Abstract
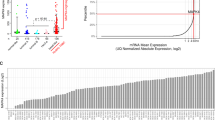
Triple-negative breast cancer (TNBC), in which cells lack expression of the estrogen receptor (ER), the progesterone receptor (PR) and the ERBB2 (also known as HER2) receptor, is the breast cancer subtype with the poorest outcome1. No targeted therapy is available against this subtype of cancer owing to a lack of validated molecular targets. We previously reported that signaling involving MYC—an essential, pleiotropic transcription factor that regulates the expression of hundreds of genes—is disproportionally higher in triple-negative (TN) tumors than in receptor-positive (RP) tumors2. Direct inhibition of the oncogenic transcriptional activity of MYC has been challenging to achieve3. Here, by conducting a shRNA screen targeting the kinome, we identified PIM1, a non-essential serine–threonine kinase, in a synthetic lethal interaction with MYC. PIM1 expression was higher in TN tumors than in RP tumors and was associated with poor prognosis in patients with hormone- and HER2-negative tumors. Small-molecule PIM kinase inhibitors halted the growth of human TN tumors with elevated MYC expression in patient-derived tumor xenograft (PDX) and MYC-driven transgenic mouse models of breast cancer by inhibiting the oncogenic transcriptional activity of MYC and restoring the function of the endogenous cell cycle inhibitor, p27. Our findings warrant clinical evaluation of PIM kinase inhibitors in patients with TN tumors that have elevated MYC expression.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Foulkes, W.D., Smith, I.E. & Reis-Filho, J.S. Triple-negative breast cancer. N. Engl. J. Med. 363, 1938–1948 (2010).
Horiuchi, D. et al. MYC pathway activation in triple-negative breast cancer is synthetic lethal with CDK inhibition. J. Exp. Med. 209, 679–696 (2012).
Prochownik, E.V. & Vogt, P.K. Therapeutic targeting of Myc. Genes Cancer 1, 650–659 (2010).
Kaelin, W.G. Jr . The concept of synthetic lethality in the context of anticancer therapy. Nat. Rev. Cancer 5, 689–698 (2005).
Mukhopadhyay, R. et al. Promotion of variant human mammary epithelial cell outgrowth by ionizing radiation: an agent-based model supported by in vitro studies. Breast Cancer Res. 12, R11 (2010).
Littlewood, T.D., Hancock, D.C., Danielian, P.S., Parker, M.G. & Evan, G.I. A modified estrogen receptor ligand-binding domain as an improved switch for the regulation of heterologous proteins. Nucleic Acids Res. 23, 1686–1690 (1995).
Kessler, J.D. et al. A SUMOylation-dependent transcriptional subprogram is required for Myc-driven tumorigenesis. Science 335, 348–353 (2012).
Sato, M. et al. MYC is a critical target of FBXW7. Oncotarget 6, 3292–3305 (2015).
Bazarov, A.V. et al. Telomerase activation by c-MYC in human mammary epithelial cells requires additional genomic changes. Cell Cycle 8, 3373–3378 (2009).
Nawijn, M.C., Alendar, A. & Berns, A. For better or for worse: the role of Pim oncogenes in tumorigenesis. Nat. Rev. Cancer 11, 23–34 (2011).
Mikkers, H. et al. Mice deficient for all PIM kinases display reduced body size and impaired responses to hematopoietic growth factors. Mol. Cell. Biol. 24, 6104–6115 (2004).
van Lohuizen, M. et al. Predisposition to lymphomagenesis in Pim1 transgenic mice: cooperation with c-Myc and N-Myc in murine leukemia virus–induced tumors. Cell 56, 673–682 (1989).
Wang, J. et al. Pim1 kinase synergizes with c-MYC to induce advanced prostate carcinoma. Oncogene 29, 2477–2487 (2010).
Wang, J. et al. Pim1 kinase is required to maintain tumorigenicity in MYC-expressing prostate cancer cells. Oncogene 31, 1794–1803 (2012).
Kirschner, A.N. et al. PIM kinase inhibitor AZD1208 for treatment of MYC-driven prostate cancer. J. Natl. Cancer Inst. 107, dju407 (2014).
Xie, Y. et al. The 44-kDa Pim-1 kinase directly interacts with tyrosine kinase ETK (BMX) and protects human prostate cancer cells from apoptosis induced by chemotherapeutic drugs. Oncogene 25, 70–78 (2006).
Malinen, M. et al. Proto-oncogene PIM1 is a novel estrogen receptor target associating with high-grade breast tumors. Mol. Cell. Endocrinol. 365, 270–276 (2013).
Chen, L.S., Redkar, S., Bearss, D., Wierda, W.G. & Gandhi, V. Pim kinase inhibitor SGI-1776 induces apoptosis in chronic lymphocytic leukemia cells. Blood 114, 4150–4157 (2009).
DeRose, Y.S. et al. Tumor grafts derived from women with breast cancer authentically reflect tumor pathology, growth, metastasis and disease outcomes. Nat. Med. 17, 1514–1520 (2011).
Lu, J. et al. Pim2 is required for maintaining multiple myeloma cell growth through modulating TSC2 phosphorylation. Blood 122, 1610–1620 (2013).
Burger, M.T. et al. Identification of N-(4-((1R,3S,5S)-3-amino-5-methylcyclohexyl)pyridin-3-yl)-6-(2,6-difluorophenyl)-5-fluoropicolinamide (PIM447), a potent and selective proviral insertion site of Moloney murine leukemia (PIM) 1, 2 and 3 kinase inhibitor in clinical trials for hematological malignancies. J. Med. Chem. 58, 8373–8386 (2015).
D'Cruz, C.M. et al. c-MYC induces mammary tumorigenesis by means of a preferred pathway involving spontaneous Kras2 mutations. Nat. Med. 7, 235–239 (2001).
Zhang, Y., Wang, Z., Li, X. & Magnuson, N.S. Pim-kinase-dependent inhibition of c-Myc degradation. Oncogene 27, 4809–4819 (2008).
Sears, R.C. The life cycle of c-Myc: from synthesis to degradation. Cell Cycle 3, 1133–1137 (2004).
Zhang, Y., Wang, Z. & Magnuson, N.S. Pim-1-kinase-dependent phosphorylation of p21Cip1 (WAF1) regulates its stability and cellular localization in H1299 cells. Mol. Cancer Res. 5, 909–922 (2007).
Morishita, D., Katayama, R., Sekimizu, K., Tsuruo, T. & Fujita, N. Pim kinases promote cell cycle progression by phosphorylating and downregulating p27Kip1 at the transcriptional and post-transcriptional levels. Cancer Res. 68, 5076–5085 (2008).
Bachmann, M. et al. The oncogenic serine–threonine kinase Pim-1 directly phosphorylates and activates the G2/M-specific phosphatase Cdc25C. Int. J. Biochem. Cell Biol. 38, 430–443 (2006).
Mochizuki, T. et al. Physical and functional interactions between Pim-1 kinase and Cdc25A phosphatase. Implications for the Pim-1-mediated activation of the c-Myc signaling pathway. J. Biol. Chem. 274, 18659–18666 (1999).
Yan, B. et al. The PIM-2 kinase phosphorylates BAD on serine 112 and reverses BAD-induced cell death. J. Biol. Chem. 278, 45358–45367 (2003).
Fox, C.J. et al. The serine–threonine kinase Pim-2 is a transcriptionally regulated apoptotic inhibitor. Genes Dev. 17, 1841–1854 (2003).
Wang, X. et al. Phosphorylation regulates c-Myc's oncogenic activity in the mammary gland. Cancer Res. 71, 925–936 (2011).
O'Donnell, K.A., Wentzel, E.A., Zeller, K.I., Dang, C.V. & Mendell, J.T. c-Myc-regulated microRNAs modulate E2F1 expression. Nature 435, 839–843 (2005).
Brasó-Maristany, F. et al. PIM1 kinase regulates cell death, tumor growth and chemotherapy response in triple-negative breast cancer. Nat. Med. 22, 1303–1313 (2016).
Mohanty, A.R. et al. Successive phosphorylation of p27KIP1 protein at Ser10 and C terminus crucially controls its potency to inactivate Cdk2. J. Biol. Chem. 287, 21757–21764 (2012).
Fujita, N., Sato, S., Katayama, K. & Tsuruo, T. Akt-dependent phosphorylation of p27Kip1 promotes binding to 14-3-3 and cytoplasmic localization. J. Biol. Chem. 277, 28706–28713 (2002).
Czabotar, P.E., Lessene, G., Strasser, A. & Adams, J.M. Control of apoptosis by the BCL-2 protein family: implications for physiology and therapy. Nat. Rev. Mol. Cell Biol. 15, 49–63 (2014).
Chu, I.M., Hengst, L. & Slingerland, J.M. The Cdk inhibitor p27 in human cancer: prognostic potential and relevance to anticancer therapy. Nat. Rev. Cancer 8, 253–267 (2008).
Cancer Genome Atlas Network. Comprehensive molecular portraits of human breast tumors. Nature 490, 61–70 (2012).
Narita, M. et al. Rb-mediated heterochromatin formation and silencing of E2F target genes during cellular senescence. Cell 113, 703–716 (2003).
Bazarov, A.V. et al. p16INK4a-mediated suppression of telomerase in normal and malignant human breast cells. Aging Cell 9, 736–746 (2010).
Yau, C. et al. A multigene predictor of metastatic outcome in early-stage hormone-receptor-negative and triple-negative breast cancer. Breast Cancer Res. 12, R85 (2010).
Esserman, L.J. et al. Chemotherapy response and recurrence-free survival in neoadjuvant breast cancer depends on biomarker profiles: results from the I-SPY 1 TRIAL (CALGB 150007/150012; ACRIN 6657). Breast Cancer Res. Treat. 132, 1049–1062 (2012).
Hatzis, C. et al. A genomic predictor of response and survival following taxane-anthracycline chemotherapy for invasive breast cancer. J. Am. Med. Assoc. 305, 1873–1881 (2011).
Davis, S. & Meltzer, P.S. GEOquery: a bridge between the Gene Expression Omnibus (GEO) and BioConductor. Bioinformatics 23, 1846–1847 (2007).
Langfelder, P. & Horvath, S. WGCNA: an R package for weighted correlation network analysis. BMC Bioinformatics 9, 559 (2008).
Langfelder, P. & Horvath, S. Fast R functions for robust correlations and hierarchical clustering. J. Stat. Softw. 46, i11 (2012).
Wickham, H. ggplot2 (Springer Science and Business Media, 2009).
Therneau, T.M. Modeling Survival Data: Extending the Cox Model (Springer Science and Business Media, 2000).
Neve, R.M. et al. A collection of breast cancer cell lines for the study of functionally distinct cancer subtypes. Cancer Cell 10, 515–527 (2006).
Chen, L.S., Redkar, S., Taverna, P., Cortes, J.E. & Gandhi, V. Mechanisms of cytotoxicity to Pim kinase inhibitor, SGI-1776, in acute myeloid leukemia. Blood 118, 693–702 (2011).
Garcia, P.D. et al. Pan–PIM kinase inhibition provides a novel therapy for treating hematologic cancers. Clin. Cancer Res. 20, 1834–1845 (2014).
Acknowledgements
This work was supported in part by grants from the US National Institutes of Health (K99CA175700 (D.H.), R00CA175700 (D.H.), 5T32DK007418 (R.C.), K99CA181490 (K.K.), ES019458 (P.Y. and Z.W.), U01CA168370 (M.T.M.), P30DK63720 (M.T.M.), R01CA180039 (Z.W.) and R01CA170447 (A.G.)), the Susan G. Komen Foundation (PDF15331114; J.R.), the UCSF Program for Breakthrough Biomedical Research (M.T.M.), an Innovative, Developmental, and Exploratory Award from the California Breast Cancer Research Program (17lB-0024; A.G.), an Era of Hope Scholar Award from the CDMRP Breast Cancer Research Program (W81XWH-12-1-0272 and W81XWH-16-1-0603; both to A.G.), an LLS Scholar Award (A.G.), a V-Foundation Award (A.G.), the Breast Cancer Research Foundation (H.S.R. and A.G.) and the Northwestern Medicine Catalyst Funds (D.H.). The authors thank A. Welm for her guidance with the use of the patient-derived orthotopic tumor xenograft models, J.W. Smyth for his assistance with the generation of the transgenic breast cancer cell lines, and D.B. Udy, C.L. Hueschen and A. Vasilopoulos for their assistance with microscopy. We thank S. Samson, C. Baas, H. Klein-Connolly and D. Roth for consumer advocacy support and feedback related to this project, and J.M. Bishop for his insights into the project and his mentorship to D.H.
Author information
Authors and Affiliations
Contributions
D.H. and A.G. conceived the project; D.H. designed and executed the shRNA screen and the subsequent biological experiments, analyzed the data and wrote the manuscript; A.V.B., M.T.M. and P.Y. provided materials for, and contributed to, designing and executing the screen; C.Y. and S.B. performed bioinformatics analyses; D.A.L., H.S.R. and Z.W. provided materials for, and contributed to, designing and executing the animal experiments involving the patient-derived orthotopic tumor xenografts; R.C., A.Y.Z., A.N.C., H.E., K.K., L.A.M., B.N.A., J.R. and R.K. contributed to executing the biological experiments; R.C., A.Y.Z., C.Y., S.B., L.A.M., B.N.A., P.Y., Z.W., O.M. and A.G. participated in the writing of the manuscript; and A.G. supervised the project.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text, Figures and Tables
Suppplementary Tables 1–2 and Supplementary Figures 1–21 (PDF 11674 kb)
Rights and permissions
About this article
Cite this article
Horiuchi, D., Camarda, R., Zhou, A. et al. PIM1 kinase inhibition as a targeted therapy against triple-negative breast tumors with elevated MYC expression. Nat Med 22, 1321–1329 (2016). https://doi.org/10.1038/nm.4213
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nm.4213