Abstract
Influenza remains an important disease in humans and animals. In contrast to measles, smallpox and poliomyelitis, influenza is caused by viruses that undergo continuous antigenic change and that possess an animal reservoir. Thus, new epidemics and pandemics are likely to occur in the future, and eradication of the disease will be difficult to achieve. Although it is not clear whether a new pandemic is imminent, it would be prudent to take into account the lessons we have learned from studying different human and animal influenza viruses. Specifically, reconstruction of the genes of the 1918 pandemic virus and studies on their contribution to virulence will be important steps toward understanding the biological capabilities of this lethal virus. Increasing the availability of new antiviral drugs and developing superior vaccines will provide us with better approaches to control influenza and to have a positive impact on disease load. A concern is that the imposition of new rules for working with infectious influenza viruses under high security and high containment conditions will stifle scientific progress. The complex questions of what makes an influenza virus transmissible from one human to another and from one species to another, as well as how the immune system interacts with the virus, will require the active collaboration and unencumbered work of many scientific groups.
Similar content being viewed by others
Main
The genome of influenza A viruses consists of eight single-stranded RNA segments, and the viral particle has two major glycoproteins on its surface: hemagglutinin and neuraminidase (Fig. 1). With at least 15 different hemagglutinin and 9 different neuraminidase subtypes, there is considerable antigenic variation among influenza viruses. Widely circulating human influenza viruses seem to have been limited to three hemagglutinin (H1, H2 and H3) and two neuraminidase (N1 and N2) subtypes; birds are the predominant hosts for the other subtype strains. As a result of changes in the surface glycoproteins of the virus, devastating epidemics and pandemics have occurred in humans; in addition, major epizootics have been reported in poultry, pigs, horses, seals and camels1. Despite great progress in studying the molecular biology of the virus2 and advances in generating influenza viruses from DNA (reverse genetics)3,4, we still lack an understanding of why some influenza virus strains are transmitted well and cause pandemics and what factors lead to disease in one infected animal species and not in another.
The pandemics
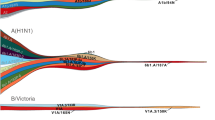
Three pandemics of influenza occurred in the last century. In 1918, the 'Spanish' influenza, a highly contagious and deadly disease, had a major global impact (Fig. 2). In fact, this pandemic (caused by an H1N1 influenza virus) is now known to have been the most deadly in recorded history, with an estimated death toll of 40 million people in less than a year (Fig. 3 and Box 1)5. Other infectious diseases have been as devastating, but in a more protracted way over a far longer period of time. Human immune deficiency virus is the prime example in recent history.
Viruses with three different hemagglutinin subtypes (H1, H2 or H3) and two neuraminidase subtypes (N1 or N2) have been identified in humans. Solid squares indicate the introduction of the pandemic H1N1, H2N2 and H3N2 strains in 1918, 1957 and 1968, respectively. In 1977, H1N1 viruses similar to those of 1950 were reintroduced. Broken lines indicate the absence of virus isolates and only indirect evidence for circulating strains based on serologic data.
Data are adapted from the National Vital Statistics Reports, Vol. 52, No. 14, February 18, 2004. (http://www.cdc.gov/nchs/data/dvs/nvsr52_14t12.pdf).
In 1957, a new influenza virus with two new surface proteins appeared (Fig. 2). The hemagglutinin glycoprotein (H2 subtype) of the 1957 'Asian' influenza virus showed only a 66% amino-acid sequence identity with the hemagglutinin H1 subtype. The N2 subtype neuraminidase of the 1957 virus shared an overall sequence identity of only 37% with the N1 subtype neuraminidase. Thus, in 1957, after 39 years of H1N1 viruses, there was little or no pre-existing immune protection in the human population against this new influenza virus. Even though five out of the eight influenza virus genes were conserved from the H1N1 strains circulating in 1956 and earlier6, the cellular (or humoral) immunity elicited by these gene products was not sufficient to effectively protect humans against the new virus that emerged in 1957. Estimates suggest that 70,000 people died in the United States alone during the Asian influenza pandemic caused by this new H2N2 virus7, and many more succumbed worldwide.
Eleven years later in 1968, another change in the surface glycoproteins again caused the virus to become pandemic, resulting in high morbidity and mortality rates worldwide (Fig. 2). It is estimated that in excess of 30,000 people were killed by the new virus in the US. In this 1968 virus, only the gene that encodes hemagglutinin, HA, and the PB1 gene, were changed6. The H3 and H2 hemagglutinins differed in more than 60% of their amino acids. The conservation of the neuraminidase in the 1968 H3N2 virus may have provided some protection to the population, which had previously been exposed to H2N2 viruses, and this may explain the lower morbidity and mortality numbers as compared with the pandemic in 1957 (ref. 7).
Strictly speaking, there was a fourth pandemic strain in the last century, an H1N1 strain which appeared in 1977 (Fig. 2). This strain was shown by oligonucleotide mapping techniques to be closely related to viruses circulating in 1950 (ref. 8). It caused disease mostly in people born after 1950, because the older population had protective immunity resulting from prior experience with H1N1 strains. Although there is no hard evidence available, the introduction of this 1977 H1N1 virus is now thought to be the result of vaccine trials in the Far East involving the challenge of several thousand military recruits with live H1N1 virus (C.M. Chu, personal communication). Unfortunately, this H1N1 strain (and its descendants) has been circulating ever since, and at present both H3N2 and H1N1 influenza viruses continue to be present in the human population (Fig. 2).
Virulence factors
Despite the fact that influenza generally has a low fatality rate, the high number of infected individuals makes influenza pandemics and epidemics a major health problem. Thus, it is paramount to understand the factors in the host and in the virus that contribute to virulence. Clearly, the outcome of an influenza virus infection is determined by both the host and virus. If the host has had prior exposure to a related strain, the effects of a highly pathogenic strain will be muted. But in an immunologically naive host, virulence is mostly determined by the virus. Even in that situation, virulence is a complex phenomenon. Early on it was recognized that many viral genes can contribute to pathogenicity and that in most instances virulence is a multigenic trait9. For an Olympic runner, strong legs alone are not sufficient to be successful. Similarly, it seems that the most successful virus is one in which all of the parts are optimal and compatible with each other. Arguably, the most virulent influenza virus, the 1918 pandemic strain, had such an optimal constellation of genes and proteins. When the reconstructed HA and NA (which encodes neuraminidase) genes of the 1918 virus are grafted onto the background of another influenza virus, the resulting strain is highly virulent in mice. In contrast, the HA and NA genes of a currently circulating influenza virus put into the same viral background results in a much less virulent strain10,11. Thus, the 1918 hemagglutinin and neuraminidase seem to make a virus intrinsically more virulent (in mice) than the hemagglutinin and neuraminidase of more current influenza virus strains. A similar result using the hemagglutinin of the 1918 virus was obtained recently12. Furthermore, when five '1918' genes are transferred to a mouse-adapted strain, this mixed strain is also highly virulent in mice10, even though the transfer of one of the '1918' genes, the NS gene, which encodes virus nonstructural protein, had previously been shown to attenuate such a virus in mice13. These results suggest that gene constellation is of great importance for the virulence of an influenza virus, and that compatibility of viral genes and proteins frequently defines the success of a virus.
However, a single gene (or a mutation in a single gene) can also markedly affect virulence. We have demonstrated that the NS gene of the 1918 virus, coding for interferon antagonist activity, is not optimal in mice when it replaces the NS gene of a mouse-adapted strain. This probably results from the fact that the viral interferon antagonist activity encoded by the NS gene shows a high degree of species-specificity (a human NS gene being more active in human cells than in mouse cells)14. An NS gene derived from a highly virulent avian H5N1 virus showed similar properties. In this case, a single mutation was shown to considerably alter the phenotype of an influenza virus in a specific host. Pigs infected with a virus carrying a single mutation in the NS1 gene (a glutamic acid present in position 92 of the NS1 protein of the virulent H5N1 virus) experienced substantially greater disease symptoms than animals infected with the control virus15. Another example of a single mutation influencing the outcome of infection in mice concerns a mutation in position 627 of the PB2 gene of an H5N1 influenza virus isolated in 1997 in Hong Kong16. As described for viruses containing genes derived from the 1918 virus or from pathogenic H5N1 viruses, gene combinations as well as specific mutations in a single gene will determine the outcome of a virus infection.
The next influenza pandemic?
“It's tough to make predictions, especially about the future.” This famous saying by Yogi Berra may also apply to influenza. The last century saw pandemic influenza viruses belonging to three subtypes, H1, H2 and H3, and indirect evidence suggests that H3 viruses were circulating from 1889–1918 (ref. 17) and that H1 viruses were possibly prevalent before 1889 (ref. 18). If this series of events over the last hundred years reflects a pattern, recycling of subtypes would be the norm in the human population and the possibility for emergence of new pandemics would be limited. On the other hand, if any subtype is able to thrive in the human population, a greater number of possibilities for novel pandemic strains exists. This is the basis for the apprehension of many people that avian influenza viruses may jump into the human population19,20,21,22.
Although H5N1 avian viruses were shown to cause death in humans in 1997 and more recently in 2004, none of these strains was easily transmitted from person to person. Also, none of the H5N1 strains showed evidence of having acquired genes from circulating human influenza viruses. Whether this is a necessary requirement for a pandemic strain to be successful is not known. It would seem probable that such a reassortment event between an avian and a human influenza virus could have happened many times over, either in humans or in animals. In fact, seroepidemiological studies conducted among the rural population in China suggest that millions of people have been infected with influenza viruses of the H4-to-H15 subtypes. Specifically, seroprevalence levels of 2–7% for H5 viruses alone have been reported23, and the seropositivity of human sera for H7, H10 and H11 viruses was estimated to be as high as 38, 17 and 15% respectively23. These findings predate the recent, highly publicized H5N1 cases.
It may be possible that infections of humans by avian influenza viruses have been ongoing for decades24 and it is only the reporting that has improved in recent years. If this were the case, the present emphasis on the imminent pandemic outbreak21 would not be justified. What is warranted, however, and where there is little or no disagreement among scientists, is a continued surveillance of influenza viruses, not only in humans but also in different animal species and commercial operations. Furthermore, the stockpiling of antiviral drugs and the development of new vaccines is highly recommended to be better prepared for a potential pandemic outbreak. Efforts are being made by the US government to address this problem (see http://www.hhs.gov/nvpo/pandemicplan/index.html and http://www.hhs.gov/news/press/2004pres/20040921a.html).
Are antiviral drugs effective against a pandemic?
Two classes of US Food and Drug Administration–approved drugs are available against influenza A viruses. Amantadine (and rimantadine) target the viral M2 protein25, which forms an ion channel required for the efficient uncoating of incoming viruses (Fig. 4) Unfortunately, there are natural influenza virus isolates (including all influenza B viruses) that are resistant to M2 blockers, and even more ominous drug resistance against amantadine has been reported to readily develop26.
After binding to sialic acid receptors, the virus is internalized by receptor-mediated endocytosis. The low pH in the endosome triggers the fusion of viral and endosomal membranes and the influx of H+ ions through the M2 channel releases the viral genes into the cytoplasm. Amantadine blocks this uncoating step. RNA replication and transcription occur in the nucleus. siRNA inhibition may affect the stability of mRNA, preventing translation of viral protein. Packaging and budding of virions occurs at the cytoplasmic membrane. Neuraminidase inhibitors block the release of the virus from the infected cell. Because sialic acid receptors are not removed by the neuraminidase, aggregates of virus stick to the cytoplasmic membrane of the infected cell and cannot move on to infect other cells.
The second class of drugs consists of inhibitors of the viral neuraminidase protein, which is needed for the efficient release of virus from the infected cell27 (Fig. 4). Both the oral oseltamivir and zanamivir (administered by inhalation) represent important tools for treating influenza virus infections28,29. In the case of oseltamivir, the drug is also approved for prophylaxis of influenza in adults and adolescents 13-years old and older. Influenza viruses recognize sialic acid-containing receptors on the cell surface through their hemagglutinin proteins. Neuraminidase is crucial for removal of these receptors so that the virus does not aggregate by binding to other (sialic acid-containing) virus particles and/or by remaining attached to the surface of infected cells. The neuraminidase is a receptor-destroying enzyme and thus has a crucial role in allowing the ready dispersal of newly generated virus. Drug resistance against neuraminidase inhibitors has been reported in the past. Most recently, however, neuraminidase-inhibitor-resistant strains were found in 18% of Japanese children treated with oseltamivir30. The future will tell whether drug-resistant virus mutants are as virulent and as transmissible in humans as wild-type viruses. It has been shown that drug-resistant mutants with altered neuraminidase activity lose their ability to do harm in mice and ferrets31.
Should there be an outbreak caused by a novel influenza virus, these drugs (M2 blockers and neuraminidase inhibitors) could provide an efficient method of controlling virus spread. Both classes of antiviral drugs have been shown (in a laboratory setting) to be effective against viruses that carry the reconstructed genes from the 1918 virus32. It is thus likely that these drugs would be effective against most emerging influenza virus pathogens. Unfortunately, logistical and financial limitations make it difficult at present to envision these drugs preventing, or even substantially affecting, the emergence of a new pandemic. This picture may change if and when serious consideration is given to stockpiling significant quantities of neuraminidase inhibitors, the drugs of choice at this moment. Only if this were done could targeted antiviral prophylaxis have a chance of success.
What are alternative antiviral approaches for the future? Short interfering RNAs (siRNAs) specific against conserved regions of influenza virus genes have successfully been used to interfere with the replication of influenza viruses in mice33,34. Such a strategy has the advantage that multiple targets (viral genes) could be selected, and if the current problems with delivery in humans were to be resolved, a viable alternative would be available for prophylaxis and therapy of influenza virus infections. In principle, many of the steps necessary for virus replication — including attachment, fusion of viral and cellular membranes, as well as the highly specific virally coded RNA-dependent RNA polymerase—are promising targets for antiviral intervention, but no practical breakthroughs seem to be on the horizon at this time.
Vaccines are the key to controlling a pandemic
The ideal way to reduce the impact of an emerging pandemic is vaccination. The present killed-virus vaccine preparations contain an H1 and an H3 virus (as well as an influenza B virus component). The strains must be frequently changed, even during an interpandemic period, because of continuing antigenic variation in the viral hemagglutinin. For manufacturing purposes, many of the vaccine strains are made to grow to higher titers by reassortment with high-yield master strains35, and similar techniques can be used to develop vaccines against emerging pandemic strains. It may be advantageous to use reverse genetics to construct the actual vaccine candidates. This technology involves the transfection of plasmids expressing influenza virus RNA. Procedures involving the transfection of 12 or more plasmids3,4 or of only 8 'ambisense' plasmids36 have been described (Fig. 5). This reverse genetics approach has advantages over the classical reassortment strategy, including the following: first, if the vaccine candidate is highly pathogenic and has a basic peptide cleavage site in its hemagglutinin protein, the gene encoding this protein can be easily modified, eliminating problems in the manufacturing process; second, the time between choosing the vaccine strain and actually having a suitable candidate for manufacture can be shortened from months to weeks; and third, the vaccine candidate is defined by cloned plasmids (defined sequences) and thus should be preferred by the regulatory agencies.
The HA and NA genes of the candidate strain are cloned. If the hemagglutinin has a multibasic cleavage site (as in the H5 hemagglutinin proteins of highly pathogenic chicken viruses), it can be altered by site-specific mutations. The other six genes of a master donor strain (for example, a high-yield donor, cold-adapted strain, or a novel attenuated strain) are also cloned into appropriate plasmids3. Transfection of these plasmids into cells (along with plasmids expressing the viral polymerase complex) leads to the rescue of a vaccine strain containing the relevant surface glycoproteins (for details and alternatives, see refs. 3,4,36). Reverse genetics techniques can be effectively used to make strains for killed or live influenza virus vaccines.
In addition, reverse genetics allows the implementation of new vaccine strategies which could not be accomplished using classical techniques. For example, changes in the gene coding for the interferon antagonist protein NS1 may lead to improved attenuated vaccine candidates37. It is also possible to envision generating live virus vaccine candidates expressing a cytokine, for example, Il-2 or GMCSF, which should considerably increase the immunogenicity of the influenza virus vaccine. Other ideas include the construction of strains that preferentially express conserved amino acids of the hemagglutinin protein to elicit a crossprotective immune response, or the generation of viruses that express a multiplicity of hemagglutinins, all from a single vaccine strain.
The first live attenuated (cold-adapted) influenza virus vaccine was recently approved by the US Food and Drug Administration, and this vaccine approach may offer advantages as it does not require needles for administration and may provide broader and longer-lasting immune protection38. As effective vaccination against new pandemic strains with killed vaccines would probably require the administration of two doses, a live-virus vaccine involving a single nasal-spray application may be a promising vaccine approach against an emerging pandemic. If vast numbers of immunologically naive individuals need to be vaccinated against a new pandemic strain in a short period of time, improved (genetically engineered) live-virus candidates would represent a major advantage. The use of powerful adjuvants and enhanced cell-culture-based production methods39 should also be considered.
Finally, we should also try to develop novel influenza virus vaccines for the poultry and pig industries. Live Newcastle disease virus vaccines are extensively used, and it is now possible to express the hemagglutinin of influenza viruses from recombinant Newcastle disease virus strains40. In the future, such a combination vaccine may well prevent influenza (as well as Newcastle disease) in commercial chicken and turkey farms, possibly controlling the emergence of pandemics in humans. Widespread vaccination of commercial pigs and of horses may similarly reduce the chances of animal influenza viruses jumping into the human population.
Scientific focus for the future
With respect to the molecular biology of influenza viruses, many of the lower-hanging scientific fruits have already been picked. The next steps must concern the study of more complex systems. Specifically, efforts should be undertaken to better understand viral pathogenicity and human immunology in response to infection. This will involve the use of animal systems and of tools—including genomics and proteomics—that allow the analysis of many parameters. Among the important questions for which we still have no answers are: what makes an influenza virus transmissible in humans and in animals? Why are some cells infected in a susceptible host and others are not? What are the specific interactions between viruses and different immune cells? It is recommended that major efforts be made to identify the viral, host and environmental factors that determine the efficient spread and disease-causing potential of influenza viruses.
One concern related to these future efforts has to do with the possible imposition, in the US as well as in Europe, of new rules and regulations for working with infectious influenza viruses. Such new regulations could have a chilling effect on scientific progress. It is feared that the quest for absolute safety assured by high-containment facilities will drive out of this field many of the good scientists who cherish the freedom of laboratory work and who love asking questions unencumbered by an unnecessary and stifling bureaucracy. In the long run, we will be made less safe by downsizing (or eliminating) a vibrant research community and we will forego the exploitation of new ideas and new strategies by pushing scientific experimentation into closed high-security and high-containment quarters controlled by policing agencies. It is hoped that a risk-benefit calculation can be performed to prevent the needless closing of important and exciting scientific frontiers.
References
Hayden, F. & Palese, P. Influenza Virus. in Clinical Virology (ed. Richman, D.D.) 891–920 (Churchill Livingstone, New York, 1997).
Basler, C. & Palese, P. Influenza Viruses. in Encyclopedia of Molecular Medicine (ed. Creighton, T.) 1741–1747 (John Wiley and Sons, New York, 2002).
Fodor, E. et al. Rescue of influenza A virus from recombinant DNA. J. Virol. 73, 9679–9682 (1999).
Neumann, G. et al. Generation of influenza A viruses entirely from cloned cDNAs. Proc. Natl. Acad. Sci. USA 96, 8804–8806 (1999).
Taubenberger, J.K., Reid, A.H., Janczewski, T.A. & Fanning, T.G. Integrating historical, clinical and molecular genetic data in order to explain the origin and virulence of the 1918 Spanish influenza virus. Philos. Trans. R. Soc. Lond. B Biol. Sci. 356, 1829–1839 (2001).
Kawaoka, Y., Krauss, S. & Webster, R.G. Avian-to-human transmission of the PB1 gene of influenza A viruses in the 1957 and 1968 pandemics. J. Virol. 63, 4603–4608 (1989).
Lipatov, A.S. et al. Influenza: emergence and control. J. Virol. 78, 8951–8959 (2004).
Nakajima, K., Desselberger, U. & Palese, P. Recent human influenza A (H1N1) viruses are closely related genetically to strains isolated in 1950. Nature 274, 334–339 (1978).
Klenk, H.D. & Rott, R. The molecular biology of influenza virus pathogenicity. Adv. Virus Res. 34, 247–281 (1988).
Tumpey, T.M. et al. Pathogenicity and immunogenicity of influenza viruses with genes from the 1918 pandemic virus. Proc. Natl. Acad. Sci. USA 101, 3166–3171 (2004).
Kash, J.C. et al. Global host immune response: pathogenesis and transcriptional profiling of type A influenza viruses expressing the hemagglutinin and neuraminidase genes from the 1918 pandemic virus. J. Virol. 78, 9499–9511 (2004).
Kobasa, D. et al. Enhanced virulence of influenza A viruses with the haemagglutinin of the 1918 pandemic virus. Nature 431, 703–707 (2004).
Basler, C.F. et al. Sequence of the 1918 pandemic influenza virus nonstructural gene (NS) segment and characterization of recombinant viruses bearing the 1918 NS genes. Proc. Natl. Acad. Sci. USA 98, 2746–2751 (2001).
Geiss, G.K. et al. Cellular transcriptional profiling in influenza A virus-infected lung epithelial cells: the role of the nonstructural NS1 protein in the evasion of the host innate defense and its potential contribution to pandemic influenza. Proc. Natl. Acad. Sci. USA 99, 10736–10741 (2002).
Seo, S.H., Hoffmann, E. & Webster, R.G. Lethal H5N1 influenza viruses escape host anti-viral cytokine responses. Nat. Med. 8, 950–954 (2002).
Hatta, M., Gao, P., Halfmann, P. & Kawaoka, Y. Molecular basis for high virulence of Hong Kong H5N1 influenza viruses. Science 293, 1773–1775 (2001).
Dowdle, W.R. Influenza A virus recycling revisited. Bull. World Health Organ. 77, 820–826 (1999).
Zamarin, D. & Palese, P. Influenza virus: Lessons learned. in International Kilmer Conference (eds Kowalski, J.B. & Morissey, J.B.) (Polyscience Publications, Station St. Martin, Laval, Quebec, Canada, in the press).
Katz, J.M. The impact of avian influenza viruses on public health. Avian Dis. 47, 914–920 (2003).
Li, K.S. et al. Genesis of a highly pathogenic and potentially pandemic H5N1 influenza virus in eastern Asia. Nature 430, 209–213 (2004).
Webby, R.J. & Webster, R.G. Are we ready for pandemic influenza? Science 302, 1519–1521 (2003).
Hatta, M. & Kawaoka, Y. The continued pandemic threat posed by avian influenza viruses in Hong Kong. Trends Microbiol. 10, 340–344 (2002).
Shortridge, K.F. Pandemic influenza: a zoonosis? Semin. Respir. Infect. 7, 11–25 (1992).
Profeta, M.L. & Palladino, G. Serological evidence of human infections with avian influenza viruses. Arch. Virol. 90, 355–360 (1986).
Hay, A., Wolstenholme, A., Skehel, J. & Smith, M. The molecular basis of the specific anti-influenza action of amantadine. EMBO J. 4, 3021–3024 (1985).
Monto, A.S. & Arden, N.H. Implications of viral resistance to amantadine in control of influenza A. Clin. Infect. Dis. 15, 362–367 (1992).
Palese, P. & Compans, R.W. Inhibition of influenza virus replication in tissue culture by 2-deoxy- 2,3-dehydro-N-trifluoroacetylneuraminic acid (FANA): mechanism of action. J. Gen. Virol., 33 159–163 (1976).
Mendel, D.B. et al. Oral administration of a prodrug of the influenza virus neuraminidase inhibitor GS 4071 protects mice and ferrets against influenza infection. Antimicrob. Agents Chemother. 42, 640–646 (1998).
Colman, P.M. A novel approach to antiviral therapy for influenza. Antimicrob. Agents Chemother. 44, 17–22 (1999).
Kiso, M. et al. Resistant influenza A viruses in children treated with oseltamivir. A descriptive study. Lancet 364, 759–765 (2004).
Carr, J. et al. Influenza virus carrying neuraminidase with reduced sensitivity to oseltamivir carboxylate has altered properties in vitro and is compromised for infectivity and replicative ability in vivo. Antiviral Res. 54, 79–88 (2002).
Tumpey, T.M. et al. Existing antivirals are effective against influenza viruses with genes from the 1918 pandemic virus. Proc. Natl. Acad. Sci. USA 99, 13849–13854 (2002).
Ge, Q. et al. Inhibition of influenza virus production in virus-infected mice by RNA interference. Proc. Natl. Acad. Sci. USA 101, 8676–8681 (2004).
Tompkins, S.M., Lo, C-Y., Tumpey, T.M. & Epstein, S.L. Protection against lethal influenza virus challenge by RNA interference in vivo. Proc. Natl. Acad. Sci. USA 101, 8682–8686 (2004).
Kilbourne, E.D. et al. Related studies of a recombinant influenza virus vaccine. I. Derivation and characterization of virus and vaccine. J. Infect. Dis. 124, 449–462 (1971).
Hoffmann, E., Neumann, G., Hobom, G., Webster, R.G. & Kawaoka, Y. “Ambisense” approach for the generation of influenza A virus: vRNA and mRNA synthesis from one template. Virology 267, 310–317 (2000).
Talon, J. et al. Influenza A and B viruses expressing altered NS1 proteins: A vaccine approach. Proc. Natl. Acad. Sci. USA 97, 4309–4314 (2000).
Kemble, G. & Greenberg, H. Novel generations of influenza vaccines. Vaccine 21, 1789–1795 (2003).
Palese, P. & Garcia-Sastre, A. Influenza vaccines: present and future. J. Clin. Invest. 110, 9–13 (2002).
Swayne, D.E. et al. Recombinant paramyxovirus type 1-avian influenza-H7 virus as a vaccine for protection of chickens against influenza and Newcastle disease. Avian Dis. 47, 1047–1050 (2003).
Taubenberger, J.K. & Palese, P. The origin and virulence of the 1918 “Spanish” influenza virus. in Contemporary Topics in Influenza Virology (ed. Kawaoka, Y.) (Horizon Scientific Press, Wymondham, Norfolk, UK, in the press).
Glezen, W.P. Emerging infections: pandemic influenza. Epidemiol. Rev. 18, 64–76 (1996).
Reid, A.H., Taubenberger, J.K. & Fanning, T.G. The 1918 Spanish influenza: integrating history and biology. Microbes Infect. 3, 81–87 (2001).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
Peter Palese is a consultant for Avimex Greenhills and Medimmune, as well as a co-author of patents and patent applications owned by Mount Sinai School of Medicine in the field of viral vaccines and antiviral drugs.
Rights and permissions
About this article
Cite this article
Palese, P. Influenza: old and new threats. Nat Med 10 (Suppl 12), S82–S87 (2004). https://doi.org/10.1038/nm1141
Published:
Issue Date:
DOI: https://doi.org/10.1038/nm1141
This article is cited by
-
Influenza pandemics and macroeconomic fluctuations 1871–2016
Cliometrica (2024)
-
Characterization of a reassortant H11N9 subtype avian influenza virus isolated from spot-billed duck in China
Virus Genes (2023)
-
Streptomyces coeruleorubidus as a potential biocontrol agent for Newcastle disease virus
BMC Veterinary Research (2022)
-
Bioinformatic analysis of PD-1 checkpoint blockade response in influenza infection
BMC Genomic Data (2022)
-
Characterization of the physicochemical properties, antioxidant activity, and antiproliferative activity of natural melanin from S. reiliana
Scientific Reports (2022)