Abstract
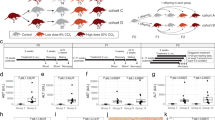
We investigated whether ancestral liver damage leads to heritable reprogramming of hepatic wound healing in male rats. We found that a history of liver damage corresponds with transmission of an epigenetic suppressive adaptation of the fibrogenic component of wound healing to the male F1 and F2 generations. Underlying this adaptation was less generation of liver myofibroblasts, higher hepatic expression of the antifibrogenic factor peroxisome proliferator-activated receptor γ (PPAR-γ) and lower expression of the profibrogenic factor transforming growth factor β1 (TGF-β1) compared to rats without this adaptation. Remodeling of DNA methylation and histone acetylation underpinned these alterations in gene expression. Sperm from rats with liver fibrosis were enriched for the histone variant H2A.Z and trimethylation of histone H3 at Lys27 (H3K27me3) at PPAR-γ chromatin. These modifications to the sperm chromatin were transmittable by adaptive serum transfer from fibrotic rats to naive rats and similar modifications were induced in mesenchymal stem cells exposed to conditioned media from cultured rat or human myofibroblasts. Thus, it is probable that a myofibroblast-secreted soluble factor stimulates heritable epigenetic signatures in sperm so that the resulting offspring better adapt to future fibrogenic hepatic insults. Adding possible relevance to humans, we found that people with mild liver fibrosis have hypomethylation of the PPARG promoter compared to others with severe fibrosis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Change history
05 October 2012
In the version of this article initially published, in Figure 6f the left-hand graph was an inadvertent duplication of the right-hand graph. The error does not alter the overall conclusions of the paper. The figure has been corrected in the HTML and PDF versions of the article.
References
Burd, D.A. et al. Fetal wound healing: an in vitro explant model. J. Pediatr. Surg. 25, 898–901 (1990).
Harrison, M.R. & Adzick, N.S. The fetus as a patient. Surgical considerations. Ann. Surg. 213, 279–291, discussion 277–278 (1991).
Kalluri, R. & Zeisberg, M. Fibroblasts in cancer. Nat. Rev. Cancer 6, 392–401 (2006).
King, T.E. Jr., Pardo, A. & Selman, M. Idiopathic pulmonary fibrosis. Lancet 378, 1949–1961 (2011).
Pinzani, M., Rombouts, K. & Colagrande, S. Fibrosis in chronic liver diseases: diagnosis and management. J. Hepatol. 42 (suppl.), S22–S36 (2005).
Matteoni, C.A. et al. Nonalcoholic fatty liver disease: a spectrum of clinical and pathological severity. Gastroenterology 116, 1413–1419 (1999).
Poynard, T., Bedossa, P. & Opolon, P. Natural history of liver fibrosis progression in patients with chronic hepatitis C. The OBSVIRC, METAVIR, CLINIVIR, and DOSVIRC groups. Lancet 349, 825–832 (1997).
Teli, M.R., Day, C.P., Burt, A.D., Bennett, M.K. & James, O.F. Determinants of progression to cirrhosis or fibrosis in pure alcoholic fatty liver. Lancet 346, 987–990 (1995).
Fischle, W., Wang, Y. & Allis, C.D. Histone and chromatin cross-talk. Curr. Opin. Cell Biol. 15, 172–183 (2003).
Jenuwein, T. & Allis, C.D. Translating the histone code. Science 293, 1074–1080 (2001).
Lavrov, S.A. & Kibanov, M.V. Noncoding RNAs and chromatin structure. Biochemistry (Mosc.) 72, 1422–1438 (2007).
Strahl, B.D. & Allis, C.D. The language of covalent histone modifications. Nature 403, 41–45 (2000).
Umlauf, D., Fraser, P. & Nagano, T. The role of long non-coding RNAs in chromatin structure and gene regulation: variations on a theme. Biol. Chem. 389, 323–331 (2008).
Barker, D.J. The origins of the developmental origins theory. J. Intern. Med. 261, 412–417 (2007).
Recknagel, R.O., Glende, E.A. Jr., Dolak, J.A. & Waller, R.L. Mechanisms of carbon tetrachloride toxicity. Pharmacol. Ther. 43, 139–154 (1989).
Williams, A.T. & Burk, R.F. Carbon tetrachloride hepatotoxicity: an example of free radical–mediated injury. Semin. Liver Dis. 10, 279–284 (1990).
Raucy, J.L., Kraner, J.C. & Lasker, J.M. Bioactivation of halogenated hydrocarbons by cytochrome P4502E1. Crit. Rev. Toxicol. 23, 1–20 (1993).
Friedman, S.L. Hepatic stellate cells: protean, multifunctional, and enigmatic cells of the liver. Physiol. Rev. 88, 125–172 (2008).
Friedman, S.L. Mechanisms of hepatic fibrogenesis. Gastroenterology 134, 1655–1669 (2008).
Yokoi, Y. et al. Immunocytochemical detection of desmin in fat-storing cells (Ito cells). Hepatology 4, 709–714 (1984).
Hazra, S. et al. Peroxisome proliferator-activated receptor gamma induces a phenotypic switch from activated to quiescent hepatic stellate cells. J. Biol. Chem. 279, 11392–11401 (2004).
Mann, J. et al. MeCP2 controls an epigenetic pathway that promotes myofibroblast transdifferentiation and fibrosis. Gastroenterology 138, 705–714 (2010).
Miyahara, T. et al. Peroxisome proliferator-activated receptors and hepatic stellate cell activation. J. Biol. Chem. 275, 35715–35722 (2000).
Gressner, A.M., Weiskirchen, R., Breitkopf, K. & Dooley, S. Roles of TGF-β in hepatic fibrosis. Front. Biosci. 7, d793–d807 (2002).
Leask, A. & Abraham, D.J. TGF-β signaling and the fibrotic response. FASEB J. 18, 816–827 (2004).
Breitling, R., Armengaud, P., Amtmann, A. & Herzyk, P. Rank products: a simple, yet powerful, new method to detect differentially regulated genes in replicated microarray experiments. FEBS Lett. 573, 83–92 (2004).
Hong, F. et al. RankProd: a bioconductor package for detecting differentially expressed genes in meta-analysis. Bioinformatics 22, 2825–2827 (2006).
Eden, S., Hashimshony, T., Keshet, I., Cedar, H. & Thorne, A.W. DNA methylation models histone acetylation. Nature 394, 842 (1998).
Zilberman, D., Coleman-Derr, D., Ballinger, T. & Henikoff, S. Histone H2A.Z and DNA methylation are mutually antagonistic chromatin marks. Nature 456, 125–129 (2008).
Kobor, M.S. & Lorincz, M.C. H2A.Z and DNA methylation: irreconcilable differences. Trends Biochem. Sci. 34, 158–161 (2009).
Creyghton, M.P. et al. H2AZ is enriched at polycomb complex target genes in ES cells and is necessary for lineage commitment. Cell 135, 649–661 (2008).
Gatewood, J.M., Cook, G.R., Balhorn, R., Schmid, C.W. & Bradbury, E.M. Isolation of four core histones from human sperm chromatin representing a minor subset of somatic histones. J. Biol. Chem. 265, 20662–20666 (1990).
Hammoud, S.S. et al. Distinctive chromatin in human sperm packages genes for embryo development. Nature 460, 473–478 (2009).
Chou, S.T. & Gibson, J.B. A histochemical study of the bile ducts in long-term biliary obstruction in the rat. J. Pathol. 103, 163–175 (1971).
De Minicis, S. et al. Gene expression profiles during hepatic stellate cell activation in culture and in vivo. Gastroenterology 132, 1937–1946 (2007).
Anway, M.D., Cupp, A.S., Uzumcu, M. & Skinner, M.K. Epigenetic transgenerational actions of endocrine disruptors and male fertility. Science 308, 1466–1469 (2005).
Ng, S.F. et al. Chronic high-fat diet in fathers programs β-cell dysfunction in female rat offspring. Nature 467, 963–966 (2010).
Carone, B.R. et al. Paternally induced transgenerational environmental reprogramming of metabolic gene expression in mammals. Cell 143, 1084–1096 (2010).
Gama-Sosa, M.A. et al. Tissue-specific differences in DNA methylation in various mammals. Biochim. Biophys. Acta 740, 212–219 (1983).
Ghosh, S. et al. Tissue specific DNA methylation of CpG islands in normal human adult somatic tissues distinguishes neural from non-neural tissues. Epigenetics 5, 527–538 (2010).
Nagase, H. & Ghosh, S. Epigenetics: differential DNA methylation in mammalian somatic tissues. FEBS J. 275, 1617–1623 (2008).
McKay, J.A. et al. Blood as a surrogate marker for tissue-specific DNA methylation and changes due to folate depletion in post-partum female mice. Mol. Nutr. Food Res. 55, 1026–1035 (2011).
Abdalla, H., Yoshizawa, Y. & Hochi, S. Active demethylation of paternal genome in mammalian zygotes. J. Reprod. Dev. 55, 356–360 (2009).
Sasaki, H. & Matsui, Y. Epigenetic events in mammalian germ-cell development: reprogramming and beyond. Nat. Rev. Genet. 9, 129–140 (2008).
Jaenisch, R. DNA methylation and imprinting: why bother? Trends Genet. 13, 323–329 (1997).
Li, E., Beard, C. & Jaenisch, R. Role for DNA methylation in genomic imprinting. Nature 366, 362–365 (1993).
Miller, D., Brinkworth, M. & Iles, D. Paternal DNA packaging in spermatozoa: more than the sum of its parts? DNA, histones, protamines and epigenetics. Reproduction 139, 287–301 (2010).
Johnson, G.D. et al. The sperm nucleus: chromatin, RNA, and the nuclear matrix. Reproduction 141, 21–36 (2011).
Jia, C. Advances in the regulation of liver regeneration. Expert Rev. Gastroenterol. Hepatol. 5, 105–121 (2011).
Oakley, F. et al. Inhibition of inhibitor of κB kinases stimulates hepatic stellate cell apoptosis and accelerated recovery from rat liver fibrosis. Gastroenterology 128, 108–120 (2005).
Oakley, F. et al. Nuclear factor-κB1 (p50) limits the inflammatory and fibrogenic responses to chronic injury. Am. J. Pathol. 166, 695–708 (2005).
Gieling, R.G. et al. The c-Rel subunit of nuclear factor-κB regulates murine liver inflammation, wound-healing, and hepatocyte proliferation. Hepatology 51, 922–931 (2010).
McKay, J.A., Wong, Y.K., Relton, C.L., Ford, D. & Mathers, J.C. Maternal folate supply and sex influence gene-specific DNA methylation in the fetal gut. Mol. Nutr. Food Res. 55, 1717–1723 (2011).
Pittenger, M.F. et al. Multilineage potential of adult human mesenchymal stem cells. Science 284, 143–147 (1999).
Acknowledgements
We thank A. Cheshire for technical assistance and N. Perkins for helpful suggestions with writing the manuscript, M. Bashton for bioinformatic support with microarray analysis and R. Kendall for assistance with rat studies. We also thank J. Kirby and J. Brain for their help with obtaining ethical approval for the retrospective patient study. This work was supported by grants from US National Institutes of Health National Institute of Alcohol Abuse and Alcoholism (to J.M. and D.A.M.) grant numbers 1U01AA018663-01 and R21AA016682; Wellcome Trust grant numbers WT086755MA (to D.A.M.) and WT074472MA (to A.M.E.) and Newcastle Biomedical Research Centre and National Institute for Health Research (to J.M.). The Centre for Brain Ageing and Vitality is funded through the Lifelong Health and Wellbeing cross council initiative by the Medical Research Council, the Biotechnology and Biological Sciences Research Council, the Engineering and Physical Sciences Research Council and the Economic and Social Research Council (to J.C.M.) and the Turkish Association For The Study Of The Liver (to M.Z.).
Author information
Authors and Affiliations
Contributions
M.Z. performed the majority of the laboratory-based experiments and the related data analyses. J.M., T.H., Y.K.W., J.C.M., A.G. and C.L.W. performed a portion of the laboratory experiments and the related data analyses. C.R.F. carried out the surgery and all the in vivo experiments relating to unilateral ureteral obstruction. M.J.B. obtained, cultured and phenotyped human mesenchymal stem cells. S.M. and Q.M.A. obtained archival human liver biopsy tissue and divided patients into groups based on their case history and disease severity. A.D.B. carried out all morphological analyses of tissue sections and scoring of patient liver pathology. A.M.E., F.O. and J.M. carried out all in vivo experiments. J.M. and D.A.M. designed the experiments and wrote the manuscript. All authors discussed the paper and commented on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7 and Supplementary Tables 1 and 2 (PDF 2369 kb)
Rights and permissions
About this article
Cite this article
Zeybel, M., Hardy, T., Wong, Y. et al. Multigenerational epigenetic adaptation of the hepatic wound-healing response. Nat Med 18, 1369–1377 (2012). https://doi.org/10.1038/nm.2893
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nm.2893
This article is cited by
-
Epigenetics as a versatile regulator of fibrosis
Journal of Translational Medicine (2023)
-
Unaltered hepatic wound healing response in male rats with ancestral liver injury
Nature Communications (2023)
-
DNA 5mC and RNA m6A modification successively facilitates the initiation and perpetuation stages of HSC activation in liver fibrosis progression
Cell Death & Differentiation (2023)
-
Sperm chromatin accessibility’s involvement in the intergenerational effects of stress hormone receptor activation
Translational Psychiatry (2023)
-
Missing Causality and Heritability of Autoimmune Hepatitis
Digestive Diseases and Sciences (2023)