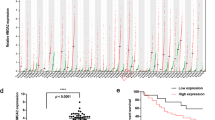
Abstract
Although there is evidence that redox regulation has an essential role in malignancies, its impact on tumor prognosis remains unclear. Here we show crosstalk between oxidative stress and the miR-200 family of microRNAs that affects tumorigenesis and chemosensitivity. miR-141 and miR-200a target p38α and modulate the oxidative stress response. Enhanced expression of these microRNAs mimics p38α deficiency and increases tumor growth in mouse models, but it also improves the response to chemotherapeutic agents. High-grade human ovarian adenocarcinomas that accumulate miR-200a have low concentrations of p38α and an associated oxidative stress signature. The miR200a-dependent stress signature correlates with improved survival of patients in response to treatment. Therefore, the role of miR-200a in stress could be a predictive marker for clinical outcome in ovarian cancer. In addition, although oxidative stress promotes tumor growth, it also sensitizes tumors to treatment, which could account for the limited success of antioxidants in clinical trials.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Hennessy, B.T., Coleman, R.L. & Markman, M. Ovarian cancer. Lancet 374, 1371–1382 (2009).
The Cancer Genome Atlas Research Network. Integrated genomic analyses of ovarian carcinoma. Nature 474, 609–615 (2011).
Allen, R.G. & Tresini, M. Oxidative stress and gene regulation. Free Radic. Biol. Med. 28, 463–499 (2000).
Gerald, D. et al. JunD reduces tumor angiogenesis by protecting cells from oxidative stress. Cell 118, 781–794 (2004).
Pouysségur, J. & Mechta-Grigoriou, F. Redox regulation of the hypoxia-inducible factor. Biol. Chem. 387, 1337–1346 (2006).
Laurent, G. et al. Oxidative stress contributes to aging by enhancing pancreatic angiogenesis and insulin signaling. Cell Metab. 7, 113–124 (2008).
Weinberg, F. & Chandel, N.S. Reactive oxygen species–dependent signaling regulates cancer. Cell. Mol. Life Sci. 66, 3663–3673 (2009).
Pani, G., Galeotti, T. & Chiarugi, P. Metastasis: cancer cell′s escape from oxidative stress. Cancer Metastasis Rev. 29, 351–378 (2010).
Toullec, A. et al. Oxidative stress promotes myofibroblast differentiation and tumour spreading. EMBO Mol. Med. 2, 211–230 (2010).
Dolado, I. et al. p38α MAP kinase as a sensor of reactive oxygen species in tumorigenesis. Cancer Cell 11, 191–205 (2007).
Kennedy, N.J., Cellurale, C. & Davis, R.J. A radical role for p38 MAPK in tumor initiation. Cancer Cell 11, 101–103 (2007).
Hui, L. et al. p38α suppresses normal and cancer cell proliferation by antagonizing the JNK-c-Jun pathway. Nat. Genet. 39, 741–749 (2007).
Wada, T. et al. Antagonistic control of cell fates by JNK and p38-MAPK signaling. Cell Death Differ. 15, 89–93 (2008).
Wagner, E.F. & Nebreda, A.R. Signal integration by JNK and p38 MAPK pathways in cancer development. Nat. Rev. Cancer 9, 537–549 (2009).
Kim, E.K. & Choi, E.J. Pathological roles of MAPK signaling pathways in human diseases. Biochim. Biophys. Acta 1802, 396–405 (2010).
Marsit, C.J., Eddy, K. & Kelsey, K.T. MicroRNA responses to cellular stress. Cancer Res. 66, 10843–10848 (2006).
Ivan, M., Harris, A.L., Martelli, F. & Kulshreshtha, R. Hypoxia response and microRNAs: no longer two separate worlds. J. Cell Mol. Med. 12, 1426–1431 (2008).
Lin, Y. et al. Involvement of microRNAs in hydrogen peroxide–mediated gene regulation and cellular injury response in vascular smooth muscle cells. J. Biol. Chem. 284, 7903–7913 (2009).
Simone, N.L. et al. Ionizing radiation–induced oxidative stress alters miRNA expression. PLoS ONE 4, e6377 (2009).
Wang, Z. et al. Profiles of oxidative stress–related microRNA and mRNA expression in auditory cells. Brain Res. 1346, 14–25 (2010).
Magenta, A. et al. miR-200c is up-regulated by oxidative stress and induces endothelial cell apoptosis and senescence via ZEB1 inhibition. Cell Death Differ. 18, 1628–1639 (2011).
Hurteau, G.J., Carlson, J.A., Spivack, S.D. & Brock, G.J. Overexpression of the microRNA hsa-miR-200c leads to reduced expression of transcription factor 8 and increased expression of E-cadherin. Cancer Res. 67, 7972–7976 (2007).
Bracken, C.P. et al. A double-negative feedback loop between ZEB1–SIP1 and the microRNA-200 family regulates epithelial-mesenchymal transition. Cancer Res. 68, 7846–7854 (2008).
Burk, U. et al. A reciprocal repression between ZEB1 and members of the miR-200 family promotes EMT and invasion in cancer cells. EMBO Rep. 9, 582–589 (2008).
Gregory, P.A. et al. The miR-200 family and miR-205 regulate epithelial to mesenchymal transition by targeting ZEB1 and SIP1. Nat. Cell Biol. 10, 593–601 (2008).
Korpal, M., Lee, E.S., Hu, G. & Kang, Y. The miR-200 family inhibits epithelial-mesenchymal transition and cancer cell migration by direct targeting of E-cadherin transcriptional repressors ZEB1 and ZEB2. J. Biol. Chem. 283, 14910–14914 (2008).
Park, S.M., Gaur, A.B., Lengyel, E. & Peter, M.E. The miR-200 family determines the epithelial phenotype of cancer cells by targeting the E-cadherin repressors ZEB1 and ZEB2. Genes Dev. 22, 894–907 (2008).
Shimono, Y. et al. Downregulation of miRNA-200c links breast cancer stem cells with normal stem cells. Cell 138, 592–603 (2009).
Wellner, U. et al. The EMT-activator ZEB1 promotes tumorigenicity by repressing stemness-inhibiting microRNAs. Nat. Cell Biol. 11, 1487–1495 (2009).
Iliopoulos, D. et al. Loss of miR-200 inhibition of Suz12 leads to polycomb-mediated repression required for the formation and maintenance of cancer stem cells. Mol. Cell 39, 761–772 (2010).
Schickel, R., Park, S.M., Murmann, A.E. & Peter, M.E. miR-200c regulates induction of apoptosis through CD95 by targeting FAP-1. Mol. Cell 38, 908–915 (2010).
Chang, C.J. et al. p53 regulates epithelial-mesenchymal transition and stem cell properties through modulating miRNAs. Nat. Cell Biol. 13, 317–323 (2011).
Kim, T. et al. p53 regulates epithelial-mesenchymal transition through microRNAs targeting ZEB1 and ZEB2. J. Exp. Med. 208, 875–883 (2011).
Ramanathan, B. et al. Resistance to paclitaxel is proportional to cellular total antioxidant capacity. Cancer Res. 65, 8455–8460 (2005).
Alexandre, J., Hu, Y., Lu, W., Pelicano, H. & Huang, P. Novel action of paclitaxel against cancer cells: bystander effect mediated by reactive oxygen species. Cancer Res. 67, 3512–3517 (2007).
Konstantinopoulos, P.A. et al. Carboplatin-induced gene expression changes in vitro are prognostic of survival in epithelial ovarian cancer. BMC Med. Genomics 1, 59 (2008).
Liu, H. et al. mRNA and microRNA expression profiles of the NCI-60 integrated with drug activities. Mol. Cancer Ther. 9, 1080–1091 (2010).
Iorio, M.V. et al. MicroRNA signatures in human ovarian cancer. Cancer Res. 67, 8699–8707 (2007).
Nam, E.J. et al. MicroRNA expression profiles in serous ovarian carcinoma. Clin. Cancer Res. 14, 2690–2695 (2008).
Hu, X. et al. A miR-200 microRNA cluster as prognostic marker in advanced ovarian cancer. Gynecol. Oncol. 114, 457–464 (2009).
Bendoraite, A. et al. Regulation of miR-200 family microRNAs and ZEB transcription factors in ovarian cancer: evidence supporting a mesothelial-to-epithelial transition. Gynecol. Oncol. 116, 117–125 (2010).
Cochrane, D.R., Howe, E.N., Spoelstra, N.S. & Richer, J.K. Loss of miR-200c: a marker of aggressiveness and chemoresistance in female reproductive cancers. J. Oncol. 2010, 821717 (2010).
Mezzanzanica, D., Bagnoli, M., De Cecco, L., Valeri, B. & Canevari, S. Role of microRNAs in ovarian cancer pathogenesis and potential clinical implications. Int. J. Biochem. Cell Biol. 42, 1262–1272 (2010).
van Jaarsveld, M.T., Helleman, J., Berns, E.M. & Wiemer, E.A. MicroRNAs in ovarian cancer biology and therapy resistance. Int. J. Biochem. Cell Biol. 42, 1282–1290 (2010).
Tothill, R.W. et al. Novel molecular subtypes of serous and endometrioid ovarian cancer linked to clinical outcome. Clin. Cancer Res. 14, 5198–5208 (2008).
Yang, H. et al. MicroRNA expression profiling in human ovarian cancer: miR-214 induces cell survival and cisplatin resistance by targeting PTEN. Cancer Res. 68, 425–433 (2008).
Wyman, S.K. et al. Repertoire of microRNAs in epithelial ovarian cancer as determined by next generation sequencing of small RNA cDNA libraries. PLoS ONE 4, e5311 (2009).
Olson, P. et al. MicroRNA dynamics in the stages of tumorigenesis correlate with hallmark capabilities of cancer. Genes Dev. 23, 2152–2165 (2009).
Gibbons, D.L. et al. Contextual extracellular cues promote tumor cell EMT and metastasis by regulating miR-200 family expression. Genes Dev. 23, 2140–2151 (2009).
Wiklund, E.D. et al. Coordinated epigenetic repression of the miR-200 family and miR-205 in invasive bladder cancer. Int. J. Cancer 128, 1327–1334 (2011).
Eitan, R. et al. Tumor microRNA expression patterns associated with resistance to platinum based chemotherapy and survival in ovarian cancer patients. Gynecol. Oncol. 114, 253–259 (2009).
Marchini, S. et al. Association between miR-200c and the survival of patients with stage I epithelial ovarian cancer: a retrospective study of two independent tumour tissue collections. Lancet Oncol. 12, 273–285 (2011).
Karres, J.S., Hilgers, V., Carrera, I., Treisman, J. & Cohen, S.M. The conserved microRNA miR-8 tunes atrophin levels to prevent neurodegeneration in Drosophila. Cell 131, 136–145 (2007).
Hyun, S. et al. Conserved MicroRNA miR-8/miR-200 and its target USH/FOG2 control growth by regulating PI3K. Cell 139, 1096–1108 (2009).
Flynt, A.S. et al. miR-8 microRNAs regulate the response to osmotic stress in zebrafish embryos. J. Cell Biol. 185, 115–127 (2009).
Naidu, S., Vijayan, V., Santoso, S., Kietzmann, T. & Immenschuh, S. Inhibition and genetic deficiency of p38 MAPK up-regulates heme oxygenase-1 gene expression via Nrf2. J. Immunol. 182, 7048–7057 (2009).
Helleman, J., Smid, M., Jansen, M.P., van der Burg, M.E. & Berns, E.M. Pathway analysis of gene lists associated with platinum-based chemotherapy resistance in ovarian cancer: the big picture. Gynecol. Oncol. 117, 170–176 (2010).
Leskelä, S. et al. The miR-200 family controls β-tubulin III expression and is associated with paclitaxel-based treatment response and progression-free survival in ovarian cancer patients. Endocr. Relat. Cancer 18, 85–95 (2011).
Raj, L. et al. Selective killing of cancer cells by a small molecule targeting the stress response to ROS. Nature 475, 231–234 (2011).
Meyniel, J.P. et al. A genomic and transcriptomic approach for a differential diagnosis between primary and secondary ovarian carcinomas in patients with a previous history of breast cancer. BMC Cancer 10, 222 (2010).
Acknowledgements
We thank O. Delattre and S. Chanock for fruitful discussions and comments on the manuscript. We acknowledge S. Alran and B. Baranger (Surgery Department of Institut Curie) and the Biological Resource Center of Institut Curie for providing human ovarian tumors and B. Hasselain and D. Hajage for advice regarding our statistical analyses. We thank the members of the animal facility and the flow-cytometry platform of Institut Curie for their expertise. The experimental work was supported by grants from Institut National de la Santé et de la Recherche Médicale, the Institut Curie, the Ligue Nationale Contre le Cancer, the Institut National du Cancer and the Association pour la Recherche Contre le Cancer. B.M. was supported by a post-doctoral fellowship from the INSERM Avenir program and the Association pour la Recherche Contre le Cancer.
Author information
Authors and Affiliations
Contributions
B.M. and F.M.-G. participated in the conception and design of the experiments. B.M., L.B., M.C., T.G. and Y.d.F. performed the experiments. X.S.-G. selected the human ovarian cancers after adapted characterization, and J.-P.M. provided transcriptome data. P.C. provided the associated clinical data from the subjects. O.M. and A.N. provided technical assistance and expertise in the ovarian tumor sample preparation. J.-P.M., B.M., L.B. and M.C. contributed to the statistical analyses of the data. F.M.-G. wrote the paper with suggestions and comments from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Methods, Supplementary Figures 1–5 and Supplementary Tables 1–6 (PDF 4127 kb)
Rights and permissions
About this article
Cite this article
Mateescu, B., Batista, L., Cardon, M. et al. miR-141 and miR-200a act on ovarian tumorigenesis by controlling oxidative stress response. Nat Med 17, 1627–1635 (2011). https://doi.org/10.1038/nm.2512
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nm.2512
This article is cited by
-
Residual ANTXR1+ myofibroblasts after chemotherapy inhibit anti-tumor immunity via YAP1 signaling pathway
Nature Communications (2024)
-
Cardiovascular complications of diabetes: role of non-coding RNAs in the crosstalk between immune and cardiovascular systems
Cardiovascular Diabetology (2023)
-
Tumor-secreted exosomal miR-141 activates tumor-stroma interactions and controls premetastatic niche formation in ovarian cancer metastasis
Molecular Cancer (2023)
-
Bioinformatic Analysis of miR-200b/429 and Hub Gene Network in Cervical Cancer
Biochemical Genetics (2023)
-
Liquid biopsy in ovarian cancer: advantages and limitations for prognosis and diagnosis
Medical Oncology (2023)