et al. PDZ domain binding selectivity is optimized across the mouse proteome. Science 317, 364–369 (2007).
Olsen, J.V. et al. Global, in vivo, and site-specific phosphorylation dynamics in signaling networks. Cell 127, 635–648 (2006).
Neumann, B. et al. High-throughput RNAi screening by time-lapse imaging of live human cells. Nat. Methods 3, 385–390 (2006).
Maslov, S. & Sneppen, K. Specificity and stability in topology of protein networks. Science 296, 910–913 (2002).
Wagner, A. Robustness against mutations in genetic networks of yeast. Nat. Genet. 24, 355–361 (2000).
Gadella, T.W.J., Jovin, T.M. & Clegg, R.M. Fluorescence lifetime imaging microscopy (FLIM)—spatial-resolution of microstructures on the nanosecond time-scale. Biophys. Chem. 48, 221–239 (1993).
Bastiaens, P.I. & Squire, A. Fluorescence lifetime imaging microscopy: spatial resolution of biochemical processes in the cell. Trends Cell Biol. 9, 48–52 (1999).
Verveer, P.J., Wouters, F.S., Reynolds, A.R. & Bastiaens, P.I.H. Quantitative imaging of lateral ErbB1 receptor signal propagation in the plasma membrane. Science 290, 1567–1570 (2000).
Reynolds, A.R., Tischer, C., Verveer, P.J., Rocks, O. & Bastiaens, P.I. EGFR activation coupled to inhibition of tyrosine phosphatases causes lateral signal propagation. Nat. Cell Biol. 5, 447–453 (2003).
Maeder, C.I. et al. Spatial regulation of Fus3 MAP kinase activity through a reaction-diffusion mechanism in yeast pheromone signalling. Nat. Cell Biol. 9, 1319–1326 (2007).
Ziauddin, J. & Sabatini, D.M. Microarrays of cells expressing defined cDNAs. Nature 411, 107–110 (2001).
Simpson, J.C., Wellenreuther, R., Poustka, A., Pepperkok, R. & Wiemann, S. Systematic subcellular localization of novel proteins identified by large-scale cDNA sequencing. EMBO Rep. 1, 287–292 (2000).
Verveer, P.J., Squire, A. & Bastiaens, P.I.H. Global analysis of fluorescence lifetime imaging microscopy data. Biophys. J. 78, 2127–2137 (2000).
Grecco, H.E., Roda-Navarro, P. & Verveer, P.J. Global analysis of time correlated single photon counting FRET-FLIM data. Opt. Express 17, 6493–6508 (2009).
Clayton, A.H., Hanley, Q.S. & Verveer, P.J. Graphical representation and multicomponent analysis of single-frequency fluorescence lifetime imaging microscopy data. J. Microsc. 213, 1–5 (2004).
Davis, T.L. et al. Autoregulation by the juxtamembrane region of the human ephrin receptor tyrosine kinase A3 (EphA3). Structure 16, 873–884 (2008).
Mikalsen, S.O. & Kaalhus, O. Properties of pervanadate and permolybdate. Connexin43, phosphatase inhibition, and thiol reactivity as model systems. J. Biol. Chem. 273, 10036–10045 (1998).
Huyer, G. et al. Mechanism of inhibition of protein-tyrosine phosphatases by vanadate and pervanadate. J. Biol. Chem. 272, 843–851 (1997).
Blom, N., Gammeltoft, S. & Brunak, S. Sequence and structure-based prediction of eukaryotic protein phosphorylation sites. J. Mol. Biol. 294, 1351–1362 (1999).
Ting, A.Y., Kain, K.H., Klemke, R.L. & Tsien, R.Y. Genetically encoded fluorescent reporters of protein tyrosine kinase activities in living cells. Proc. Natl. Acad. Sci. USA 98, 15003–15008 (2001).
Szymkiewicz, I. et al. CIN85 participates in Cbl-b–mediated down-regulation of receptor tyrosine kinases. J. Biol. Chem. 277, 39666–39672 (2002).
Beaujolais, S.A. et al. Large-scale characterization of HeLa cell nuclear phosphoproteins. Proc. Natl. Acad. Sci. USA 101, 12130–12135 (2004).
Glister, M.L., Yang, Y., Uren, J. & Schlessinger, J. Activation of the nonreceptor protein tyrosine kinase Ack by multiple extracellular stimuli. Proc. Natl. Acad. Sci. USA 103, 9796–9801 (2006).
Li, W. et al. Srcasm modulates EGF and Src-kinase signaling in keratinocytes. J. Biol. Chem. 280, 6036–6046 (2005).
Shibamoto, S. et al. Tyrosine phosphorylation of beta-catenin and plakoglobin enhanced by hepatocyte growth factor and epidermal growth factor in human carcinoma cells. Cell Adhes. Commun. 1, 295–305 (1994).
Pennock, S. & Wang, Z. A tale of two Cbls: interplay of c-Cbl and Cbl-b in epidermal growth factor receptor downregulation. Mol. Cell. Biol. 28, 3020–3037 (2008).
Tyson, J.J., Chen, K.C. & Novak, B. Sniffers, buzzers, toggles and blinkers: dynamics of regulatory and signaling pathways in the cell. Curr. Opin. Cell Biol. 15, 221–231 (2003).
Sachs, K., Perez, O., Pe'er, D., Lauffenburger, D.A. & Nolan, G.P. Causal protein-signaling networks derived from multiparameter single-cell data. Science 308, 523–529 (2005).
Erfle, H. et al. Reverse transfection on cell arrays for high content screening microscopy. Nat. Protocols 2, 392–399 (2007).
van Munster, E.B. & Gadella, T.W.J. Suppression of photobleaching-induced artifacts in frequency-domain FLIM by permutation of the recording order. Cytometry A 58A, 185–194 (2004).
Griesbeck, O., Baird, G.S., Campbell, R.E., Zacharias, D.A. & Tsien, R.Y. Reducing the environmental sensitivity of yellow fluorescent protein. Mechanism and applications. J. Biol. Chem. 276, 29188–29194 (2001).
Lew, E.D., Furdui, C.M., Anderson, K.S. & Schlessinger, J. The precise sequence of FGF receptor autophosphorylation is kinetically driven and is disrupted by oncogenic mutations. Sci. Signal. 2, ra6 (2009).
Acknowledgements
P.R.-N. was supported by a Marie Curie Intra-European fellowship for career development (FP7-PEOPLE 2007-2-1-EIF). This work was supported by Interaction Proteome (Integrated Project from FP6) and the Centre for Systems Biology in Dortmund (cofinanced by the European Regional Development Fund and the State of North Rhine-Westfalia). S. Dhe-Paganon (University of Toronto) provided the pET28a-LIC-EPHA3c plasmid.
Author information
Authors and Affiliations
Contributions
P.I.H.B. devised the method. A.S. and R.P. contributed the initial implementation. H.E.G., P.R.-N. and A.G. designed, supervised, performed and analyzed the experiments. A.S., T.F., D.C.T. and J.H. contributed experiments. A.S. and H.E.G. built the instrument. H.E.G., P.R-N. and P.I.H.B. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–10, Supplementary Tables 1–2, Supplementary Protocols and Supplementary Discussion (PDF 3761 kb)
Rights and permissions
About this article
Cite this article
Grecco, H., Roda-Navarro, P., Girod, A. et al. In situ analysis of tyrosine phosphorylation networks by FLIM on cell arrays. Nat Methods 7, 467–472 (2010). https://doi.org/10.1038/nmeth.1458
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.1458
This article is cited by
-
A self-organized synthetic morphogenic liposome responds with shape changes to local light cues
Nature Communications (2021)
-
A conformational sensor based on genetic code expansion reveals an autocatalytic component in EGFR activation
Nature Communications (2018)
-
Spatial cycles mediated by UNC119 solubilisation maintain Src family kinases plasma membrane localisation
Nature Communications (2017)
-
Developments in preclinical cancer imaging: innovating the discovery of therapeutics
Nature Reviews Cancer (2014)
-
Small molecule inhibition of the KRAS–PDEδ interaction impairs oncogenic KRAS signalling
Nature (2013)