Abstract
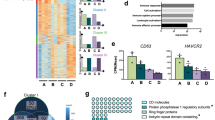
The latency of human immunodeficiency virus type 1 (HIV-1) in resting primary CD4+ T cells is the major barrier for the eradication of the virus in patients on suppressive highly active antiretroviral therapy (HAART). Even with optimal HAART treatment, replication-competent HIV-1 still exists in resting primary CD4+ T cells1,2,3,4. Multiple restriction factors that act upon various steps of the viral life cycle could contribute to viral latency. Here we show that cellular microRNAs (miRNAs) potently inhibit HIV-1 production in resting primary CD4+ T cells. We have found that the 3′ ends of HIV-1 messenger RNAs are targeted by a cluster of cellular miRNAs including miR-28, miR-125b, miR-150, miR-223 and miR-382, which are enriched in resting CD4+ T cells as compared to activated CD4+ T cells. Specific inhibitors of these miRNAs substantially counteracted their effects on the target mRNAs, measured either as HIV-1 protein translation in resting CD4+ T cells transfected with HIV-1 infectious clones, or as HIV-1 virus production from resting CD4+ T cells isolated from HIV-1–infected individuals on suppressive HAART. Our data indicate that cellular miRNAs are pivotal in HIV-1 latency and suggest that manipulation of cellular miRNAs could be a novel approach for purging the HIV-1 reservoir.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Finzi, D. et al. Identification of a reservoir for HIV-1 in patients on highly active antiretroviral therapy. Science 278, 1295–1300 (1997).
Wong, J.K. et al. Recovery of replication-competent HIV despite prolonged suppression of plasma viremia. Science 278, 1291–1295 (1997).
Siliciano, J.D. et al. Long-term follow-up studies confirm the stability of the latent reservoir for HIV-1 in resting CD4+ T cells. Nat. Med. 9, 727–728 (2003).
Persaud, D., Zhou, Y., Siliciano, J.M. & Siliciano, R.F. Latency in human immunodeficiency virus type 1 infection: no easy answers. J. Virol. 77, 1659–1665 (2003).
Kasschau, K.D. et al. P1/HC-Pro, a viral suppressor of RNA silencing, interferes with Arabidopsis development and miRNA function. Dev. Cell 4, 205–217 (2003).
Dunoyer, P., Lecellier, C.H., Parizotto, E.A., Himber, C. & Voinnet, O. Probing the microRNA and small interfering RNA pathways with virus-encoded suppressors of RNA silencing. Plant Cell 16, 1235–1250 (2004).
Lecellier, C.H. et al. A cellular microRNA mediates antiviral defense in human cells. Science 308, 557–560 (2005).
Pfeffer, S. et al. Identification of virus-encoded microRNAs. Science 304, 734–736 (2004).
Gupta, A., Gartner, J.J., Sethupathy, P., Hatzigeorgiou, A.G. & Fraser, N.W. Anti-apoptotic function of a microRNA encoded by the HSV-1 latency-associated transcript. Nature 442, 82–85 (2006).
Bennasser, Y., Le, S.Y., Benkirane, M. & Jeang, K.T. Evidence that HIV-1 encodes an siRNA and a suppressor of RNA silencing. Immunity 22, 607–619 (2005).
Jopling, C.L., Yi, M., Lancaster, A.M., Lemon, S.M. & Sarnow, P. Modulation of hepatitis C virus RNA abundance by a liver-specific MicroRNA. Science 309, 1577–1581 (2005).
Triboulet, R. et al. Suppression of microRNA-silencing pathway by HIV-1 during virus replication. Science 315, 1579–1582 (2007).
Furtado, M.R. et al. Persistence of HIV-1 transcription in peripheral-blood mononuclear cells in patients receiving potent antiretroviral therapy. N. Engl. J. Med. 340, 1614–1622 (1999).
Chun, T.W. et al. Gene expression and viral prodution in latently infected, resting CD4+ T cells in viremic versus aviremic HIV-infected individuals. Proc. Natl. Acad. Sci. USA 100, 1908–1913 (2003).
Lassen, K.G., Ramyar, K.X., Bailey, J.R., Zhou, Y. & Siliciano, R.F. Nuclear retention of multiply spliced HIV-1 RNA in resting CD4+ T cells. PLoS Pathog. 2, e68 (2006).
Zhang, L. et al. Quantifying residual HIV-1 replication in patients receiving combination antiretroviral therapy. N. Engl. J. Med. 340, 1605–1613 (1999).
Patterson, B.K. et al. Persistence of intracellular HIV-1 mRNA correlates with HIV-1–specific immune responses in infected subjects on stable HAART. AIDS 15, 1635–1641 (2001).
Kim, Y.K. et al. Recruitment of TFIIH to the HIV LTR is a rate-limiting step in the emergence of HIV from latency. EMBO J. 25, 3596–3604 (2006).
Williams, S.A. et al. NF-κB p50 promotes HIV latency through HDAC recruitment and repression of transcriptional initiation. EMBO J. 25, 139–149 (2006).
Lassen, K.G., Bailey, J.R. & Siliciano, R.F. Analysis of human immunodeficiency virus type 1 transcriptional elongation in resting CD4+ T cells in vivo. J. Virol. 78, 9105–9114 (2004).
Lewis, B.P., Burge, C.B. & Bartel, D.P. Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell 120, 15–20 (2005).
Didiano, D. & Hobert, O. Perfect seed pairing is not a generally reliable predictor for miRNA-target interactions. Nat. Struct. Mol. Biol. 13, 849–851 (2006).
Chen, C. et al. Real-time quantification of microRNAs by stem-loop RT-PCR. Nucleic Acids Res. 33, e179 (2005).
Pomerantz, R.J., Seshamma, T. & Trono, D. Efficient replication of human immunodeficiency virus type 1 requires a threshold level of Rev: potential implications for latency. J. Virol. 66, 1809–1813 (1992).
Schwartz, S., Felber, B.K., Benko, D.M., Fenyo, E.M. & Pavlakis, G.N. Cloning and functional analysis of multiply spliced mRNA species of human immunodeficiency virus type 1. J. Virol. 64, 2519–2529 (1990).
Chen, K. et al. α-interferon potently enhances the anti-human immunodeficiency virus type 1 activity of APOBEC3G in resting primary CD4 T cells. J. Virol. 80, 7645–7657 (2006).
Wang, F.X. et al. IL-7 is a potent and proviral strain-specific inducer of latent HIV-1 cellular reservoirs of infected individuals on virally suppressive HAART. J. Clin. Invest. 115, 128–137 (2005).
Dornadula, G. et al. Residual HIV-1 RNA in blood plasma of patients taking suppressive highly active antiretroviral therapy. J. Am. Med. Assoc. 282, 1627–1632 (1999).
Butler, S.L., Hansen, M.S. & Bushman, F.D. A quantitative assay for HIV DNA integration in vivo. Nat. Med. 7, 631–634 (2001).
Acknowledgements
We thank K. Zhang for his technical help, J. DeSimone and D. Horn for their assistance in enrolling patients and Y. Wang for conducting X-ray irradiation. This work was supported by grants from the US National Institutes of Health (AI058798 and AI052732) to H.Z. and the National Basic Research Program of China (grant 2004CB518801) to W.H.
Author information
Authors and Affiliations
Contributions
J.H. carried out most experiments. F.W., E.A., H.T., Z.L. and W.H. participated in some of the experiments such as purification of the cells, generation of some plasmid constructs, and DNA or RNA sequencing. K.C. performed some immunoblotting. K.S. and G.V. placed HIV-1–infected individuals on HAART and H.Z. directed and supervised the experiments and interpretation of data. The manuscript was prepared by J.H. and H.Z.
Corresponding author
Ethics declarations
Competing interests
H.Z. and J.H. are going to apply for a patent for the antisense inhibitors to miR-28, miR-125b, miR-150, miR-225 and miR-382 for their possible usage in activating HIV-1 latency.
Supplementary information
Supplementary Text and Figures
Supplementary Figs. 1-10 and Supplementary Tables 1-4 (PDF 1270 kb)
Rights and permissions
About this article
Cite this article
Huang, J., Wang, F., Argyris, E. et al. Cellular microRNAs contribute to HIV-1 latency in resting primary CD4+ T lymphocytes. Nat Med 13, 1241–1247 (2007). https://doi.org/10.1038/nm1639
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nm1639