Abstract
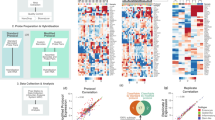
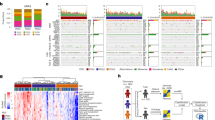
Colorectal cancer (CRC) is a frequently lethal disease with heterogeneous outcomes and drug responses. To resolve inconsistencies among the reported gene expression–based CRC classifications and facilitate clinical translation, we formed an international consortium dedicated to large-scale data sharing and analytics across expert groups. We show marked interconnectivity between six independent classification systems coalescing into four consensus molecular subtypes (CMSs) with distinguishing features: CMS1 (microsatellite instability immune, 14%), hypermutated, microsatellite unstable and strong immune activation; CMS2 (canonical, 37%), epithelial, marked WNT and MYC signaling activation; CMS3 (metabolic, 13%), epithelial and evident metabolic dysregulation; and CMS4 (mesenchymal, 23%), prominent transforming growth factor–β activation, stromal invasion and angiogenesis. Samples with mixed features (13%) possibly represent a transition phenotype or intratumoral heterogeneity. We consider the CMS groups the most robust classification system currently available for CRC—with clear biological interpretability—and the basis for future clinical stratification and subtype-based targeted interventions.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Hoadley, K.A. et al. Multiplatform analysis of 12 cancer types reveals molecular classification within and across tissues of origin. Cell 158, 929–944 (2014).
Cancer Genome Atlas Network. Comprehensive molecular characterization of human colon and rectal cancer. Nature 487, 330–337 (2012).
Roepman, P. et al. Colorectal cancer intrinsic subtypes predict chemotherapy benefit, deficient mismatch repair and epithelial-to-mesenchymal transition. Int. J. Cancer 134, 552–562 (2014).
Budinska, E. et al. Gene expression patterns unveil a new level of molecular heterogeneity in colorectal cancer. J. Pathol. 231, 63–76 (2013).
Schlicker, A. et al. Subtypes of primary colorectal tumors correlate with response to targeted treatment in colorectal cell lines. BMC Med. Genomics 5, 66 (2012).
Sadanandam, A. et al. A colorectal cancer classification system that associates cellular phenotype and responses to therapy. Nat. Med. 19, 619–625 (2013).
De Sousa E Melo, F. et al. Poor-prognosis colon cancer is defined by a molecularly distinct subtype and develops from serrated precursor lesions. Nat. Med. 19, 614–618 (2013).
Marisa, L. et al. Gene expression classification of colon cancer into molecular subtypes: characterization, validation and prognostic value. PLoS Med. 10, e1001453 (2013).
Perez-Villamil, B. et al. Colon cancer molecular subtypes identified by expression profiling and associated to stroma, mucinous type and different clinical behavior. BMC Cancer 12, 260 (2012).
Van Cutsem, E. et al. Randomized phase III trial comparing biweekly infusional fluorouracil/leucovorin alone or with irinotecan in the adjuvant treatment of stage III colon cancer: PETACC-3. J. Clin. Oncol. 27, 3117–3125 (2009).
Van Dongen, S. Graph clustering via a discrete uncoupling process. SIAM J. Matrix Anal. Appl. 30, 121–141 (2008).
Enright, A.J., Van Dongen, S. & Ouzounis, C.A. An efficient algorithm for large-scale detection of protein families. Nucleic Acids Res. 30, 1575–1584 (2002).
Llosa, N.J. et al. The vigorous immune microenvironment of microsatellite instable colon cancer is balanced by multiple counter-inhibitory checkpoints. Cancer Discov. 5, 43–51 (2015).
Brunelli, L., Caiola, E., Marabese, M., Broggini, M. & Pastorelli, R. Capturing the metabolomic diversity of KRAS mutants in non-small-cell lung cancer cells. Oncotarget 5, 4722–4731 (2014).
Son, J. et al. Glutamine supports pancreatic cancer growth through a KRAS-regulated metabolic pathway. Nature 496, 101–105 (2013).
Kamphorst, J.J. et al. Hypoxic and Ras-transformed cells support growth by scavenging unsaturated fatty acids from lysophospholipids. Proc. Natl. Acad. Sci. USA 110, 8882–8887 (2013).
Ying, H. et al. Oncogenic Kras maintains pancreatic tumors through regulation of anabolic glucose metabolism. Cell 149, 656–670 (2012).
Lei, Z. et al. Identification of molecular subtypes of gastric cancer with different responses to PI3-kinase inhibitors and 5-fluorouracil. Gastroenterology 145, 554–565 (2013).
Cancer Genome Atlas Research Network. Comprehensive molecular characterization of gastric adenocarcinoma. Nature 513, 202–209 (2014).
Carter, S.L. et al. Absolute quantification of somatic DNA alterations in human cancer. Nat. Biotechnol. 30, 413–421 (2012).
Zhang, B. et al. Proteogenomic characterization of human colon and rectal cancer. Nature 513, 382–387 (2014).
Li, Y., Choi, P.S., Casey, S.C., Dill, D.L. & Felsher, D.W. MYC through miR-17–92 suppresses specific target genes to maintain survival, autonomous proliferation and a neoplastic state. Cancer Cell 26, 262–272 (2014).
Park, S.-M., Gaur, A.B., Lengyel, E. & Peter, M.E. The miR-200 family determines the epithelial phenotype of cancer cells by targeting the E-cadherin repressors ZEB1 and ZEB2. Genes Dev. 22, 894–907 (2008).
Carmona, F.J. et al. A comprehensive DNA methylation profile of epithelial-to-mesenchymal transition. Cancer Res. 74, 5608–5619 (2014).
Tran, B. et al. Impact of BRAF mutation and microsatellite instability on the pattern of metastatic spread and prognosis in metastatic colorectal cancer. Cancer 117, 4623–4632 (2011).
Gavin, P.G. et al. Mutation profiling and microsatellite instability in stage II and III colon cancer: an assessment of their prognostic and oxaliplatin predictive value. Clin. Cancer Res. 18, 6531–6541 (2012).
Popovici, V. et al. Context-dependent interpretation of the prognostic value of BRAF and KRAS mutations in colorectal cancer. BMC Cancer 13, 439 (2013).
Sinicrope, F.A. et al. Molecular markers identify subtypes of stage III colon cancer associated with patient outcomes. Gastroenterology 148, 88–99 (2015).
Le, D.T. et al. PD-1 blockade in tumors with mismatch-repair deficiency. N. Engl. J. Med. 372, 2509–2520 (2015).
Yoshihara, K. et al. Inferring tumor purity and stromal and immune cell admixture from expression data. Nat. Commun. 4, 2612 (2013).
Derry, J.M.J. et al. Developing predictive molecular maps of human disease through community-based modeling. Nat. Genet. 44, 127–130 (2012).
Johnson, W.E., Li, C. & Rabinovic, A. Adjusting batch effects in microarray expression data using empirical Bayes methods. Biostatistics 8, 118–127 (2007).
McCall, M.N., Bolstad, B.M. & Irizarry, R.A. Frozen robust multiarray analysis (fRMA). Biostatistics 11, 242–253 (2010).
Zilliox, M.J. & Irizarry, R.A. A gene expression bar code for microarray data. Nat. Methods 4, 911–913 (2007).
Tusher, V.G., Tibshirani, R. & Chu, G. Significance analysis of microarrays applied to the ionizing radiation response. Proc. Natl. Acad. Sci. USA 98, 5116–5121 (2001).
Tibshirani, R., Hastie, T., Narasimhan, B. & Chu, G. Diagnosis of multiple cancer types by shrunken centroids of gene expression. Proc. Natl. Acad. Sci. USA 99, 6567–6572 (2002).
Brunet, J.-P., Tamayo, P., Golub, T.R. & Mesirov, J.P. Metagenes and molecular pattern discovery using matrix factorization. Proc. Natl. Acad. Sci. USA 101, 4164–4169 (2004).
Marron, J.S., Todd, M.J. & Ahn, J. Distance-weighted discrimination. J. Am. Stat. Assoc. 102, 1267–1271 (2007).
Gautier, L., Cope, L., Bolstad, B.M. & Irizarry, R.A. affy–analysis of Affymetrix GeneChip data at the probe level. Bioinformatics 20, 307–315 (2004).
Li, B. & Dewey, C.N. RSEM: accurate transcript quantification from RNA-seq data with or without a reference genome. BMC Bioinformatics 12, 323 (2011).
Kauffmann, A., Gentleman, R. & Huber, W. arrayQualityMetrics–a bioconductor package for quality assessment of microarray data. Bioinformatics 25, 415–416 (2009).
Tian, S. et al. A robust genomic signature for the detection of colorectal cancer patients with microsatellite instability phenotype and high mutation frequency. J. Pathol. 228, 586–595 (2012).
Breiman, L. Random forest. Mach. Learn. 45, 5–32 (2001).
Chen, X. & Ishwaran, H. Random forests for genomic data analysis. Genomics 99, 323–329 (2012).
Murray, J.S., Dunson, D.B., Carin, L. & Lucas, J.E. Bayesian Gaussian copula factor models for mixed data. J. Am. Stat. Assoc. 108, 656–665 (2013).
Ghosh, J. & Dunson, D.B. Default prior distributions and efficient posterior computation in Bayesian factor analysis. J. Comput. Graph. Stat. 18, 306–320 (2009).
Tamborero, D. et al. Comprehensive identification of mutational cancer driver genes across 12 tumor types. Sci. Rep. 3, 2650 (2013).
Umar, A. et al. Revised Bethesda guidelines for hereditary nonpolyposis colorectal cancer (Lynch syndrome) and microsatellite instability. J. Natl. Cancer Inst. 96, 261–268 (2004).
Lindor, N.M. et al. Immunohistochemistry versus microsatellite instability testing in phenotyping colorectal tumors. J. Clin. Oncol. 20, 1043–1048 (2002).
Weisenberger, D.J. et al. CpG island methylator phenotype underlies sporadic microsatellite instability and is tightly associated with BRAF mutation in colorectal cancer. Nat. Genet. 38, 787–793 (2006).
Kosinski, C. et al. Gene expression patterns of human colon tops and basal crypts and BMP antagonists as intestinal stem cell niche factors. Proc. Natl. Acad. Sci. USA 104, 15418–15423 (2007).
Van der Flier, L.G. et al. The intestinal Wnt/TCF signature. Gastroenterology 132, 628–632 (2007).
Zeller, K.I., Jegga, A.G., Aronow, B.J., O'Donnell, K.A. & Dang, C.V. An integrated database of genes responsive to the Myc oncogenic transcription factor: identification of direct genomic targets. Genome Biol. 4, R69 (2003).
Loboda, A. et al. EMT is the dominant program in human colon cancer. BMC Med. Genomics 4, 9 (2011).
Merlos-Suárez, A. et al. The intestinal stem cell signature identifies colorectal cancer stem cells and predicts disease relapse. Cell Stem Cell 8, 511–524 (2011).
Mlecnik, B. et al. Biomolecular network reconstruction identifies T cell homing factors associated with survival in colorectal cancer. Gastroenterology 138, 1429–1440 (2010).
Tosolini, M. et al. Clinical impact of different classes of infiltrating T cytotoxic and helper cells (TH1, TH2, Treg, TH17) in patients with colorectal cancer. Cancer Res. 71, 1263–1271 (2011).
Galon, J. et al. Type, density and location of immune cells within human colorectal tumors predict clinical outcome. Science 313, 1960–1964 (2006).
Ascierto, M.L. et al. Molecular signatures mostly associated with NK cells are predictive of relapse-free survival in breast cancer patients. J. Transl. Med. 11, 145 (2013).
Gu-Trantien, C. et al. CD4+ follicular helper T cell infiltration predicts breast cancer survival. J. Clin. Invest. 123, 2873–2892 (2013).
Keerthivasan, S. et al. β-Catenin promotes colitis and colon cancer through imprinting of proinflammatory properties in T cells. Sci. Transl. Med. 6, 225ra28 (2014).
Stockis, J. et al. Comparison of stable human Treg and TH clones by transcriptional profiling. Eur. J. Immunol. 39, 869–882 (2009).
Fridlender, Z.G. et al. Transcriptomic analysis comparing tumor-associated neutrophils with granulocytic myeloid-derived suppressor cells and normal neutrophils. PLoS ONE 7, e31524 (2012).
Efron, B. & Tibshirani, R. On testing the significance of sets of genes. Ann. Appl. Stat. 1, 107–129 (2007).
Wang, L. et al. miR-143 acts as a tumor suppressor by targeting N-RAS and enhances temozolomide-induced apoptosis in glioma. Oncotarget 5, 5416–5427 (2014).
Johnson, S.M. et al. RAS is regulated by the let-7 microRNA family. Cell 120, 635–647 (2005).
Kim, T. et al. p53 regulates epithelial-mesenchymal transition through microRNAs targeting ZEB1 and ZEB2. J. Exp. Med. 208, 875–883 (2011).
Lewis, B.P., Burge, C.B. & Bartel, D.P. Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell 120, 15–20 (2005).
Gentleman, R.C. et al. Bioconductor: open software development for computational biology and bioinformatics. Genome Biol. 5, R80 (2004).
Acknowledgements
The authors would like to acknowledge the goodwill and generosity of the colorectal research community who made this study possible. J.G. and S.H.F. are supported by the Integrative Cancer Biology Program of the National Cancer Institute (grant U54CA149237). R.D. is supported by La Caixa International Program for Cancer Research & Education. L.V. is supported by grants from the Dutch Cancer Society (UVA2011-4969 and UVA2014-7245), Worldwide Cancer Research (14-1164), the Maag Lever Darm Stichting (MLDS) (MLDS-CDG 14-03) and the European Research Council (ERG-StG 638193). J.P.M. is supported by grants from the Dutch Cancer Society (UVA2012-573, UVA2013-6331 and UVA2015-7587) and the MLDS (FP012). S.K. is supported by the US National Institutes of Health (grants R01CA172670, R01CA184843, R01 CA187238 and P30CA016672 (Biostatistic and Bioinformatic Core)). A. Sadanandam and G.N. acknowledge support from the National Health Service. S.T. is supported by the Katholieke Universiteit Leuven GOA/12/2106 grant, the EU FP7 Coltheres grant, the Research Foundation Flanders and the Belgian National Cancer Plan.
Author information
Authors and Affiliations
Contributions
J.G., R.D., J.P.M., A. Sadanandam, L.W., M.D., S.K., L.M., L.V., S.T. and S.H.F. conceived and designed the study. A.d.R., P.R., P.L.-P., I.M.S., E.F., F.D.S.E.M., E.M., D.B., K.H., J.W.G., B.B., D.H., J.T., R.B., J.P.M., A. Sadanandam, L.W., M.D., S.K., L.V., V.B. and S.T. provided study materials. J.G., R.D., P.A., B.B., S.G., E.F., D.B., K.H., D.M., G.C.M. and B.M.B. collected and assembled the data. J.G., R.D., X.W., A.d.R., A. Schlicker, C.S., L.M., G.N., P.A., B.M.B., J.M., T.L., L.V., A. Schlicker, J.S.M., B.P.-V., R.S. and M.D. analysed and interpreted the data. J.G., R.D., X.W., A.d.R., A. Sadanandam, C.S., L.M., J.T., R.S., J.P.M., A. Schlicker, M.D., S.K., L.V. and S.T. wrote the manuscript. All authors contributed to the final approval of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
I.M.S. and P.R. are employees of Agendia. R.B. is a shareholder of Agendia.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–13 (PDF 5271 kb)
Supplementary Table 1
Summary of individual groups subtyping strategy (XLSX 17 kb)
Supplementary Table 2
Summary of clinical, pathological and molecular associations of individual groups' subtypes (XLSX 17 kb)
Supplementary Table 3
Data sets and variables used for correlative analyses (XLSX 19 kb)
Supplementary Table 4
Report of Random Forest CMS classifier during training and validation steps (XLSX 11 kb)
Supplementary Table 5
Clinicopathological and molecular associations of CMS groups (XLSX 24 kb)
Supplementary Table 6
Adjusted P values for enrichment in selected copy number counts across CMS groups (XLSX 15 kb)
Supplementary Table 7
Adjusted P values for enrichment in reverse-phase protein array measurements across CMS groups (XLSX 21 kb)
Supplementary Table 8
Adjusted P values for enrichment in cancer drivers mutations across CMS groups (XLSX 22 kb)
Supplementary Table 9
Adjusted P values for gene set mRNA enrichment analysis (XLSX 20 kb)
Supplementary Table 10
Comparison of TCGA proteomic subtypes and CMS groups (XLSX 11 kb)
Supplementary Table 11
Adjusted P values for gene set protein enrichment analysis (XLSX 18 kb)
Supplementary Table 12
Differential microRNA expression levels across CMS groups (XLSX 53 kb)
Supplementary Table 13
Univariate and multivariate survival models (XLSX 19 kb)
Supplementary Table 14
Major clinicopathological and molecular features of classified and undeterminate samples (XLSX 15 kb)
Supplementary Table 15
Major clinicopathological and molecular features of samples with network labels (consensus samples) versus samples with classifier labels (non-consensus classified samples) for each CMS group (XLSX 18 kb)
Supplementary Table 16
Final performance metrics of CMS classifiers (Random Forest and Single Sample Predictor) applied to consensus samples (XLSX 13 kb)
Rights and permissions
About this article
Cite this article
Guinney, J., Dienstmann, R., Wang, X. et al. The consensus molecular subtypes of colorectal cancer. Nat Med 21, 1350–1356 (2015). https://doi.org/10.1038/nm.3967
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nm.3967
This article is cited by
-
Ensemble methods of rank-based trees for single sample classification with gene expression profiles
Journal of Translational Medicine (2024)
-
A complex of cadherin 17 with desmocollin 1 and p120-catenin regulates colorectal cancer migration and invasion according to the cell phenotype
Journal of Experimental & Clinical Cancer Research (2024)
-
An integrated framework for prognosis prediction and drug response modeling in colorectal liver metastasis drug discovery
Journal of Translational Medicine (2024)
-
Feature-specific quantile normalization and feature-specific mean–variance normalization deliver robust bi-directional classification and feature selection performance between microarray and RNAseq data
BMC Bioinformatics (2024)
-
Towards precision oncology discovery: four less known genes and their unknown interactions as highest-performed biomarkers for colorectal cancer
npj Precision Oncology (2024)