Abstract
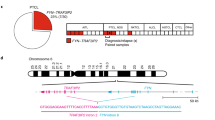
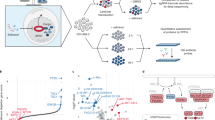
Anaplastic large cell lymphoma (ALCL) is an aggressive non-Hodgkin's lymphoma found in children and young adults. ALCLs frequently carry a chromosomal translocation that results in expression of the oncoprotein nucleophosmin–anaplastic lymphoma kinase (NPM-ALK). The key molecular downstream events required for NPM-ALK–triggered lymphoma growth have been only partly unveiled. Here we show that the activator protein 1 family members JUN and JUNB promote lymphoma development and tumor dissemination through transcriptional regulation of platelet-derived growth factor receptor-β (PDGFRB) in a mouse model of NPM-ALK–triggered lymphomagenesis. Therapeutic inhibition of PDGFRB markedly prolonged survival of NPM-ALK transgenic mice and increased the efficacy of an ALK-specific inhibitor in transplanted NPM-ALK tumors. Notably, inhibition of PDGFRA and PDGFRB in a patient with refractory late-stage NPM-ALK+ ALCL resulted in rapid, complete and sustained remission. Together, our data identify PDGFRB as a previously unknown JUN and JUNB target that could be a highly effective therapy for ALCL.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Kadin, M.E. Ki-1/CD30+ (anaplastic) large-cell lymphoma: maturation of a clinicopathologic entity with prospects of effective therapy. J. Clin. Oncol. 12, 884–887 (1994).
Kadin, M.E. Primary Ki-1-positive anaplastic large-cell lymphoma: a distinct clinicopathologic entity. Ann. Oncol. 5 (suppl. 1), 25–30 (1994).
Stein, H. et al. CD30+ anaplastic large cell lymphoma: a review of its histopathologic, genetic, and clinical features. Blood 96, 3681–3695 (2000).
Morris, S.W. et al. Fusion of a kinase gene, ALK, to a nucleolar protein gene, NPM, in non-Hodgkin's lymphoma. Science 263, 1281–1284 (1994).
Chen, Y. et al. Oncogenic mutations of ALK kinase in neuroblastoma. Nature 455, 971–974 (2008).
George, R.E. et al. Activating mutations in ALK provide a therapeutic target in neuroblastoma. Nature 455, 975–978 (2008).
Janoueix-Lerosey, I. et al. Somatic and germline activating mutations of the ALK kinase receptor in neuroblastoma. Nature 455, 967–970 (2008).
Soda, M. et al. Identification of the transforming EML4-ALK fusion gene in non–small cell lung cancer. Nature 448, 561–566 (2007).
Gambacorti-Passerini, C., Messa, C. & Pogliani, E.M. Crizotinib in anaplastic large-cell lymphoma. N. Engl. J. Med. 364, 775–776 (2011).
Kwak, E.L. et al. Anaplastic lymphoma kinase inhibition in non–small cell lung cancer. N. Engl. J. Med. 363, 1693–1703 (2010).
Choi, Y.L. et al. EML4-ALK mutations in lung cancer that confer resistance to ALK inhibitors. N. Engl. J. Med. 363, 1734–1739 (2010).
Mathas, S. et al. Aberrantly expressed c-JUN and JunB are a hallmark of Hodgkin lymphoma cells, stimulate proliferation and synergize with NF-κB. EMBO J. 21, 4104–4113 (2002).
Staber, P.B. et al. The oncoprotein NPM-ALK of anaplastic large-cell lymphoma induces JUNB transcription via ERK1/2 and JunB translation via mTOR signaling. Blood 110, 3374–3383 (2007).
Chiarle, R. et al. NPM-ALK transgenic mice spontaneously develop T cell lymphomas and plasma cell tumors. Blood 101, 1919–1927 (2003).
Kent, W.J. et al. The human genome browser at UCSC. Genome Res. 12, 996–1006 (2002).
Cartharius, K. et al. MatInspector and beyond: promoter analysis based on transcription factor binding sites. Bioinformatics 21, 2933–2942 (2005).
The ENCODE Project Consortium. A user's guide to the encyclopedia of DNA elements (ENCODE). PLoS Biol. 9, e1001046 (2011).
Andrae, J., Gallini, R. & Betsholtz, C. Role of platelet-derived growth factors in physiology and medicine. Genes Dev. 22, 1276–1312 (2008).
Claesson-Welsh, L. Signal transduction by the PDGF receptors. Prog. Growth Factor Res. 5, 37–54 (1994).
Nissen, L.J. et al. Angiogenic factors FGF2 and PDGF-BB synergistically promote murine tumor neovascularization and metastasis. J. Clin. Invest. 117, 2766–2777 (2007).
Pardanani, A. & Tefferi, A. Imatinib targets other than bcr/abl and their clinical relevance in myeloid disorders. Blood 104, 1931–1939 (2004).
Ergin, M. et al. Inhibition of tyrosine kinase activity induces caspase-dependent apoptosis in anaplastic large cell lymphoma with NPM-ALK (p80) fusion protein. Exp. Hematol. 29, 1082–1090 (2001).
Gunby, R.H. et al. Structural insights into the ATP binding pocket of the anaplastic lymphoma kinase by site-directed mutagenesis, inhibitor binding analysis, and homology modeling. J. Med. Chem. 49, 5759–5768 (2006).
Hantschel, O., Rix, U. & Superti-Furga, G. Target spectrum of the BCR-ABL inhibitors imatinib, nilotinib and dasatinib. Leuk. Lymphoma 49, 615–619 (2008).
Stover, E.H. et al. The small molecule tyrosine kinase inhibitor AMN107 inhibits TEL–PDGFRB and FIP1L1–PDGFR-α in vitro and in vivo. Blood 106, 3206–3213 (2005).
Qiu, L. et al. Autocrine release of interleukin-9 promotes Jak3-dependent survival of ALK+ anaplastic large-cell lymphoma cells. Blood 108, 2407–2415 (2006).
Savan, R. et al. A novel role for IL-22R1 as a driver of inflammation. Blood 117, 575–584 (2011).
Pietras, K. et al. Inhibition of PDGF receptor signaling in tumor stroma enhances antitumor effect of chemotherapy. Cancer Res. 62, 5476–5484 (2002).
Sumida, T. et al. Anti-stromal therapy with imatinib inhibits growth and metastasis of gastric carcinoma in an orthotopic nude mouse model. Int. J. Cancer 128, 2050–2062 (2011).
Cheng, M. et al. CEP-28122, a highly potent and selective orally active inhibitor of anaplastic lymphoma kinase with antitumor activity in experimental models of human cancers. Mol. Cancer Ther. 11, 670–679 (2012).
Iqbal, J. et al. Molecular signatures to improve diagnosis in peripheral T cell lymphoma and prognostication in angioimmunoblastic T cell lymphoma. Blood 115, 1026–1036 (2010).
Piccaluga, P.P. et al. Gene expression analysis of peripheral T cell lymphoma, unspecified, reveals distinct profiles and new potential therapeutic targets. J. Clin. Invest. 117, 823–834 (2007).
Eckerle, S. et al. Gene expression profiling of isolated tumour cells from anaplastic large cell lymphomas: insights into its cellular origin, pathogenesis and relation to Hodgkin lymphoma. Leukemia 23, 2129–2138 (2009).
Piva, R. et al. Gene expression profiling uncovers molecular classifiers for the recognition of anaplastic large-cell lymphoma within peripheral T cell neoplasms. J. Clin. Oncol. 28, 1583–1590 (2010).
Wolfer, A. et al. Inactivation of Notch1 in immature thymocytes does not perturb CD4 or CD8 T cell development. Nat. Immunol. 2, 235–241 (2001).
Behrens, A. et al. Impaired postnatal hepatocyte proliferation and liver regeneration in mice lacking c-jun in the liver. EMBO J. 21, 1782–1790 (2002).
Kenner, L. et al. Mice lacking JunB are osteopenic due to cell-autonomous osteoblast and osteoclast defects. J. Cell Biol. 164, 613–623 (2004).
Ovcharenko, I., Nobrega, M.A., Loots, G.G. & Stubbs, L. ECR Browser: a tool for visualizing and accessing data from comparisons of multiple vertebrate genomes. Nucleic Acids Res. 32, W280–W286 (2004).
Gal-Yam, E.N. et al. Frequent switching of Polycomb repressive marks and DNA hypermethylation in the PC3 prostate cancer cell line. Proc. Natl. Acad. Sci. USA 105, 12979–12984 (2008).
Pflegerl, P. et al. Epidermal loss of JunB leads to a SLE phenotype due to hyper IL-6 signaling. Proc. Natl. Acad. Sci. USA 106, 20423–20428 (2009).
Acknowledgements
We thank E.F. Wagner for providing the JUNBflox/flox and JUNflox/flox mice, A. Ostman for helpful discussions and technical help with PDGFRB analysis, and S. Reichmann for technical help with soluble-CD30 and PDGFB assays. L.K. is supported by the Fonds zur Förderung der wissenschaftlichen Forschung (FWF, P-18478-B12) and the Genome Research in Austria InflammoBiota project. R.M., V.S. and R.E. are supported by the FWF (SFB-F28) and V.S. is supported by FWF project 19723. G.E. is supported by an Elise Richter Fellowship (FWF V102-B12) and an FP7 Marie Curie International Reintegration Grant (IRG 230984). H.D. is supported by the Herzfelder Family Foundation and the Niederösterreichische Forschungs- und Bildungsges.m.b.H. P.W.V. and P.B.S. were supported by Jubiläumsfonds of the Österreichische Nationalbank (P-12147), and P.W.V. was awarded an E. Schroedinger Grant (J 2922). T.W. is supported by the Else-Kröner Fresenius Stiftung. S.P. and G.I. are supported by Italian Association for Cancer Research (AIRC) Special Program in Clinical Molecular Oncology, Milan (5x1000 No. 10007). G.I. is supported by Regione Piemonte (ONCOPROT, CIPE 25/2005); ImmOnc ('Innovative approaches to bust the immune responses', Programma Operativo Regionale, Piattaforme Innovative BIO F.E.S.R. 2007/13, Asse 1 'Ricerca e innovazione' della LR 34/2004) and the Oncology Program of Compagnia di San Paolo, Torino. R.P. is supported by AIRC grant IG-8675. R.E. is supported by the FWF Doktoratskolleg-plus Inflammation and Immunity grant, the Comprehensive Cancer Center Vienna Research Grant and the Austrian Federal Ministry of Science and Research GENAU Austromouse grant. J.M.P. is supported by the Austrian Academy of Sciences and an Advanced Grant from the European Research Council. This study was performed on behalf of the European Research Initiative on ALCL (ERIA).
Author information
Authors and Affiliations
Contributions
H.D., P.W.V., K.K., O.M., P.V., T.W., M. Shehata, V.S., G.H., G.E., J.M.P., U.J., R.M., G.I., R.E. and L.K. designed experiments. D.L., H.D., K.K., P.W.V., M. Schlederer, O.M., A.-I.S., M.R.H., S.H., L.A., C.T., P.B.S., I.S.-K., M.A., S.P., P.P.P., K.M., I.L., S.K., M. Shehata, M.T., S.L., S.D.T., R.P., E.M., G.E., R.M., M.C. and B.A.R. performed experiments and collected and analyzed data. H.D., V.S., R.M., J.M.P., G.I., R.E. and L.K. wrote the manuscript. All authors discussed the results and edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Case Report, Supplementary Methods, Supplementary Figs. 1–8 (PDF 1297 kb)
Rights and permissions
About this article
Cite this article
Laimer, D., Dolznig, H., Kollmann, K. et al. PDGFR blockade is a rational and effective therapy for NPM-ALK–driven lymphomas. Nat Med 18, 1699–1704 (2012). https://doi.org/10.1038/nm.2966
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nm.2966
This article is cited by
-
ALK fusions in the pan-cancer setting: another tumor-agnostic target?
npj Precision Oncology (2023)
-
PDGFRβ promotes oncogenic progression via STAT3/STAT5 hyperactivation in anaplastic large cell lymphoma
Molecular Cancer (2022)
-
AP-1 family transcription factors: a diverse family of proteins that regulate varied cellular activities in classical hodgkin lymphoma and ALK+ ALCL
Experimental Hematology & Oncology (2021)
-
Super-enhancer-based identification of a BATF3/IL-2R−module reveals vulnerabilities in anaplastic large cell lymphoma
Nature Communications (2021)
-
ALK expressed in a gastrointestinal stromal tumor harboring PDGFRA p. D842V mutation:a case report
Diagnostic Pathology (2020)