Abstract
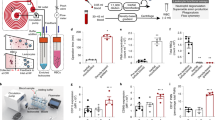
Neutrophils have key roles in modulating the immune response. We present a robust methodology for rapidly isolating neutrophils directly from whole blood with 'on-chip' processing for mRNA and protein isolation for genomics and proteomics. We validate this device with an ex vivo stimulation experiment and by comparison with standard bulk isolation methodologies. Last, we implement this tool as part of a near-patient blood processing system within a multi-center clinical study of the immune response to severe trauma and burn injury. The preliminary results from a small cohort of subjects in our study and healthy controls show a unique time-dependent gene expression pattern clearly demonstrating the ability of this tool to discriminate temporal transcriptional events of neutrophils within a clinical setting.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
References
Nathan, C. Neutrophils and immunity: challenges and opportunities. Nat. Rev. Immunol. 6, 173–182 (2006).
Cassatella, M.A., Gasperini, S. & Russo, M.P. Cytokine expression and release by neutrophils. Ann. NY Acad. Sci. 832, 233–242 (1997).
McDonald, P.P., Bald, A. & Cassatella, M.A. Activation of the NF-κB pathway by inflammatory stimuli in human neutrophils. Blood 89, 3421–3433 (1997).
Burczynski, M.E. & Dorner, A.J. Transcriptional profiling of peripheral blood cells in clinical pharmacogenomic studies. Pharmacogenomics 7, 187–202 (2006).
Calvano, S.E. et al. A network-based analysis of systemic inflammation in humans. Nature 437, 1032–1037 (2005).
Laudanski, K. et al. Cell-specific expression and pathway analyses reveal alterations in trauma-related human T cell and monocyte pathways. Proc. Natl. Acad. Sci. USA 103, 15564–15569 (2006).
Nauseef, W.M. Isolation of human neutrophils from venous blood. Methods Mol. Biol. 412, 15–20 (2007).
Elghetany, M.T. & Davis, B.H. Impact of preanalytical variables on granulocytic surface antigen expression: a review. Cytometry B Clin. Cytom. 65, 1–5 (2005).
Cassatella, M.A. The production of cytokines by polymorphonuclear neutrophils. Immunol. Today 16, 21–26 (1995).
Cheng, X. et al. A microchip approach for practical label-free CD4+ T-cell counting of HIV-infected subjects in resource-poor settings. J. Acquir. Immune Defic. Syndr. 45, 257–261 (2007).
Nagrath, S. et al. Isolation of rare circulating tumour cells in cancer patients by microchip technology. Nature 450, 1235–1239 (2007).
Lyons, P.A. et al. Microarray analysis of human leucocyte subsets: the advantages of positive selection and rapid purification. BMC Genomics 8, 64 (2007).
Schroeder, A. et al. The RIN: an RNA integrity number for assigning integrity values to RNA measurements. BMC Mol. Biol. 7, 3 (2006).
DeForge, L.E., Kenney, J.S., Jones, M.L., Warren, J.S. & Remick, D.G. Biphasic production of IL-8 in lipopolysaccharide (LPS)-stimulated human whole blood. J. Immunol. 148, 2133–2141 (1992).
Pelletier, M. et al. Evidence for a cross-talk between human neutrophils and TH17 cells. Blood 115, 335–343 (2009).
De, A.K. et al. Selective activation of peripheral blood T cell subsets by endotoxin infusion in healthy human subjects corresponds to differential chemokine activation. J. Immunol. 175, 6155–6162 (2005).
Gosselin, E.J., Wardwell, K., Rigby, W.F. & Guyre, P.M. Induction of MHC class II on human polymorphonuclear neutrophils by granulocyte/macrophage colony-stimulating factor, IFN-γ and IL-3. J. Immunol. 151, 1482–1490 (1993).
Cobb, J.P. et al. Application of genome-wide expression analysis to human health and disease. Proc. Natl. Acad. Sci. USA 102, 4801–4806 (2005).
Singh, R. et al. Microarray-based comparison of three amplification methods for nanogram amounts of total RNA. Am. J. Physiol. Cell Physiol. 288, C1179–C1189 (2005).
Kotz, K., Cheng, X. & Toner, M. Cell capture using a microfluidic device. J. Vis. Exp. published online, doi:10.3791/320 (1 October 2007).
Covert, M.W., Leung, T.H., Gaston, J.E. & Baltimore, D. Achieving stability of lipopolysaccharide-induced NF-κB activation. Science 309, 1854–1857 (2005).
Theilgaard-Mönch, K. et al. The transcriptional program of terminal granulocytic differentiation. Blood 105, 1785–1796 (2005).
Kobayashi, S.D., Voyich, J.M., Braughton, K.R. & DeLeo, F.R. Down-regulation of proinflammatory capacity during apoptosis in human polymorphonuclear leukocytes. J. Immunol. 170, 3357–3368 (2003).
Fessler, M.B., Malcolm, K.C., Duncan, M.W. & Worthen, G.S. A genomic and proteomic analysis of activation of the human neutrophil by lipopolysaccharide and its mediation by p38 mitogen-activated protein kinase. J. Biol. Chem. 277, 31291–31302 (2002).
Zhang, X. et al. Gene expression in mature neutrophils: early responses to inflammatory stimuli. J. Leukoc. Biol. 75, 358–372 (2004).
Malcolm, K.C., Arndt, P.G., Manos, E.J., Jones, D.A. & Worthen, G.S. Microarray analysis of lipopolysaccharide-treated human neutrophils. Am. J. Physiol. Lung Cell. Mol. Physiol. 284, L663–L670 (2003).
Kobayashi, S.D., Voyich, J.M., Whitney, A.R. & DeLeo, F.R. Spontaneous neutrophil apoptosis and regulation of cell survival by granulocyte macrophage-colony stimulating factor. J. Leukoc. Biol. 78, 1408–1418 (2005).
Ong, S.E. & Mann, M. Mass spectrometry–based proteomics turns quantitative. Nat. Chem. Biol. 1, 252–262 (2005).
de Godoy, L.M. et al. Comprehensive mass-spectrometry–based proteome quantification of haploid versus diploid yeast. Nature 455, 1251–1254 (2008).
Zhang, X. et al. Proteomic analysis of macrophages stimulated by lipopolysaccharide: Lipopolysaccharide inhibits the cleavage of nucleophosmin. Electrophoresis 27, 1659–1668 (2006).
Berger, T. et al. Lipocalin 2–deficient mice exhibit increased sensitivity to Escherichia coli infection but not to ischemia-reperfusion injury. Proc. Natl. Acad. Sci. USA 103, 1834–1839 (2006).
Meheus, L.A. et al. Identification by microsequencing of lipopolysaccharide-induced proteins secreted by mouse macrophages. J. Immunol. 151, 1535–1547 (1993).
Kotz, K., Cheng, X. & Toner, M. PDMS device fabrication and surface modification. J. Vis. Exp. published online, doi:10.3791/319 (1 October 2007).
Li, C. & Wong, W.H. Model-based analysis of oligonucleotide arrays: expression index computation and outlier detection. Proc. Natl. Acad. Sci. USA 98, 31–36 (2001).
Tusher, V.G., Tibshirani, R. & Chu, G. Significance analysis of microarrays applied to the ionizing radiation response. Proc. Natl. Acad. Sci. USA 98, 5116–5121 (2001).
Zimmer, J.S., Monroe, M.E., Qian, W.J. & Smith, R.D. Advances in proteomics data analysis and display using an accurate mass and time tag approach. Mass Spectrom. Rev. 25, 450–482 (2006).
Qian, W.J., Jacobs, J.M., Liu, T., Camp, D.G. II & Smith, R.D. Advances and challenges in liquid chromatography–mass spectrometry–based proteomics profiling for clinical applications. Mol. Cell. Proteomics 5, 1727–1744 (2006).
Petyuk, V.A. et al. Spatial mapping of protein abundances in the mouse brain by voxelation integrated with high-throughput liquid chromatography–mass spectrometry. Genome Res. 17, 328–336 (2007).
Polpitiya, A.D. et al. DAnTE: a statistical tool for quantitative analysis of -omics data. Bioinformatics 24, 1556–1558 (2008).
Acknowledgements
We thank O. Hurtado, K. Eken, K. Richter and A. Gupta for microfabrication support. We thank C. Vanderburg for help with nucleic acid analysis and the use of the Harvard NeuroDiscovery Center Agilent Bioanalyzer 2100. We thank the University of Florida technical staff (A. Abouhamze, C. Tannahill and R. Ungaro) for managing clinical implementation of the microfluidic devices. K.T.K. was supported by a US National Institutes of Health (NIH) training grant T32 GM-007035-32. These studies were supported by the US NIH Inflammation and the Host Response to Injury Large Scale Collaborative Project, U54 GM-062119, BioMEMS Resource Center P41 EB-002503 and Proteomics Research Resource for Integrative Biology RR018522. The ex vivo stimulation studies and genomics protocol development were partially supported by US National Institutes of Health grants R01-GM-36214 and P01 HG000205, respectively. The proteomics work was performed in the Environmental Molecular Sciences Laboratory, a US Department of Energy Office of Biological and Environmental Research national scientific user facility on the Pacific Northwest National Laboratory (PNNL) campus. PNNL is multiprogram national laboratory operated by Battelle for the Department of Energy under contract number DE-AC05-76RLO 1830.
Author information
Authors and Affiliations
Consortia
Contributions
K.T.K. performed and analyzed experiments. W. Xiao and W.-J.Q. preformed microarray and proteomics analyses. K.T.K., W. Xu, J.W., M.N.M., W. Xu, A.R., E.A.W., L.L.M., D.I., B.H.B., R.W.D. & M.T. designed genomic experiments. K.T.K., W.-J.Q., D.G.C. II and R.D.S. designed proteomic experiments. J.G., S.P.F., A.E.R. and R.G.T. aided in clinical sample studies at Massachusetts General Hospital. K.T.K., C.M.-G., A.D., L.L.M., W. Xiao, M.N.M., J.W., W.-J.Q., B.O.P., D.G.C. II, A.E.R., P.E.B. and M.T. designed, conducted and analyzed the ex vivo stimulation experiment. K.T.K., C.M.-G., W. Xiao, M.N.M. and L.L.M. wrote the manuscript. All authors contributed to the final editing of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–3, Supplementary Tables 1–2,4,6-7,9-12 and Supplementary Methods (PDF 857 kb)
Supplementary Table 3
Significantly perturbed genes and proteins following ex vivo stimulation of whole blood by LPS or GM+I. (PDF 1485 kb)
Supplementary Table 5
Gene expression across all subjects for the genes in Figure 2f comparing microfluidic isolation with Ficoll-dextran isolation (PDF 607 kb)
Supplementary Table 8
Significantly perturbed genes after severe trauma injury (PDF 5588 kb)
Rights and permissions
About this article
Cite this article
Kotz, K., Xiao, W., Miller-Graziano, C. et al. Clinical microfluidics for neutrophil genomics and proteomics. Nat Med 16, 1042–1047 (2010). https://doi.org/10.1038/nm.2205
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nm.2205
This article is cited by
-
Microfluidic devices for neutrophil chemotaxis studies
Journal of Translational Medicine (2020)
-
Chemotaxing neutrophils enter alternate branches at capillary bifurcations
Nature Communications (2020)
-
Discovery and Validation of a Novel Neutrophil Activation Marker Associated with Obesity
Scientific Reports (2019)
-
Assaying leukocyte hallmarks during sepsis
Nature Biomedical Engineering (2019)
-
Whole-blood sorting, enrichment and in situ immunolabeling of cellular subsets using acoustic microstreaming
Microsystems & Nanoengineering (2018)