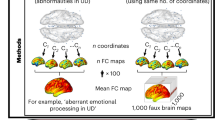
Abstract
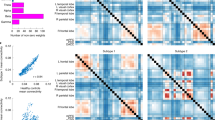
Biomarkers have transformed modern medicine but remain largely elusive in psychiatry, partly because there is a weak correspondence between diagnostic labels and their neurobiological substrates. Like other neuropsychiatric disorders, depression is not a unitary disease, but rather a heterogeneous syndrome that encompasses varied, co-occurring symptoms and divergent responses to treatment. By using functional magnetic resonance imaging (fMRI) in a large multisite sample (n = 1,188), we show here that patients with depression can be subdivided into four neurophysiological subtypes ('biotypes') defined by distinct patterns of dysfunctional connectivity in limbic and frontostriatal networks. Clustering patients on this basis enabled the development of diagnostic classifiers (biomarkers) with high (82–93%) sensitivity and specificity for depression subtypes in multisite validation (n = 711) and out-of-sample replication (n = 477) data sets. These biotypes cannot be differentiated solely on the basis of clinical features, but they are associated with differing clinical-symptom profiles. They also predict responsiveness to transcranial magnetic stimulation therapy (n = 154). Our results define novel subtypes of depression that transcend current diagnostic boundaries and may be useful for identifying the individuals who are most likely to benefit from targeted neurostimulation therapies.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Change history
19 December 2016
In the version of this article initially published online, the abstract contained two typos reading, “Like to other neuropsychiatric disorders,…” and “transcranial-magnetic-stimulation therapy…” . These errors have been corrected in the print, PDF and HTML versions of this article.
References
Insel, T.R. & Cuthbert, B.N. Medicine. Brain disorders? Precisely. Science 348, 499–500 (2015).
Nestler, E.J. & Hyman, S.E. Animal models of neuropsychiatric disorders. Nat. Neurosci. 13, 1161–1169 (2010).
Carroll, B.J. et al. A specific laboratory test for the diagnosis of melancholia. Standardization, validation, and clinical utility. Arch. Gen. Psychiatry 38, 15–22 (1981).
Gold, P.W. & Chrousos, G.P. Organization of the stress system and its dysregulation in melancholic and atypical depression: high vs low CRH/NE states. Mol. Psychiatry 7, 254–275 (2002).
Lewy, A.J., Sack, R.L., Miller, L.S. & Hoban, T.M. Antidepressant and circadian phase-shifting effects of light. Science 235, 352–354 (1987).
Clementz, B.A. et al. Identification of distinct psychosis biotypes using brain-based biomarkers. Am. J. Psychiatry 173, 373–384 (2016).
Hill, S.K. et al. Neuropsychological impairments in schizophrenia and psychotic bipolar disorder: findings from the Bipolar-Schizophrenia Network on Intermediate Phenotypes (B-SNIP) study. Am. J. Psychiatry 170, 1275–1284 (2013).
Wager, T.D. et al. An fMRI-based neurologic signature of physical pain. N. Engl. J. Med. 368, 1388–1397 (2013).
Liston, C. et al. Default mode network mechanisms of transcranial magnetic stimulation in depression. Biol. Psychiatry 76, 517–526 (2014).
Chen, C.-H. et al. Brain imaging correlates of depressive symptom severity and predictors of symptom improvement after antidepressant treatment. Biol. Psychiatry 62, 407–414 (2007).
Salvadore, G. et al. Increased anterior cingulate cortical activity in response to fearful faces: a neurophysiological biomarker that predicts rapid antidepressant response to ketamine. Biol. Psychiatry 65, 289–295 (2009).
Fox, M.D., Buckner, R.L., White, M.P., Greicius, M.D. & Pascual-Leone, A. Efficacy of transcranial magnetic stimulation targets for depression is related to intrinsic functional connectivity with the subgenual cingulate. Biol. Psychiatry 72, 595–603 (2012).
Downar, J. et al. Anhedonia and reward-circuit connectivity distinguish nonresponders from responders to dorsomedial prefrontal repetitive transcranial magnetic stimulation in major depression. Biol. Psychiatry 76, 176–185 (2014).
McGrath, C.L. et al. Toward a neuroimaging treatment selection biomarker for major depressive disorder. JAMA Psychiatry 70, 821–829 (2013).
Greicius, M.D. et al. Resting-state functional connectivity in major depression: abnormally increased contributions from subgenual cingulate cortex and thalamus. Biol. Psychiatry 62, 429–437 (2007).
Drevets, W.C. et al. Subgenual prefrontal cortex abnormalities in mood disorders. Nature 386, 824–827 (1997).
Pezawas, L. et al. 5-HTTLPR polymorphism impacts human cingulate-amygdala interactions: a genetic susceptibility mechanism for depression. Nat. Neurosci. 8, 828–834 (2005).
Mayberg, H.S. et al. Deep brain stimulation for treatment-resistant depression. Neuron 45, 651–660 (2005).
Sheline, Y.I. et al. The default mode network and self-referential processes in depression. Proc. Natl. Acad. Sci. USA 106, 1942–1947 (2009).
Knutson, B., Bhanji, J.P., Cooney, R.E., Atlas, L.Y. & Gotlib, I.H. Neural responses to monetary incentives in major depression. Biol. Psychiatry 63, 686–692 (2008).
Cook, S.C. & Wellman, C.L. Chronic stress alters dendritic morphology in rat medial prefrontal cortex. J. Neurobiol. 60, 236–248 (2004).
Liston, C. et al. Stress-induced alterations in prefrontal cortical dendritic morphology predict selective impairments in perceptual attentional set-shifting. J. Neurosci. 26, 7870–7874 (2006).
Gourley, S.L., Swanson, A.M. & Koleske, A.J. Corticosteroid-induced neural remodeling predicts behavioral vulnerability and resilience. J. Neurosci. 33, 3107–3112 (2013).
Dias-Ferreira, E. et al. Chronic stress causes frontostriatal reorganization and affects decision-making. Science 325, 621–625 (2009).
Power, J.D., Barnes, K.A., Snyder, A.Z., Schlaggar, B.L. & Petersen, S.E. Spurious but systematic correlations in functional connectivity MRI networks arise from subject motion. Neuroimage 59, 2142–2154 (2012).
Satterthwaite, T.D. et al. Impact of in-scanner head motion on multiple measures of functional connectivity: relevance for studies of neurodevelopment in youth. Neuroimage 60, 623–632 (2012).
Van Dijk, K.R.A., Sabuncu, M.R. & Buckner, R.L. The influence of head motion on intrinsic functional connectivity MRI. Neuroimage 59, 431–438 (2012).
Power, J.D. et al. Functional network organization of the human brain. Neuron 72, 665–678 (2011).
Ravasz, E., Somera, A.L., Mongru, D.A., Oltvai, Z.N. & Barabási, A.L. Hierarchical organization of modularity in metabolic networks. Science 297, 1551–1555 (2002).
Rihel, J. et al. Zebrafish behavioral profiling links drugs to biological targets and rest/wake regulation. Science 327, 348–351 (2010).
Wager, T.D., Davidson, M.L., Hughes, B.L., Lindquist, M.A. & Ochsner, K.N. Prefrontal-subcortical pathways mediating successful emotion regulation. Neuron 59, 1037–1050 (2008).
Milad, M.R. & Quirk, G.J. Neurons in medial prefrontal cortex signal memory for fear extinction. Nature 420, 70–74 (2002).
Phelps, E.A., Delgado, M.R., Nearing, K.I. & LeDoux, J.E. Extinction learning in humans: role of the amygdala and vmPFC. Neuron 43, 897–905 (2004).
Graybiel, A.M., Aosaki, T., Flaherty, A.W. & Kimura, M. The basal ganglia and adaptive motor control. Science 265, 1826–1831 (1994).
Pizzagalli, D.A. et al. Reduced caudate and nucleus accumbens response to rewards in unmedicated individuals with major depressive disorder. Am. J. Psychiatry 166, 702–710 (2009).
Ferenczi, E.A. et al. Prefrontal cortical regulation of brainwide circuit dynamics and reward-related behavior. Science 351, aac9698 (2016).
Schultz, W., Dayan, P. & Montague, P.R. A neural substrate of prediction and reward. Science 275, 1593–1599 (1997).
Cardinal, R.N., Parkinson, J.A., Hall, J. & Everitt, B.J. Emotion and motivation: the role of the amygdala, ventral striatum, and prefrontal cortex. Neurosci. Biobehav. Rev. 26, 321–352 (2002).
Gottfried, J.A., O'Doherty, J. & Dolan, R.J. Encoding predictive reward value in human amygdala and orbitofrontal cortex. Science 301, 1104–1107 (2003).
Schultz, W. Behavioral theories and the neurophysiology of reward. Annu. Rev. Psychol. 57, 87–115 (2006).
Rosa, M.J. et al. Sparse network-based models for patient classification using fMRI. Neuroimage 105, 493–506 (2015).
Craddock, R.C., Holtzheimer, P.E. III, Hu, X.P. & Mayberg, H.S. Disease state prediction from resting state functional connectivity. Magn. Reson. Med. 62, 1619–1628 (2009).
Zeng, L.L. et al. Identifying major depression using whole-brain functional connectivity: a multivariate pattern analysis. Brain 135, 1498–1507 (2012).
Nielsen, J.A. et al. Multisite functional connectivity MRI classification of autism: ABIDE results. Front. Hum. Neurosci. 7, 599 (2013).
Plitt, M., Barnes, K.A. & Martin, A. Functional connectivity classification of autism identifies highly predictive brain features but falls short of biomarker standards. Neuroimage Clin. 7, 359–366 (2014).
Hastie, T., Tibshirani, R. & Friedman, J. The Elements of Statistical Learning: Data Mining, Inference, and Prediction (Springer-Verlag, 2009).
George, M.S. et al. Daily repetitive transcranial magnetic stimulation (rTMS) improves mood in depression. Neuroreport 6, 1853–1856 (1995).
Pascual-Leone, A., Rubio, B., Pallardó, F. & Catalá, M.D. Rapid-rate transcranial magnetic stimulation of left dorsolateral prefrontal cortex in drug-resistant depression. Lancet 348, 233–237 (1996).
Huang, Y.-Z., Rothwell, J.C., Edwards, M.J. & Chen, R.-S. Effect of physiological activity on an NMDA-dependent form of cortical plasticity in human. Cereb. Cortex 18, 563–570 (2008).
Davidson, R.J., Pizzagalli, D., Nitschke, J.B. & Putnam, K. Depression: perspectives from affective neuroscience. Annu. Rev. Psychol. 53, 545–574 (2002).
Oathes, D.J., Patenaude, B., Schatzberg, A.F. & Etkin, A. Neurobiological signatures of anxiety and depression in resting-state functional magnetic resonance imaging. Biol. Psychiatry 77, 385–393 (2015).
Goodkind, M. et al. Identification of a common neurobiological substrate for mental illness. JAMA Psychiatry 72, 305–315 (2015).
Baker, J.T. et al. Disruption of cortical association networks in schizophrenia and psychotic bipolar disorder. JAMA Psychiatry 71, 109–118 (2014).
Sullivan, P.F., Neale, M.C. & Kendler, K.S. Genetic epidemiology of major depression: review and meta-analysis. Am. J. Psychiatry 157, 1552–1562 (2000).
Ripke, S. et al. A mega-analysis of genome-wide association studies for major depressive disorder. Mol. Psychiatry 18, 497–511 (2013).
Pankevich, D.E., Altevogt, B.M., Dunlop, J., Gage, F.H. & Hyman, S.E. Improving and accelerating drug development for nervous system disorders. Neuron 84, 546–553 (2014).
Krishnan, V. et al. Molecular adaptations underlying susceptibility and resistance to social defeat in brain reward regions. Cell 131, 391–404 (2007).
Chaudhury, D. et al. Rapid regulation of depression-related behaviours by control of midbrain dopamine neurons. Nature 493, 532–536 (2013).
Tye, K.M. et al. Amygdala circuitry mediating reversible and bidirectional control of anxiety. Nature 471, 358–362 (2011).
Power, J.D., Barnes, K.A., Snyder, A.Z., Schlaggar, B.L. & Petersen, S.E. Steps toward optimizing motion artifact removal in functional connectivity MRI; a reply to Carp. Neuroimage 76, 439–441 (2013).
Carp, J. Optimizing the order of operations for movement scrubbing: Comment on Power et al. Neuroimage 76, 436–438 (2013).
Jo, H.J. et al. Effective preprocessing procedures virtually eliminate distance-dependent motion artifacts in resting state fMRI. J. Appl. Math. 2013, 935154 (2013).
Jo, H.J., Saad, Z.S., Simmons, W.K., Milbury, L.A. & Cox, R.W. Mapping sources of correlation in resting state FMRI, with artifact detection and removal. Neuroimage 52, 571–582 (2010).
Power, J.D., Schlaggar, B.L. & Petersen, S.E. Recent progress and outstanding issues in motion correction in resting state fMRI. Neuroimage 105, 536–551 (2015).
Desikan, R.S. et al. An automated labeling system for subdividing the human cerebral cortex on MRI scans into gyral based regions of interest. Neuroimage 31, 968–980 (2006).
Shirer, W.R., Ryali, S., Rykhlevskaia, E., Menon, V. & Greicius, M.D. Decoding subject-driven cognitive states with whole-brain connectivity patterns. Cereb. Cortex 22, 158–165 (2012).
Chang, C.-C. & Lin, C.-J. LIBSVM: A library for support vector machines. ACM Trans. Intell. Syst. Technol. 2, 27 (2011).
Dosenbach, N.U.F. et al. Prediction of individual brain maturity using fMRI. Science 329, 1358–1361 (2010).
Van Dijk, K.R.A. et al. Intrinsic functional connectivity as a tool for human connectomics: theory, properties, and optimization. J. Neurophysiol. 103, 297–321 (2010).
Salomons, T.V. et al. Resting-state cortico-thalamic-striatal connectivity predicts response to dorsomedial prefrontal rTMS in major depressive disorder. Neuropsychopharmacology 39, 488–498 (2014).
Bakker, N. et al. rTMS of the dorsomedial prefrontal cortex for major depression: safety, tolerability, effectiveness, and outcome predictors for 10 Hz versus intermittent theta-burst stimulation. Brain Stimul. 8, 208–215 (2015).
Acknowledgements
We wish to thank all investigators who volunteered to share MRI data via the 1000 Functional Connectomes Project (http://fcon_1000.projects.nitrc.org/index.html), which was supported by grants from the NIMH, NIDA, Autism Speaks, NINDS and HHMI. Principal investigators from sites that provided data used here include: R.L. Buckner (Harvard–MGH), F.X. Castellanos (NYU), A.C. Evans (ICBM), B. Leventhal (Nathan Kline Institute), S.J. Li (Medical College of Wisconsin), M.J. Lowe (Cleveland Clinic), H.M. Mayberg (Emory), M.P. Milham (Nathan Kline Institute), V. Riedl (Munchen), C. Sorg (Munchen), A. Villringer (Leipzig) and Y.F. Zang (Beijing Normal University). We also thank the following investigators at the University of New Mexico who provided public access to MRI data from patients diagnosed with schizophrenia through the Center of Biomedical Research Excellence in Brain Function and Mental Illness (COBRE): C. Aine, V. Calhoun, J. Canive, F. Hanlon, R. Jung, K. Kiehl, A. Mayer, N. Perrone-Bizzozero, J. Stephen and C. Tesche, who were supported by NIH COBRE grant 1P20RR021938-01A2. We also thank D. Fair (OHSU) and J. Power (NIMH, Weill Cornell) for providing comments on the data analysis, as well as members of the Liston Lab and Sackler Institute, for their helpful comments on the manuscript. H.S.M. was supported by a grant from the NIMH (P50 MH077083). C.L. was supported by grants from the Dana Foundation, Hartwell Foundation, International Mental Health Research Organization, Klingenstein-Simons Foundations, NARSAD and NIMH (R00 MH097822, R01 MH109685).
Author information
Authors and Affiliations
Contributions
J.D., K.D., F.M., D.J.O., A.E., A.F.S., K.S., J.K., H.S.M., F.M.G., G.S.A., M.D.F., A.P.-L., H.U.V., B.J.C., M.J.D. and C.L. collected the data. L.G. consulted on all statistical analyses. C.L. designed the protocol for analyzing data pooled across multiple sites and identifying clusters. A.T.D., R.F. and C.L. designed and implemented the preprocessing pipeline and methods for validating clusters and optimizing classifiers, and C.L. developed and implemented the method for clustering and classification in a low-dimensional connectivity-feature space by using canonical correlation analysis (Figs. 1,2,3). J.D., K.D. and F.M. collected the TMS data. C.L. analyzed the TMS response data and other clinical data (Figs. 2 and 4) and tested the subtype classifiers on subjects with other diagnoses (Fig. 5). A.T.D., Y.M. and C.L. implemented the permutation testing. A.T.D., B.Z. and C.L. created the figures and wrote the manuscript. All authors discussed the results and conclusions and edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
C.L. and A.T.D. have no competing interests. A.E. has received a research grant from Brain Resource. A.F.S. has served as a consultant to BrainCells, CeNeRx, BioPharma, CNS Response, Corcept Therapeutics, Eli Lilly, Forest Laboratories, GlaxoSmithKline, InnoPharma, Lundbeck, Merck, Neuronetics, Novartis, Pathway Diagnostics, Pfizer, PharmaNeuroBoost, Quintiles, Sanofi, Synosis, Takeda, and Xytis; has equity in Amnestix, BrainCells, CeNeRx, Corcept Therapeutics, Forest Laboratories, Merck, Neurocrine Biosciences, Pfizer, PharmaNeuroBoost, Somaxon Pharmaceuticals and Synosis; has pharmacogenetic-use patents on the prediction of antidepressant response; and has received speaking fees from GlaxoSmithKline and Roche. G.S.A. has received grant support from Forest Pharmaceuticals; has consulted for Hoffman–LaRoche, Lilly, Pfizer and Otsuka; and has served at the speakers′ bureaus of AstraZeneca, Avanir, Forest, Merck, Novartis and Sunovion. M.J.D. has received research grants from Neuronetics. All other authors report no biomedical financial interests or potential conflicts of interest.
Supplementary information
Supplementary Figures and Tables
Supplementary Figures 1–7 and Supplementary Tables 1–7 (PDF 7125 kb)
Rights and permissions
About this article
Cite this article
Drysdale, A., Grosenick, L., Downar, J. et al. Resting-state connectivity biomarkers define neurophysiological subtypes of depression. Nat Med 23, 28–38 (2017). https://doi.org/10.1038/nm.4246
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nm.4246
This article is cited by
-
Interaction of 5-HTTLPR and SLE disease status on resting-state brain function
Arthritis Research & Therapy (2024)
-
Limited generalizability of multivariate brain-based dimensions of child psychiatric symptoms
Communications Psychology (2024)
-
Major depressive disorder: hypothesis, mechanism, prevention and treatment
Signal Transduction and Targeted Therapy (2024)
-
Addiction biotypes: a paradigm shift for future treatment strategies?
Molecular Psychiatry (2024)
-
Novel mechanism-based treatments for pediatric anxiety and depressive disorders
Neuropsychopharmacology (2024)