Abstract
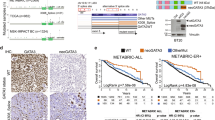
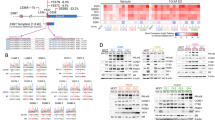
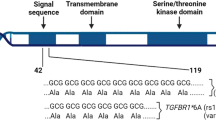
Therapies targeting estrogen receptor α (ERα, encoded by ESR1) have transformed the treatment of breast cancer. However, large numbers of women relapse, highlighting the need for the discovery of new regulatory targets modulating ERα pathways1,2,3,4,5. An siRNA screen identified kinases whose silencing alters the estrogen response including those previously implicated in regulating ERα activity (such as mitogen-activated protein kinase and AKT). Among the most potent regulators was lemur tyrosine kinase-3 (LMTK3), for which a role has not previously been assigned. In contrast to other modulators of ERα activity, LMTK3 seems to have been subject to Darwinian positive selection, a noteworthy result given the unique susceptibility of humans to ERα+ breast cancer. LMTK3 acts by decreasing the activity of protein kinase C (PKC) and the phosphorylation of AKT (Ser473), thereby increasing binding of forkhead box O3 (FOXO3) to the ESR1 promoter. LMTK3 phosphorylated ERα, protecting it from proteasomal degradation in vitro. Silencing of LMTK3 reduced tumor volume in an orthotopic mouse model and abrogated proliferation of ERα+ but not ERα− cells, indicative of its role in ERα activity. In human cancers, LMTK3 abundance and intronic polymorphisms were significantly associated with disease-free and overall survival and predicted response to endocrine therapies. These findings yield insights into the natural history of breast cancer in humans and reveal LMTK3 as a new therapeutic target.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Ali, S. & Coombes, R.C. Endocrine-responsive breast cancer and strategies for combating resistance. Nat. Rev. Cancer 2, 101–112 (2002).
Hammond, M.E. et al. American Society of Clinical Oncology/College Of American Pathologists guideline recommendations for immunohistochemical testing of estrogen and progesterone receptors in breast cancer. J. Clin. Oncol 28, 2784–2795.
Lannigan, D.A. Estrogen receptor phosphorylation. Steroids 68, 1–9 (2003).
Li, C. et al. Essential phosphatases and a phospho-degron are critical for regulation of SRC-3/AIB1 coactivator function and turnover. Mol. Cell 31, 835–849 (2008).
Wu, R.C. et al. Selective phosphorylations of the SRC-3/AIB1 coactivator integrate genomic reponses to multiple cellular signaling pathways. Mol. Cell 15, 937–949 (2004).
Cordera, F. & Jordan, V.C. Steroid receptors and their role in the biology and control of breast cancer growth. Semin. Oncol. 33, 631–641 (2006).
Stack, G. et al. Structure and function of the pS2 gene and estrogen receptor in human breast cancer cells. Cancer Treat. Res. 40, 185–206 (1988).
Campbell, R.A. et al. Phosphatidylinositol 3-kinase/AKT-mediated activation of estrogen receptor alpha: a new model for anti-estrogen resistance. The Journal of Biological Chemistry 276, 9817–9824 (2001).
Chen, D. et al. Phosphorylation of human estrogen receptor alpha at serine 118 by two distinct signal transduction pathways revealed by phosphorylation-specific antisera. Oncogene 21, 4921–4931 (2002).
Jiang, J. et al. Phosphorylation of estrogen receptor-alpha at Ser167 is indicative of longer disease-free and overall survival in breast cancer patients. Clin. Cancer Res. 13, 5769–5776 (2007).
Kato, S. et al. Activation of the estrogen receptor through phosphorylation by mitogen-activated protein kinase. Science 270, 1491–1494 (1995).
Sarwar, N. et al. Phosphorylation of ERalpha at serine 118 in primary breast cancer and in tamoxifen-resistant tumours is indicative of a complex role for ERalpha phosphorylation in breast cancer progression. Endocr. Relat. Cancer 13, 851–861 (2006).
Ghosh, M.G., Thompson, D.A. & Weigel, R.J. PDZK1 and GREB1 are estrogen-regulated genes expressed in hormone-responsive breast cancer. Cancer Res. 60, 6367–6375 (2000).
Beniashvili, D.S. An overview of the world literature on spontaneous tumors in nonhuman primates. J. Med. Primatol. 18, 423–437 (1989).
McClure, H.M. Tumors in nonhuman primates: observations during a six-year period in the Yerkes primate center colony. Am. J. Phys. Anthropol. 38, 425–429 (1973).
Puente, X.S. et al. Comparative analysis of cancer genes in the human and chimpanzee genomes. BMC Genomics 7, 15 (2006).
Seibold, H.R. & Wolf, R.H. Neoplasms and proliferative lesions in 1065 nonhuman primate necropsies. Lab. Anim. Sci 23, 533–539 (1973).
Waters, D.J. et al. Workgroup 4: spontaneous prostate carcinoma in dogs and nonhuman primates. Prostate 36, 64–67 (1998).
Inoue, T. et al. BREK/LMTK2 is a myosin VI-binding protein involved in endosomal membrane trafficking. Genes Cells 13, 483–495 (2008).
Robinson, D.R., Wu, Y.M. & Lin, S.F. The protein tyrosine kinase family of the human genome. Oncogene 19, 5548–5557 (2000).
Tomomura, M. et al. Structural and functional analysis of the apoptosis-associated tyrosine kinase (AATYK) family. Neuroscience 148, 510–521 (2007).
Naik, S., Dothager, R.S., Marasa, J., Lewis, C.L. & Piwnica-Worms, D. Vascular Endothelial Growth Factor Receptor-1 Is Synthetic Lethal to Aberrant β-Catenin Activation in Colon Cancer. Clin. Cancer Res. 15, 7529–7537 (2009).
Tyner, J.W. et al. RNAi screen for rapid therapeutic target identification in leukemia patients. Proc. Natl. Acad. Sci. USA 106, 8695–8700 (2009).
Eeckhoute, J. et al. Positive cross-regulatory loop ties GATA-3 to estrogen receptor alpha expression in breast cancer. Cancer Res. 67, 6477–6483 (2007).
Guo, S. & Sonenshein, G.E. Forkhead box transcription factor FOXO3a regulates estrogen receptor alpha expression and is repressed by the Her-2/neu/phosphatidylinositol 3-kinase/Akt signaling pathway. Mol. Cell. Biol. 24, 8681–8690 (2004).
Morelli, C. et al. Akt2 inhibition enables the forkhead transcription factor FoxO3a to have a repressive role in estrogen receptor alpha transcriptional activity in breast cancer cells. Mol. Cell. Biol. 30, 857–870.
Zou, Y. et al. Forkhead box transcription factor FOXO3a suppresses estrogen-dependent breast cancer cell proliferation and tumorigenesis. Breast Cancer Res 10, R21 (2008).
Madureira, P.A. et al. The Forkhead box M1 protein regulates the transcription of the estrogen receptor alpha in breast cancer cells. J. Biol. Chem. 281, 25167–25176 (2006).
Castles, C.G., Oesterreich, S., Hansen, R. & Fuqua, S.A. Auto-regulation of the estrogen receptor promoter. J. Steroid Biochem. Mol. Biol. 62, 155–163 (1997).
Brunet, A. et al. Akt promotes cell survival by phosphorylating and inhibiting a Forkhead transcription factor. Cell 96, 857–868 (1999).
Marsaud, V., Gougelet, A., Maillard, S. & Renoir, J.M. Various phosphorylation pathways, depending on agonist and antagonist binding to endogenous estrogen receptor alpha (ERalpha), differentially affect ERalpha extractability, proteasome-mediated stability, and transcriptional activity in human breast cancer cells. Mol. Endocrinol. 17, 2013–2027 (2003).
Belguise, K. & Sonenshein, G.E. PKCtheta promotes c-Rel-driven mammary tumorigenesis in mice and humans by repressing estrogen receptor alpha synthesis. J. Clin. Invest. 117, 4009–4021 (2007).
Gschwendt, M. et al. Inhibition of protein kinase C mu by various inhibitors. Differentiation from protein kinase c isoenzymes. FEBS Lett 392, 77–80 (1996).
Castagna, M. et al. Direct activation of calcium-activated, phospholipid-dependent protein kinase by tumor-promoting phorbol esters. J. Biol. Chem. 257, 7847–7851 (1982).
Powe, D.G. et al. Investigating AP-2 and YY1 protein expression as a cause of high HER2 gene transcription in breast cancers with discordant HER2 gene amplification. Breast Cancer Res. 11, R90 (2009).
Brunner, N. et al. MCF7/LCC9: an antiestrogen-resistant MCF-7 variant in which acquired resistance to the steroidal antiestrogen ICI 182,780 confers an early cross-resistance to the nonsteroidal antiestrogen tamoxifen. Cancer Res. 57, 3486–3493 (1997).
Lasfargues, E.Y., Coutinho, W.G. & Redfield, E.S. Isolation of two human tumor epithelial cell lines from solid breast carcinomas. J. Natl. Cancer Inst. 61, 967–978 (1978).
Tolhurst, R.S. et al. Transient over-expression of estrogen receptor-alpha in breast cancer cells promotes cell survival and estrogen-independent growth. Breast Cancer Res. Treat. published online, doi:10.1007/s10549-010-1122-6 (22 August 2010).
Anzick, S.L. et al. AIB1, a steroid receptor coactivator amplified in breast and ovarian cancer. Science 277, 965–968 (1997).
Messier, W. & Stewart, C.B. Episodic adaptive evolution of primate lysozymes. Nature 385, 151–154 (1997).
Acknowledgements
We would like to thank P. Hexley and J. Elliott of the Flow Cytometry Core Facility (Imperial College London) for analyzing FACS data, J. Gronau for helping with confocal microscopy, P. Trivedi for performing the immunohistochemistry staining and T. Rapoz D'Silva for helping in the laboratory. We are grateful to G. Dranitsaris (Princess Margaret Hospital, Toronto) for statistical advice. Li93 software was a kind gift from W. Hsiung-Li (University of Chicago). The single-nucleotide polymorphism analyses were funded in part by US National Institutes of Health grant 5 P30CA14089-27I. S.A. and R.C.C. are supported by grants from Cancer Research UK and the Department of Health–funded Imperial College Cancer Medicine Centre (ECMC) grant. We thank R. and E. Girling and friends for their support.
Author information
Authors and Affiliations
Contributions
G.G. and J.S. conceived of the study, initiated, designed, supervised and conducted most of the experiments and wrote the manuscript. W.M., S.A., H.-J.L., C.T.-S. and R.C.C. contributed to manuscript editing. G.G. and J.J. performed the kinome screening. A.F., B.A.S., A.P., L.C. and H.Z. performed in vitro experiments (including proliferation assays, quantitative RT-PCR and FACS). A.F. and J.S. performed the immunohistochemistry scoring. D.Y., W.Z. and H.J.L. generated the single-nucleotide polymorphism data. W.M. produced all the evolutionary data. A.R.G. and I.O.E. performed the statistical analysis of the clinical data. All authors discussed the results, conceived further experiments, commented on the manuscript and approved the final submitted version.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Methods, Supplementary Figures 1–11 and Supplementary Tables 1–5 (PDF 1033 kb)
Rights and permissions
About this article
Cite this article
Giamas, G., Filipović, A., Jacob, J. et al. Kinome screening for regulators of the estrogen receptor identifies LMTK3 as a new therapeutic target in breast cancer. Nat Med 17, 715–719 (2011). https://doi.org/10.1038/nm.2351
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nm.2351
This article is cited by
-
A revised nomenclature for the lemur family of protein kinases
Communications Biology (2024)
-
Diving into the dark kinome: lessons learned from LMTK3
Cancer Gene Therapy (2022)
-
Computational discovery of novel human LMTK3 inhibitors by high throughput virtual screening using NCI database
Korean Journal of Chemical Engineering (2022)
-
LMTK3 inhibition affects microtubule stability
Molecular Cancer (2021)
-
The Eleanor ncRNAs activate the topological domain of the ESR1 locus to balance against apoptosis
Nature Communications (2019)