Abstract
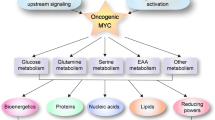
Expression of the oncogenic transcription factor MYC is disproportionately elevated in triple-negative breast cancer (TNBC), as compared to estrogen receptor–, progesterone receptor– or human epidermal growth factor 2 receptor–positive (RP) breast cancer1,2. We and others have shown that MYC alters metabolism during tumorigenesis3,4. However, the role of MYC in TNBC metabolism remains mostly unexplored. We hypothesized that MYC-dependent metabolic dysregulation is essential for the growth of MYC-overexpressing TNBC cells and may identify new therapeutic targets for this clinically challenging subset of breast cancer. Using a targeted metabolomics approach, we identified fatty acid oxidation (FAO) intermediates as being dramatically upregulated in a MYC-driven model of TNBC. We also identified a lipid metabolism gene signature in patients with TNBC that were identified from The Cancer Genome Atlas database and from multiple other clinical data sets, implicating FAO as a dysregulated pathway that is critical for TNBC cell metabolism. We found that pharmacologic inhibition of FAO catastrophically decreased energy metabolism in MYC-overexpressing TNBC cells and blocked tumor growth in a MYC-driven transgenic TNBC model and in a MYC-overexpressing TNBC patient–derived xenograft. These findings demonstrate that MYC-overexpressing TNBC shows an increased bioenergetic reliance on FAO and identify the inhibition of FAO as a potential therapeutic strategy for this subset of breast cancer.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Horiuchi, D. et al. MYC pathway activation in triple-negative breast cancer is synthetic lethal with CDK inhibition. J. Exp. Med. 209, 679–696 (2012).
Koboldt, D.C. et al. Comprehensive molecular portraits of human breast tumors. Nature 490, 61–70 (2012).
Hu, S. et al. [13C]pyruvate imaging reveals alterations in glycolysis that precede c-Myc–induced tumor formation and regression. Cell Metab. 14, 131–142 (2011).
Wahlström, T. & Henriksson, M.A. Impact of MYC in regulation of tumor cell metabolism. Biochim. Biophys. Acta 1849, 563–569 (2015).
D'Cruz, C.M. et al. c-MYC induces mammary tumorigenesis by means of a preferred pathway involving spontaneous Kras2 mutations. Nat. Med. 7, 235–239 (2001).
Pfefferle, A.D. et al. Transcriptomic classification of genetically engineered mouse models of breast cancer identifies human subtype counterparts. Genome Biol. 14, R125 (2013).
Carracedo, A. et al. A metabolic prosurvival role for PML in breast cancer. J. Clin. Invest. 122, 3088–3100 (2012).
Zaugg, K. et al. Carnitine palmitoyltransferase 1C promotes cell survival and tumor growth under conditions of metabolic stress. Genes Dev. 25, 1041–1051 (2011).
Bonnefont, J.P. et al. Carnitine palmitoyltransferases 1 and 2: biochemical, molecular and medical aspects. Mol. Aspects Med. 25, 495–520 (2004).
Esserman, L.J. et al. Chemotherapy response and recurrence-free survival in neoadjuvant breast cancer depends on biomarker profiles: results from the I-SPY 1 TRIAL (CALGB 150007/150012; ACRIN 6657). Breast Cancer Res. Treat. 132, 1049–1062 (2012).
Hatzis, C. et al. A genomic predictor of response and survival following taxane-anthracycline chemotherapy for invasive breast cancer. JAMA 305, 1873–1881 (2011).
Yau, C. et al. A multigene predictor of metastatic outcome in early-stage hormone receptor–negative and triple-negative breast cancer. Breast Cancer Res. 12, R85 (2010).
Chin, K. et al. Genomic and transcriptional aberrations linked to breast cancer pathophysiologies. Cancer Cell 10, 529–541 (2006).
Morris, E.M. et al. PGC-1α overexpression results in increased hepatic fatty acid oxidation with reduced triacylglycerol accumulation and secretion. Am. J. Physiol. Gastrointest. Liver Physiol. 303, G979–G992 (2012).
Neve, R.M. et al. A collection of breast cancer cell lines for the study of functionally distinct cancer subtypes. Cancer Cell 10, 515–527 (2006).
Holubarsch, C.J.F. et al. A double-blind randomized multicenter clinical trial to evaluate the efficacy and safety of two doses of etomoxir, in comparison with placebo, in patients with moderate congestive heart failure: the ERGO (etomoxir for the recovery of glucose oxidation) study. Clin. Sci. 113, 205–212 (2007).
Cowley, G.S. et al. Parallel genome-scale loss-of-function screens in 216 cancer cell lines for the identification of context-specific genetic dependencies. Sci. Data 1, 140035 (2014).
Jeon, S.M., Chandel, N.S. & Hay, N. AMPK regulates NADPH homeostasis to promote tumor cell survival during energy stress. Nature 485, 661–665 (2012).
Schafer, Z.T. et al. Antioxidant and oncogene rescue of metabolic defects caused by loss of matrix attachment. Nature 461, 109–113 (2009).
Shen, L. et al. Metabolic reprogramming in triple-negative breast cancer through Myc suppression of TXNIP. Proc. Natl. Acad. Sci. USA 112, 5425–5430 (2015).
Bensaad, K. et al. Fatty acid uptake and lipid storage induced by HIF-1α contribute to cell growth and survival after hypoxia-reoxygenation. Cell Rep. 9, 349–365 (2014).
DeRose, Y.S. et al. Tumor grafts derived from women with breast cancer authentically reflect tumor pathology, growth, metastasis and disease outcomes. Nat. Med. 17, 1514–1520 (2011).
Davis, S. & Meltzer, P.S. GEOquery: a bridge between the Gene Expression Omnibus (GEO) and BioConductor. Bioinformatics 23, 1846–1847 (2007).
Langfelder, P. & Horvath, S. WGCNA: an R package for weighted correlation network analysis. BMC Bioinformatics 9, 559 (2008).
Langfelder, P. & Horvath, S. Fast R functions for robust correlations and hierarchical clustering. J. Stat. Softw. 46, i11 (2012).
Mukhopadhyay, R. et al. Promotion of variant human mammary epithelial cell outgrowth by ionizing radiation: an agent-based model supported by in vitro studies. Breast Cancer Res. 12, R11 (2010).
Narita, M. et al. Rb-mediated heterochromatin formation and silencing of E2F target genes during cellular senescence. Cell 113, 703–716 (2003).
Littlewood, T.D., Hancock, D.C., Danielian, P.S., Parker, M.G. & Evan, G.I. A modified estrogen receptor ligand-binding domain as an improved switch for the regulation of heterologous proteins. Nucleic Acids Res. 23, 1686–1690 (1995).
Smyth, G.K. in Bioinformatics and Computational Biology Solutions Using R Bioconductor (eds. Gentleman, R., Carey, V.J., Huber, W., Irizarry, R.A. & Dudoit, S.) 397–420 (Springer, 2005).
Väremo, L., Nielsen, J. & Nookaew, I. Enriching the gene set analysis of genome-wide data by incorporating directionality of gene expression and combining statistical hypotheses and methods. Nucleic Acids Res. 41, 4378–4391 (2013).
Acknowledgements
This work was supported, in part, by the US Department of Defense–Congressionally Directed Medical Research Programs' Era of Hope Scholar award W81XWH-12-1-0272 (A.G.), the US National Institutes of Health (NIH) grant R01 CA170447 (A.G.), the Diabetes, Endocrinology and Metabolism training grant T32 DK007418 (R.C.), the Molecular and Cellular Mechanisms of Cancer training grant T32 CA108462 (A.Y.Z.) and the Atwater Foundation (A.G.). The authors thank A. Welm for guidance in the use of patient-derived xenografts, D. Lowe and A. Beardsley for technical guidance and helpful discussions, K.A. Fontaine for helpful discussions and comments on the manuscript, and S. Samson for a helpful consumer advocate perspective and guidance on our work.
Author information
Authors and Affiliations
Contributions
R.C. designed and conducted all of the experiments, except the initial MTB-TOM metabolomic and HMECMYC-ER studies, and wrote the manuscript. A.Y.Z. performed the in vivo studies and provided intellectual input and valuable discussion. R.A.K. performed the mass spectrometry and metabolomic analyses. S.B. performed bioinformatic analyses of the data from the gene expression and metabolomic analyses. C.M. performed the in vitro proliferation and viability studies, did the matrix detachment studies and assisted in TUNEL quantification. B.A. performed the HMECMYC-ER studies and provided valuable discussion. H.E. performed the orthotopic tumor transplants. S.K. supervised the FAO activity analyses. A.T. provided the CPT1B-knockdown data, interpreted data and was provided valuable discussion. G.K. constructed the HCI-002 etomoxir study tissue microarray and performed Ki-67 staining and quantification. D.K.N. supervised the mass spectrometry and metabolomic analyses, and provided valuable discussion. A.G. supervised all of the studies, and provided valuable discussion and intellectual input. All authors edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–9 (PDF 4310 kb)
Supplementary Table 1
Metabolomic changes in MTB-TOM tumors compared to non-tumor mammary glands. (XLSX 120 kb)
Supplementary Table 2
List of genes associated with fatty-acid metabolism by the Gene Ontology Database and their expression in TN compared to RP tumors from TCGA RNASeq data (771 patients). (XLSX 77 kb)
Supplementary Table 3
Correlation analyses of TN and RP tumors to TCGA TN fattyacid metabolism centroid. (XLSX 10 kb)
Supplementary Table 4
Univariate analysis indicating the effect of fatty-acid metabolism gene expression on survival of all, TN and RP patients in a pooled neoadjuvant chemotherapy (taxaneanthracycline) treated cohorts. (XLSX 92 kb)
Supplementary Table 5
Metabolomic changes in HCI-002 untreated tumors compared to 60 mg/kg etomoxir-treated tumors. (XLSX 105 kb)
Supplementary Table 6
Metabolomic changes in MTB-TOM untreated tumors compared to 20 mg/kg etomoxir-treated tumors. (XLSX 70 kb)
Supplementary Data
Supplementary Source Data (PDF 1468 kb)
Rights and permissions
About this article
Cite this article
Camarda, R., Zhou, A., Kohnz, R. et al. Inhibition of fatty acid oxidation as a therapy for MYC-overexpressing triple-negative breast cancer. Nat Med 22, 427–432 (2016). https://doi.org/10.1038/nm.4055
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nm.4055
This article is cited by
-
Recent advances in the potential role of RNA N4-acetylcytidine in cancer progression
Cell Communication and Signaling (2024)
-
Imaging the metabolic reprograming of fatty acid synthesis pathway enables new diagnostic and therapeutic opportunity for breast cancer
Cancer Cell International (2023)
-
Lipid metabolic reprogramming in tumor microenvironment: from mechanisms to therapeutics
Journal of Hematology & Oncology (2023)
-
The impact of lipid metabolism on breast cancer: a review about its role in tumorigenesis and immune escape
Cell Communication and Signaling (2023)
-
Mitoribosomal synthetic lethality overcomes multidrug resistance in MYC-driven neuroblastoma
Cell Death & Disease (2023)