Abstract
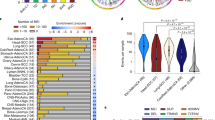
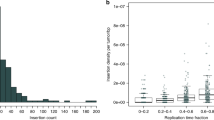
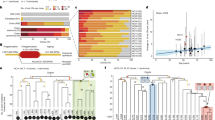
Pancreatic ductal adenocarcinoma (PDAC) is typically diagnosed after the disease has metastasized; it is among the most lethal forms of cancer. We recently described aberrant expression of an open reading frame 1 protein, ORF1p, encoded by long interspersed element-1 (LINE-1; L1) retrotransposon, in PDAC1. To test whether LINE-1 expression leads to somatic insertions of this mobile DNA, we used a targeted method to sequence LINE-1 insertion sites in matched PDAC and normal samples. We found evidence of 465 somatic LINE-1 insertions in 20 PDAC genomes, which were absent from corresponding normal samples. In cases in which matched normal tissue, primary PDAC and metastatic disease sites were available, insertions were found in primary and metastatic tissues in differing proportions. Two adenocarcinomas secondarily involving the pancreas, but originating in the stomach and duodenum, acquired insertions with a similar discordance between primary and metastatic sites. Together, our findings show that LINE-1 contributes to the genetic evolution of PDAC and suggest that somatic insertions are acquired discontinuously in gastrointestinal neoplasms.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Rodic´, N. et al. Long interspersed element-1 protein expression is a hallmark of many human cancers. Am. J. Pathol. 184, 1280–1286 (2014).
Vogelstein, B. et al. Cancer genome landscapes. Science 339, 1546–1558 (2013).
Jones, S. et al. Core signaling pathways in human pancreatic cancers revealed by global genomic analyses. Science 321, 1801–1806 (2008).
Yachida, S. et al. Distant metastasis occurs late during the genetic evolution of pancreatic cancer. Nature 467, 1114–1117 (2010).
Burns, K.H. & Boeke, J.D. Human transposon tectonics. Cell 149, 740–752 (2012).
Beck, C.R., Garcia-Perez, J.L., Badge, R.M. & Moran, J.V. LINE-1 elements in structural variation and disease. Annu. Rev. Genomics Hum. Genet. 12, 187–215 (2011).
Hancks, D.C. & Kazazian, H.H. Jr. Active human retrotransposons: variation and disease. Curr. Opin. Genet. Dev. 22, 191–203 (2012).
Iskow, R.C. et al. Natural mutagenesis of human genomes by endogenous retrotransposons. Cell 141, 1253–1261 (2010).
Ewing, A.D. & Kazazian, H.H. Jr. High-throughput sequencing reveals extensive variation in human-specific L1 content in individual human genomes. Genome Res. 20, 1262–1270 (2010).
Murphy, K.M. et al. Evaluation of candidate genes MAP2K4, MADH4, ACVR1B, and BRCA2 in familial pancreatic cancer: deleterious BRCA2 mutations in 17%. Cancer Res. 62, 3789–3793 (2002).
Jones, S. et al. Exomic sequencing identifies PALB2 as a pancreatic cancer susceptibility gene. Science 324, 217 (2009).
Whitcomb, D.C. et al. Hereditary pancreatitis is caused by a mutation in the cationic trypsinogen gene. Nat. Genet. 14, 141–145 (1996).
Whelan, A.J., Bartsch, D. & Goodfellow, P.J. Brief report: a familial syndrome of pancreatic cancer and melanoma with a mutation in the CDKN2 tumor-suppressor gene. N. Engl. J. Med. 333, 975–977 (1995).
Ostertag, E.M. & Kazazian, H.H. Jr. Biology of mammalian L1 retrotransposons. Annu. Rev. Genet. 35, 501–538 (2001).
Ostertag, E.M. & Kazazian, H.H. Jr. Twin priming: a proposed mechanism for the creation of inversions in L1 retrotransposition. Genome Res. 11, 2059–2065 (2001).
Goodier, J.L., Ostertag, E.M. & Kazazian, H.H. Jr. Transduction of 3′-flanking sequences is common in L1 retrotransposition. Hum. Mol. Genet. 9, 653–657 (2000).
Beck, C.R. et al. LINE-1 retrotransposition activity in human genomes. Cell 141, 1159–1170 (2010).
Solyom, S. et al. Extensive somatic L1 retrotransposition in colorectal tumors. Genome Res. 22, 2328–2338 (2012).
Shukla, R. et al. Endogenous retrotransposition activates oncogenic pathways in hepatocellular carcinoma. Cell 153, 101–111 (2013).
Lee, E. et al. Landscape of somatic retrotransposition in human cancers. Science 337, 967–971 (2012).
Helman, E. et al. Somatic retrotransposition in human cancer revealed by whole-genome and exome sequencing. Genome Res. 24, 1053–1063 (2014).
Tubio, J.M. et al. Mobile DNA in cancer. Extensive transduction of nonrepetitive DNA mediated by L1 retrotransposition in cancer genomes. Science 345, 1251343 (2014).
Stephens, P.J. et al. Massive genomic rearrangement acquired in a single catastrophic event during cancer development. Cell 144, 27–40 (2011).
Evrony, G.D. et al. Single-neuron sequencing analysis of L1 retrotransposition and somatic mutation in the human brain. Cell 151, 483–496 (2012).
Huang, C.R. et al. Mobile interspersed repeats are major structural variants in the human genome. Cell 141, 1171–1182 (2010).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S.L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009).
Ji, H. et al. An integrated software system for analyzing ChIP-chip and ChIP-seq data. Nat. Biotechnol. 26, 1293–1300 (2008).
Amikura, K., Kobari, M. & Matsuno, S. The time of occurrence of liver metastasis in carcinoma of the pancreas. Pancreatol. 17, 139–146 (1995).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. USA 102, 15545–15550 (2005).
Mootha, V.K. et al. PGC-1alpha-responsive genes involved in oxidative phosphorylation are coordinately downregulated in human diabetes. Nat. Genet. 34, 267–273 (2003).
Acknowledgements
This work was started by funding from the Sol Goldman Pancreatic Cancer Research Center (K.H.B. and N.R.) and supported also by the Fred and Janet Sanfilippo Fund in the Department of Pathology at the Johns Hopkins University School of Medicine (N.R.); a Burroughs Wellcome Fund Career Award for Biomedical Scientists Program (K.H.B.); and US National Institutes of Health awards F31CA180682 (A.M.-M.), R01CA163705 (K.H.B.), R01GM103999 (K.H.B.), P50CA62924 (R.H.H. and C.A.I.-D.), R01CA179991 (C.A.I.-D.), as well as the National Institute of General Medical Sciences Center for Systems Biology of Retrotransposition P50GM107632 (K.H.B. and J.D.B.). Computational resources were provided through the National Science Foundation–funded MRI-R2 project #DBI-0959894. The authors would like to thank H. Kazazian, S. Solyom and A. Ewing for their helpful discussion. This work is dedicated to Dr. Frank Kretzer.
Author information
Authors and Affiliations
Contributions
N.R., R.H.H., C.A.I.-D., J.D.B. and K.H.B. conceived of the project; N.R., A.M.-M. and C.A.I.-D. obtained tissues and reviewed histology; J.P.S., A.M., P.S. and P.M. designed and performed molecular-biology assays; N.R., R.S., M.S.T. and N.J.B. performed and reviewed immunostains; Z.A.K., C.R.H. and D.A. designed and performed sequence analysis; N.R., J.P.S., J.D.B. and K.H.B. interpreted data; J.P.S. summarized data for the supplementary tables; N.R. and K.H.B. wrote the manuscript. All authors contributed edits and approved of the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figure 1 (PDF 454 kb)
Supplementary Tables
Supplementary Tables 1–6 (XLSX 185 kb)
Rights and permissions
About this article
Cite this article
Rodić, N., Steranka, J., Makohon-Moore, A. et al. Retrotransposon insertions in the clonal evolution of pancreatic ductal adenocarcinoma. Nat Med 21, 1060–1064 (2015). https://doi.org/10.1038/nm.3919
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nm.3919
This article is cited by
-
Transposable elements as essential elements in the control of gene expression
Mobile DNA (2023)
-
LINE-1 ORF1p as a candidate biomarker in high grade serous ovarian carcinoma
Scientific Reports (2023)
-
Comprehensive proteogenomic characterization of early duodenal cancer reveals the carcinogenesis tracks of different subtypes
Nature Communications (2023)
-
The role of transposable elements in aging and cancer
Biogerontology (2023)
-
LINE-1 promotes tumorigenicity and exacerbates tumor progression via stimulating metabolism reprogramming in non-small cell lung cancer
Molecular Cancer (2022)