Abstract
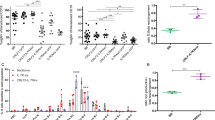
The B cell–specific transcription factor BACH2 is required for affinity maturation of B cells. Here we show that Bach2-mediated activation of p53 is required for stringent elimination of pre-B cells that failed to productively rearrange immunoglobulin VH-DJH gene segments. After productive VH-DJH gene rearrangement, pre-B cell receptor signaling ends BACH2-mediated negative selection through B cell lymphoma 6 (BCL6)-mediated repression of p53. In patients with pre-B acute lymphoblastic leukemia, the BACH2-mediated checkpoint control is compromised by deletions, rare somatic mutations and loss of its upstream activator, PAX5. Low levels of BACH2 expression in these patients represent a strong independent predictor of poor clinical outcome. In this study, we demonstrate that Bach2+/+ pre-B cells resist leukemic transformation by Myc through Bach2-dependent upregulation of p53 and do not initiate fatal leukemia in transplant-recipient mice. Chromatin immunoprecipitation sequencing and gene expression analyses carried out by us revealed that BACH2 competes with BCL6 for promoter binding and reverses BCL6-mediated repression of p53 and other cell cycle checkpoint–control genes. These findings identify BACH2 as a crucial mediator of negative selection at the pre-B cell receptor checkpoint and a safeguard against leukemogenesis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Osmond, D.G. Proliferation kinetics and the lifespan of B cells in central and peripheral lymphoid organs. Curr. Opin. Immunol. 3, 179–185 (1991).
Sakaguchi, N. & Melchers, F. Lambda 5, a new light-chain–related locus selectively expressed in pre-B lymphocytes. Nature 324, 579–582 (1986).
Kitamura, D., Roes, J., Kühn, R. & Rajewsky, K.A. B cell–deficient mouse by targeted disruption of the membrane exon of the immunoglobulin mu chain gene. Nature 350, 423–426 (1991).
Duy, C. et al. BCL6 is critical for the development of a diverse primary B cell repertoire. J. Exp. Med. 207, 1209–1221 (2010).
Nahar, R. et al. Pre-B cell receptor-mediated activation of BCL6 induces pre-B cell quiescence through transcriptional repression of MYC. Blood 118, 4174–4178 (2011).
van Zelm, M.C. et al. Ig gene rearrangement steps are initiated in early human precursor B cell subsets and correlate with specific transcription factor expression. J. Immunol. 175, 5912–5922 (2005).
Fuxa, M. et al. Pax5 induces V-to-DJ rearrangements and locus contraction of the Ig heavy-chain gene. Genes Dev. 18, 411–422 (2004).
Oyake, T. et al. Bach proteins belong to a novel family of BTB-basic leucine zipper transcription factors that interact with MafK and regulate transcription through the NF-E2 site. Mol. Cell Biol. 16, 6083–6095 (1996).
Muto, A. et al. The transcriptional programme of antibody class switching involves the repressor Bach2. Nature 429, 566–571 (2004).
Duy, C. et al. BCL6 enables Ph+ acute lymphoblastic leukaemia cells to survive BCR-ABL1 kinase inhibition. Nature 473, 384–388 (2011).
Rouault, J.P. et al. Identification of BTG2, an antiproliferative p53-dependent component of the DNA damage cellular response pathway. Nat. Genet. 14, 482–486 (1996).
Muljo, S.A. & Schlissel, M.S. A small molecule Abl kinase inhibitor induces differentiation of Abelson virus–transformed pre-B cell lines. Nat. Immunol. 4, 31–37 (2003).
Malin, S. et al. Role of STAT5 in controlling cell survival and immunoglobulin gene recombination during pro-B cell development. Nat. Immunol. 11, 171–179 (2010).
Ehlich, A., Martin, V., Müller, W. & Rajewsky, K. Analysis of the B-cell progenitor compartment at the level of single cells. Curr. Biol. 4, 573–583 (1994).
Mullighan, C.G. et al. Genome-wide analysis of genetic alterations in acute lymphoblastic leukaemia. Nature 446, 758–764 (2007).
Geng, H. et al. Integrative epigenomic analysis identifies biomarkers and therapeutic targets in adult B-acute lymphoblastic leukemia. Cancer Discov. 2, 1004–1023 (2012).
Kawamata, N., Pennella, M.A., Woo, J.L., Berk, A.J. & Koeffler, H.P. Dominant-negative mechanism of leukemogenic PAX5 fusions. Oncogene 31, 966–977 (2012).
Mullighan, C.G. et al. Deletion of IKZF1 and prognosis in acute lymphoblastic leukemia. N. Engl. J. Med. 360, 470–480 (2009).
Merup, M. et al. 6q deletions in acute lymphoblastic leukemia and non-Hodgkin's lymphomas. Blood 91, 3397–3400 (1998).
Hoelbl, A. et al. Stat5 is indispensable for the maintenance of bcr/abl-positive leukaemia. EMBO Mol. Med. 2, 98–110 (2010).
Drayton, S. et al. Tumor suppressor p16INK4a determines sensitivity of human cells to transformation by cooperating cellular oncogenes. Cancer Cell 4, 301–310 (2003).
Schmitt, C.A., McCurrach, M.E., de Stanchina, E., Wallace-Brodeur, R.R. & Lowe, S.W. INK4a/ARF mutations accelerate lymphomagenesis and promote chemoresistance by disabling p53. Genes Dev. 13, 2670–2677 (1999).
Zindy, F. et al. Myc signalling via the ARF tumor suppressor regulates p53-dependent apoptosis and immortalization. Genes Dev. 12, 2424–2433 (1998).
Trageser, D. et al. Pre-B cell receptor-mediated cell cycle arrest in Philadelphia chromosome-positive acute lymphoblastic leukemia requires IKAROS function. J. Exp. Med. 206, 1739–1753 (2009).
Plaisier, S.B., Taschereau, R., Wong, J.A. & Graeber, T.G. Rank-rank hypergeometric overlap: identification of statistically significant overlap between gene-expression signatures. Nucleic Acids Res. 38, e169 (2010).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. USA 102, 15545–15550 (2005).
Acknowledgements
We would like to thank H. Ye (Albert Einstein College of Medicine) for sharing Bcl6−/− mice and wild-type controls and L. Hennighausen (US National Institute of Diabetes and Digestive and Kidney Diseases) for Stat5flox/flox mice. Samples used in this research include those obtained from the Newcastle Haematology Biobank (http://www.ncl.ac.uk/nbb/collections/nhb/index.htm). We thank D.B. Kohn (University of California Los Angeles) for providing us with the retroviral envelope and packaging vectors used in this study. This work is supported by grants from the US National Institutes of Health National Cancer Institute through R01CA137060, R01CA139032, R01CA157644, R01CA172558, R01CA172558 and R01CA169458 (to M.M.), Translational Research Program grants from the Leukemia and Lymphoma Society (grants 6132-09 and 6097-10), a Leukemia and Lymphoma Society Specialized Center of Research (grant 7005-11, B.J. Druker (principal investigator)), the William Lawrence and Blanche Hughes Foundation and a Stand Up To Cancer–American Association for Cancer Research Innovative Research Grant (IRG00909 to M.M.), the California Institute for Regenerative Medicine (CIRM; TR2-01816 to M.M.) and Leukaemia and Lymphoma Research (V.R. and A.G.H.). A.M. and M.M. are Scholars of the Leukemia and Lymphoma Society. V.R. is a Leukaemia and Lymphoma Research Bennett Fellow.
Author information
Authors and Affiliations
Contributions
S.S. and M.M. designed experiments and interpreted data. M.M. and S.S. also conceived the study and wrote the paper. S.S., C. Huang, H.G., Z.C., C.N., B.T., C. Hurtz, M.F.S., D.N., G.B.T. and V.R. performed experiments and interpreted data. H.G., R.H., H.K., V.R., H.P.K., W.L.C., C.L.W., A.G.H. and A.M. provided and characterized patient samples and clinical outcome data. T.G.G., H.P.K., K.I. and A.M. provided important reagents and mouse samples.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Figures and Supplementary Tables
Supplementary Figures 1–22 and Supplementary Tables 1–8 (PDF 4990 kb)
Rights and permissions
About this article
Cite this article
Swaminathan, S., Huang, C., Geng, H. et al. BACH2 mediates negative selection and p53-dependent tumor suppression at the pre-B cell receptor checkpoint. Nat Med 19, 1014–1022 (2013). https://doi.org/10.1038/nm.3247
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nm.3247
This article is cited by
-
BACH2-mediated CD28 and CD40LG axes contribute to pathogenesis and progression of T-cell lymphoblastic leukemia
Cell Death & Disease (2024)
-
Isoform-specific knockdown of long and intermediate prolactin receptors interferes with evolution of B-cell neoplasms
Communications Biology (2023)
-
Disruption to the FOXO-PRDM1 axis resulting from deletions of chromosome 6 in acute lymphoblastic leukaemia
Leukemia (2023)
-
3-Methyladenine but not antioxidants to overcome BACH2-mediated bortezomib resistance in mantle cell lymphoma
Cancer Cell International (2021)
-
Regeneration of immunocompetent B lymphopoiesis from pluripotent stem cells guided by transcription factors
Cellular & Molecular Immunology (2021)