Abstract
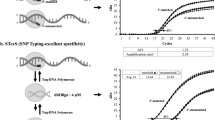
Genetic diagnoses, such as single nucleotide polymorphism (SNP) typing, allow elucidation of gene-based physiological differences, such as susceptibility to diseases and response to drugs, among individuals. Many detection technologies, including allele-specific hybridization, allele-specific primer extension and oligonucleotide ligation, are being used to discriminate SNP alleles1,2. These methods still have many unsolved practical issues1,2,3. In general they require adequate and specific hybridizations of primer or probe DNAs with target DNAs. This frequently needs optimization of the probe/primer structures and operating conditions. In nature, highly homology-sensitive hybridization is assisted by a nucleic acid chaperone that reduces the energy barrier associated with breakage and reassociation of nucleic base pairs4,5. Here we report a simple, quick, precise but enzyme-free method for SNP analysis. The method uses cationic comb-type copolymers (CCCs) producing high nucleic acid chaperone activities. A single-base mismatch in 20-mer DNA can be detected within a few minutes at ambient temperatures (25–37 °C). Even without careful optimization processes, the method has the sensitivity to detect the mismatches causing subtle changes (ΔTm ≈ 1 °C) in duplex thermal stability. CCCs may have various bioanalytical applications where precise hybridization of nucleic acids is needed.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Brennan, M.D. High throughput genotyping technologies for pharmacogenomics. Am. J. Pharmacogenomics 1, 295–302 (2001).
Kwok, P.Y. Methods for genotyping single nucleotide polymorphisms. Annu. Rev. Genom. Hum. Genet. 2, 235–258 (2001).
Chicurel, M. Faster, better, cheaper genotyping. Nature 412, 580–582 (2001).
Tsuchihashi, Z. & Brown, P.O. DNA strand exchange and selective DNA annealing promoted by the human immunodeficiency virus type 1 nucleocapsid protein. J. Virol. 68, 5863–5870 (1994).
Rein, A., Henderson, L.E. & Levin, J.G. Nucleic-acid-chaperone activity of retroviral nucleocapsid proteins: significance for viral replication. Trends Biochem. Sci. 23, 297–301 (1998).
Reynaldo, L.P., Vologodskii, A.V., Neri, B.P. & Lyamichev, V.I. The kinetic of oligonucleotide replacements. J. Mol. Biol. 297, 511–520 (2000).
Panyutin, I.G. & Hsieh, P. Formation of a single base mismatch impedes spontaneous DNA branch migration. J. Mol. Biol. 230, 413–424 (1993).
Li, Q., Luan, G., Guo, Q. & Liang, J. A new class of homogeneous nucleic acid probes based on specific displacement hybridisation. Nucleic Acids Res. 30, e5 (2002).
Kim, W.J., Ishihara, T., Akaike, T. & Maruyama, A. Comb-type cationic copolymer expedites DNA strand exchange while stabilizing DNA duplex. Chem. Eur. J. 7, 176–180 (2001).
Kim, W.J., Akaike, T. & Maruyama, A. DNA strand exchange stimulated by spontaneous complex formation with cationic comb-type copolymer. J. Am. Chem. Soc. 124, 12676–12677 (2002).
Maruyama, A., Katoh, M., Ishihara, T. & Akaike, T. Comb-type polycations effectively stabilize DNA triplex. Bioconjugate Chem. 8, 3–6 (1997).
Torigoe, H., Ferdous, A., Watanabe, H., Akaike, T. & Maruyama, A. Poly(L-lysine)-graft-dextran copolymer promotes pyrimidine motif triplex DNA formation at physiological pH. J. Biol. Chem. 274, 6161–6167 (1999).
Ferdous, A., Watanabe, H., Akaike, T. & Maruyama, A. Poly(L-lysine)-graft-dextran copolymer: amazing effects on triplex stabilization under physiological pH and ionic conditions (in vitro). Nucleic Acids Res. 26, 3949–3954 (1998).
Urbaneja, M.A., Wu, M., Casas-Finet, J.R. & Karpel, R.L. HIV-1 Nucleocapsid protein as a nucleic acid chaperone: spectroscopic study of its helix-destabilizing properties, structural binding specificity, and annealing activity. J. Mol. Biol. 318, 749–764 (2002).
Bazemore, L.R., Takahashi, M. & Radding, C.M. Kinetic analysis of pairing and strand exchange catalyzed by RecA. J. Biol. Chem. 272, 14672–14682 (1997).
Wilson, J.F. et al. Population genetic structure of variable drug response. Nature Genet. 29, 265–269 (2001).
Morais, S.M.F. et al. The major genetic defect responsible for polymorphism of S-mephenytoin metabolism in humans. J. Biol. Chem. 269, 15419–15422 (1994).
Goldstein, J.A. & Morais, S.M.F. Biochemistry and molecular biology of the human CYP2C subfamily. Pharmacogenetics 4, 285–299 (2002).
Elghanian, R., Storhoff, J.J., Mucic, R.C., Letsinger, R.L. & Mirkin, C.A. Selective colorimetric detection of polynucleotides based on the distance-dependent optical properties of gold nanoparticles. Science 277, 1078–1081 (1997).
Kajiyama, T. et al. Genotyping on a thermal gradient DNA chip. Genome Res. 13, 467–475 (2003).
Santalucia, J. Jr. A unified view of polymer, dumbbell, and oligonucleotide DNA nearest-neighbor thermodynamics. Proc. Natl Acad. Sci. USA 95, 1460–1465 (1998).
Chee, M. et al. Accessing genetic information with high-density DNA arrays. Science 274, 610–614 (1996).
Han, M., Gao, X., Su, J.Z. & Nie, S. Quantum-dot-tagged microbeads for multiplexed optical coding of biomolecules. Nature Biotechnol. 19, 631–635 (2001).
Yang, L., Tran, D.K. & Wang, X. BADGE, Beads array for the detection of gene expression, a high-throughput diagnostic bioassay. Genome Res. 11, 1888–1898 (2001).
Boon, E.M., Ceres, D.M., Drummond, T.G., Hill, M.G. & Barton, J.K. Mutation detection by electrocatalysis at DNA-modified electrodes. Nature Biotechnol. 18, 1096–1100 (2000).
Mazumder, A., Majlessi, M. & Becker, M.M. A high throughput method to investigate oligodeoxyribonucleotide hybridisation kinetics and thermodynamics. Nucleic Acids Res. 26, 1996–2000 (1998).
Jenison, R., Yang, S., Haeberli, A. & Polisky, B. Interference-based detection of nucleic acid targets on optically coated silicon. Nature Biotechnol. 19, 62–65 (2001).
Cao, Y.C., Jin, R. & Mirkin, C.A. Nanoparticles with Raman spectroscopic fingerprints for DNA and RNA detection. Science 297, 1536–1540 (2002).
Takahashi, T., Ueno, A. & Mihara, H. Nucleobase amino acids incorporated into the HIV-1 nucleocapsid protein increased the binding affinity and specificity for a hairpin RNA. ChemBioChem 3, 543–549 (2002).
Acknowledgements
We thank H. Mihara for discussion and the gift of NCp7, and D. W. Grainger and K. Yamana for critical reading of the manuscript. We acknowledge FASMAC, for oligonucleotide syntheses and K. Tajima for experimental assistance. This work was supported in part by grant-in-aids (11167225 and 12480260) for scientific research from Ministry of Education, Culture, Sports, Science, and Technology of Japan. W.J.K. was supported by the Japanese Government Scholarship.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information, Fig. S1
Supplementary Information, Fig. S2 (PDF 4171 kb)
Supplementary Information, Fig. S3
Supplementary Information, Fig. S4
Supplementary Information, Fig. S5
Supplementary Information, Fig. S6
Rights and permissions
About this article
Cite this article
Kim, W., Sato, Y., Akaike, T. et al. Cationic comb-type copolymers for DNA analysis. Nature Mater 2, 815–820 (2003). https://doi.org/10.1038/nmat1021
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmat1021
This article is cited by
-
Cationic comb-type copolymer as an artificial chaperone
Polymer Journal (2019)
-
Equilibrious Strand Exchange Promoted by DNA Conformational Switching
Scientific Reports (2013)
-
Targeted polymeric gene delivery for anti-angiogenic tumor therapy
Macromolecular Research (2007)