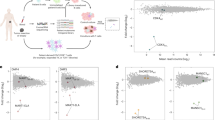
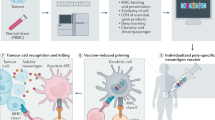
Abstract
Tumor-specific neo-antigens that arise as a consequence of mutations1,2 are thought to be important for the therapeutic efficacy of cancer immunotherapies3,4,5. Accumulating evidence suggests that neo-antigens may be commonly recognized by intratumoral CD8+ T cells3,4,5,6,7, but it is unclear whether neo-antigen–specific CD4+ T cells also frequently reside within human tumors. In view of the accepted role of tumor-specific CD4+ T-cell responses in tumor control8,9,10, we addressed whether neo-antigen–specific CD4+ T-cell reactivity is a common property in human melanoma.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Change history
18 August 2016
In the version of this article initially published, the article did not mention some restrictions on the availability of reagents. The following text has been added to the HTML and PDF versions of the paper: “The retroviral vectors containing BCL-6 and BCL-xL have been generated by a for-profit company, AIMM Therapeutics, which makes the plasmids available. Obtaining the plasmids requires an MTA (http://www.aimmtherapeutics.com/partnering/academic-collaboration/) that includes financial obligations.” The error has been corrected in the HTML and PDF versions of the article.
References
Alexandrov, L.B. et al. Signatures of mutational processes in human cancer. Nature 500, 415–421 (2013).
Vogelstein, B. et al. Cancer genome landscapes. Science 339, 1546–1558 (2013).
Robbins, P.F. et al. Mining exomic sequencing data to identify mutated antigens recognized by adoptively transferred tumor-reactive T cells. Nat. Med. 19, 747–752 (2013).
van Rooij, N. et al. Tumor exome analysis reveals neoantigen-specific T-cell reactivity in an ipilimumab-responsive melanoma. J. Clin. Oncol. 31, e439–e442 (2013).
Lu, Y.C. et al. Mutated PPP1R3B is recognized by T cells used to treat a melanoma patient who experienced a durable complete tumor regression. J. Immunol. 190, 6034–6042 (2013).
Wick, D.A. et al. Surveillance of the tumor mutanome by T cells during progression from primary to recurrent ovarian cancer. Clin. Cancer Res. 20, 1125–1134 (2014).
Rajasagi, M. et al. Systematic identification of personal tumor-specific neoantigens in chronic lymphocytic leukemia. Blood 124, 453–462 (2014).
Kenter, G.G. et al. Vaccination against HPV-16 oncoproteins for vulvar intraepithelial neoplasia. N. Engl. J. Med. 361, 1838–1847 (2009).
Quezada, S.A. et al. Tumor-reactive CD4+ T cells develop cytotoxic activity and eradicate large established melanoma after transfer into lymphopenic hosts. J. Exp. Med. 207, 637–650 (2010).
Ossendorp, F., Mengede, E., Camps, M., Filius, R. & Melief, C.J. Specific T helper cell requirement for optimal induction of cytotoxic T lymphocytes against major histocompatibility complex class II negative tumors. J. Exp. Med. 187, 693–702 (1998).
Friedman, K.M. et al. Tumor-specific CD4+ melanoma tumor-infiltrating lymphocytes. J. Immunother. 35, 400–408 (2012).
Kitano, S. et al. Enhancement of tumor-reactive cytotoxic CD4+ T cell responses after ipilimumab treatment in four advanced melanoma patients. Cancer Immunol. Res. 1, 235–244 (2013).
Tran, E. et al. Cancer immunotherapy based on mutation-specific CD4+ T cells in a patient with epithelial cancer. Science 344, 641–645 (2014).
Champiat, S., Ferte, C., Lebel-Binay, S., Eggermont, A. & Soria, J.C. Exomics and immunogenics: bridging mutational load and immune checkpoints efficacy. OncoImmunology 3, e27817 (2014).
Britten, C.M. et al. The regulatory landscape for actively personalized cancer immunotherapies. Nat. Biotechnol. 31, 880–882 (2013).
Overwijk, W.W., Wang, E., Marincola, F.M., Rammensee, H.G. & Restifo, N.P. Mining the mutanome: developing highly personalized Immunotherapies based on mutational analysis of tumors. J. Immunother. Cancer 1, 11 (2013).
Verdegaal, E.M. et al. Successful treatment of metastatic melanoma by adoptive transfer of blood-derived polyclonal tumor-specific CD4+ and CD8+ T cells in combination with low-dose interferon-alpha. Cancer Immunol. Immunother. 60, 953–963 (2011).
Kwakkenbos, M.J. et al. Genetic manipulation of B cells for the isolation of rare therapeutic antibodies from the human repertoire. Methods 65, 38–43 (2014).
Kwakkenbos, M.J. et al. Generation of stable monoclonal antibody-producing B cell receptor-positive human memory B cells by genetic programming. Nat. Med. 16, 123–128 (2010).
Mason, D. A very high level of crossreactivity is an essential feature of the T-cell receptor. Immunol. Today 19, 395–404 (1998).
Deffrennes, V. et al. Constitutive expression of MHC class II genes in melanoma cell lines results from the transcription of class II transactivator abnormally initiated from its B cell-specific promoter. J. Immunol. 167, 98–106 (2001).
Kvistborg, P. et al. TIL therapy broadens the tumor-reactive CD8(+) T cell compartment in melanoma patients. OncoImmunology 1, 409–418 (2012).
Kvistborg, P. et al. Anti-CTLA-4 therapy broadens the melanoma-reactive CD8+ T cell response. Sci. Transl. Med. 6, 254ra128 (2014).
Lu, Y.C. et al. Efficient identification of mutated cancer antigens recognized by T cells associated with durable tumor regressions. Clin. Cancer Res. 20, 3401–3410 (2014).
Jandus, C. et al. Tumor antigen-specific FOXP3+ CD4 T cells identified in human metastatic melanoma: peptide vaccination results in selective expansion of Th1-like counterparts. Cancer Res. 69, 8085–8093 (2009).
Mautner, J., Jaffee, E.M. & Pardoll, D.M. Tumor-specific CD4+ T cells from a patient with renal cell carcinoma recognize diverse shared antigens. Int. J. Cancer 115, 752–759 (2005).
Wang, R.F., Wang, X., Atwood, A.C., Topalian, S.L. & Rosenberg, S.A. Cloning genes encoding MHC class II-restricted antigens: mutated CDC27 as a tumor antigen. Science 284, 1351–1354 (1999).
Hunder, N.N. et al. Treatment of metastatic melanoma with autologous CD4+ T cells against NY-ESO-1. N. Engl. J. Med. 358, 2698–2703 (2008).
Varela, I. et al. Exome sequencing identifies frequent mutation of the SWI/SNF complex gene PBRM1 in renal carcinoma. Nature 469, 539–542 (2011).
Li, H. & Durbin, R. Fast and accurate long-read alignment with Burrows-Wheeler transform. Bioinformatics 26, 589–595 (2010).
Trapnell, C., Pachter, L. & Salzberg, S.L. TopHat: discovering splice junctions with RNA-Seq. Bioinformatics 25, 1105–1111 (2009).
Trapnell, C. et al. Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat. Biotechnol. 28, 511–515 (2010).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Bolotin, D.A. et al. MiTCR: software for T-cell receptor sequencing data analysis. Nat. Methods 10, 813–814 (2013).
Acknowledgements
We are grateful to A. Pfauth, F. van Diepen and C. Bachas for flow cytometric support; W. van de Kasteele and T. de Jong for technical assistance; N. van Rooij and B. Heemskerk for handling of patient material; and R. Kluin, M. Nieuwland and R. van Kerkhoven for support with next generation sequencing. We thank G. Bendle and P. Kvistborg for critical reading of the manuscript, and members from the Schumacher and Haanen laboratories for useful discussions. This work was supported by Worldwide Cancer Research grant 14-0321 (to C.L. and T.N.M.S.), the PhD Fellowship Program of Boehringer Ingelheim Fonds–Foundation for Basic Research in Biomedicine (to C.L.), Dutch Cancer Society grants: UVA 2010-4822 (to H.S.), NKI 2012-5463 (to J.B.A.G.H., S.H.v.d.B. and T.N.M.S.) and UL 2012-5544 (to E.M.E.V.), The Wellcome Trust Research Training Fellowship for Clinicians (to S.B.), The Wellcome Trust Grant WT098051 (to M.R.S.), the Anticancer Fund (to E.M.E.V., M.V.) the K.G. Jebsen Foundation (to T.N.M.S.), EU FP7 SUPERSIST, and the Stand Up To Cancer–Cancer Research Institute Cancer Immunology Translational Cancer Research Grant (to T.N.M.S.). SU2C is a program of the Entertainment Industry Foundation administered by the American Association for Cancer Research.
Author information
Authors and Affiliations
Contributions
C.L., M.M.v.B. and L.B. designed, performed, analyzed and interpreted experiments and wrote the paper. E.M.E.V. and M.V. designed, performed, analyzed, and interpreted the analysis of subject BO. R.S. generated BCL-6/BCL-XL–immortalized B-cell lines and provided essential culture reagents and advice for the development of the B-cell screening platform. J.J.A.C. developed bioinformatics scripts for TCR sequence identification in NGS data. H.H. and D.e.A. synthesized peptide libraries. S.B., M.R.S. and A.V. performed and analyzed exome and RNA sequencing and developed the pipeline for identification of somatic mutations. J.B.A.G.H. supervised the clinical treatment of NKI TIL patients, supplied patient material and provided clinical interpretation of results. H.S. developed the BCL-6/BCL-XL immortalization technology and provided essential culture reagents and advice for the development of the B-cell screening platform. S.H.v.d.B. supervised, analyzed and interpreted the analysis of subject BO. T.N.M.S. supervised the project, designed and interpreted all experiments, and wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The technology described in this manuscript is the subject of an EU patent application co-owned by NKI-AVL and AIMM Therapeutics B.V. Based on NKI-AVL policy on management of intellectual property, C.L., M.M.v.B., L.B. and T.N.M.S would be entitled to a portion of received royalty income. H.S. and R.S. own certificates of AIMM Therapeutics B.V. (minority stake). T.N.M.S. is a scientific advisor of AIMM Therapeutics B.V. and indirectly owns stock of AIMM Therapeutics B.V. (minority stake).
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7 and Supplementary Table 1. (PDF 1874 kb)
Supplementary Data Set
Source data for all supplementary figures. (XLS 157 kb)
Rights and permissions
About this article
Cite this article
Linnemann, C., van Buuren, M., Bies, L. et al. High-throughput epitope discovery reveals frequent recognition of neo-antigens by CD4+ T cells in human melanoma. Nat Med 21, 81–85 (2015). https://doi.org/10.1038/nm.3773
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nm.3773
This article is cited by
-
CD4+ T cells in cancer
Nature Cancer (2023)
-
Human CD4 cytotoxic T lymphocytes mediate potent tumor control in humanized immune system mice
Communications Biology (2023)
-
A20 promotes colorectal cancer immune evasion by upregulating STC1 expression to block “eat-me” signal
Signal Transduction and Targeted Therapy (2023)
-
The molecular and functional landscape of resistance to immune checkpoint blockade in melanoma
Nature Communications (2023)
-
Microbial peptides activate tumour-infiltrating lymphocytes in glioblastoma
Nature (2023)