Abstract
Antigen-specific human monoclonal antibodies (mAbs) are key candidates for therapeutic agents. However, the availability of a suitable screening system for antigen-specific antibody–secreting cells (ASCs) is limited in humans. Here we present a unique method for detecting individual ASCs using microwell array chips, which enables the analysis of live cells on a single-cell basis and offers a rapid, efficient and high-throughput (up to 234,000 individual cells) system for identifying and recovering objective ASCs. We applied the system to detect and retrieve ASCs for hepatitis B virus and influenza viruses from human peripheral blood lymphocytes and produced human mAbs with virus-neutralizing activities within a week. Furthermore, we show that the system is useful for detecting ASCs for multiple antigens as well as for selection of ASCs secreting high-affinity antibodies on a chip. Our method can open the way for the generation of therapeutic antibodies for individual patients.
Similar content being viewed by others
Main
Pandemic outbreaks of infections, such as that of the severe acute respiratory syndrome in 2003 and influenza in 1918, have caused high mortality. Vaccines are effective for preventing infectious diseases. However, it takes a long time to develop and produce them. Antibody therapeutics is another method used for prevention and therapy for newly emerging viruses. A rapid and effective method of selecting and producing objective antibodies is required for preventing a pandemic outbreak of an infection. A principal concern in developing antibody therapeutics is the immunogenicity of mAbs in patients1.
To overcome this problem, various procedures for obtaining fully human antibodies have been developed, including phage display libraries of artificial antibody fragments of human origin2,3,4. Other methods select antibodies that are naturally produced in humans. Attempts have been made to establish hybridomas producing human mAbs5 or Epstein-Barr virus–immortalized human ASCs6,7,8. Furthermore, protocols for isolating antibody heavy and light chain variable region (VH and VL) complementary DNA pairs by RT-PCR directly from single B lineage cells have also been designed9,10,11. Although feasible, these systems are limited by the throughput for the selection of antigen-specific B cells.
Previously, we proposed a cell-based microarray system for developing antigen-specific human mAbs directly from human peripheral blood lymphocytes (PBLs)12,13,14. In this study, we tried to establish a rapid and efficient method for obtaining antigen-specific antibodies using a cell-based microarray system, which detects the secretion of antibodies with desired specificity from single cells in a large population of primary lymphocytes.
Note: Supplementary information is available on the Nature Medicine website.
Results
Detection of antigen-specific ASCs on a chip
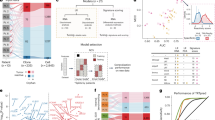
We prepared an array of single live cells by applying ASCs to a microarray chip (Fig. 1a). The chip surface was coated with antibodies against immunoglobulin, and the antibodies secreted by an ASC were trapped on the surface around the well. Binding of the antigen to the specific antibodies formed distinct circular spots, which were easily distinguishable from nonspecific signals. We then retrieved the detected antigen-specific ASCs, recovered antibody cDNA, and produced antibodies within a week15. We named this method 'immunospot array assay on a chip' (ISAAC).
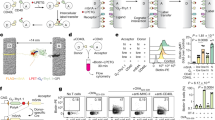
(a) Schematic model depicting the method used for detection and retrieval of single ASCs on a cell-based microwell array chip (ISAAC). Antigen specificity of the produced antibodies could be determined within a week after the detection of antigen-specific ASCs on the chip. InV, influenza virus. (b) Detection of antibody-secreting cell lines with ISAAC. Antibody nonsecreting cells (110TC), HEL-specific ASCs (HyHEL10/110TC) or HBs-antigen-specific ASCs (HB0116/X63) were arrayed on microwell array chips. Signals for antigen-specific antibodies secreted from individual cells were detected (top). Cells were then stained with Oregon Green (center). Antibody signals and cell signals were merged in the bottom panels. (c) Detection of single HEL-specific ASCs in splenocytes from HEL-immunized mice by ISAAC. Antibody secretion was detected with HBs antigen (top left, negative control) or HEL (top center). Then, the cells were stained with Oregon Green, and antibody signals (top panels) and cell signals were merged (bottom). Expanded views of the region marked with yellow boxes are shown (right). (d) Comparison of detection efficiency of the ISAAC and ELISPOT assays for HEL-specific ASCs from HEL-immunized murine spleens. Antigen-specific ASCs were detected with labeled HEL either with ISAAC or by ELISPOT assay. As negative controls, ASCs were analyzed with HBs antigen. The x axis shows the number of analyzed cells, and the y axis shows the number of detected HEL-specific ASCs. Data represent means ± s.d.
First, we tested the feasibility of ISAAC for detecting ASCs on a single-cell basis using the 110TC, HyHEL10/110TC and HB0116/X63 cell lines, which produce no antibodies, antibody to hen egg lysozyme (HEL) and antibody to hepatitis B virus surface antigen (HBs antigen), respectively (Fig. 1b). We cultured the cells on a chip for 3 h and detected the secreted antibodies with biotinylated antigens (HEL or HBs antigen) and Cy3 conjugates of streptavidin (Cy3-streptavidin). We observed signals from the secreted antibodies outside of wells that contained either HyHEL10/110TC or HB0116/X63 cells, but not outside of wells containing 110TC cells (Fig. 1b). Labeled HEL did not detect HBs antigen–specific ASCs, and labeled HBs antigen did not detect HEL-specific ASCs (data not shown). To identify the cells, we stained them on the chips with Oregon Green. Not all of the cells secreted antibodies (more than 15%; Fig. 1b). The results corresponded with those of an ELISPOT assay, which showed that about 10% of the cells secreted antibodies (data not shown). These results indicate that the ISAAC detects ASCs at the single-cell level in culture within only a 3-h period and in an antigen-specific manner, implying the high specificity and speed of the method.
We then verified whether ISAAC can efficiently detect antigen-specific ASCs in lymphoid tissues using HEL-immunized mice. We arrayed CD138+ plasma cells on a chip and detected HEL-specific ASCs using labeled HEL. We observed strong signals of HEL that bound the secreted antibodies on the chip (Fig. 1c and Supplementary Fig. 1). The signals were specific, as we did not observe any spots with labeled HBs antigen as a negative control (Fig. 1c). We did not detect HEL-specific signals when we analyzed CD138+ cells from unimmunized mice (data not shown). When we analyzed approximately 10,000 cells, ISAAC detected an average of 27 spots of HEL-bound IgG (Supplementary Table 1). We simultaneously performed an ELISPOT assay with the same cell preparations. We found that the number of HEL-specific ASCs detected by ISAAC is about four times higher than that detected by ELISPOT assay (Fig. 1d). The results show that ISAAC can detect antigen-specific ASCs in primary lymphocytes at the single-cell level with high efficiency and sensitivity.
Detection of virus-specific ASCs from human PBLs
We then attempted to determine whether ISAAC can be used to obtain antigen-specific human mAbs directly from human peripheral blood B cells. We vaccinated healthy volunteers with HBs vaccine (yeast-derived recombinant HBs antigen) and prepared CD138+ plasma cells from PBLs on day 7 after the boost. The cells were arrayed on a chip coated with antibodies to human IgG. We specifically detected secreted antibodies to HBs antigen with labeled HBs antigen, but not with labeled HEL (Fig. 2a and Supplementary Fig. 2). We detected HBs antigen–specific ASCs at a frequency of 0.17% on day 7 after the boost. Before the boost with HBs antigen, HBs antigen–specific ASCs were barely detectable (data not shown). We retrieved 104 single detected cells and amplified antibody cDNAs. We amplified 68 pairs of antibody VH and VL cDNAs, inserted them into expression vectors and transfected them into Chinese hamster ovary (CHO) cells. ELISA and competitive binding assay showed that 45 of the pairs of antibody cDNAs yielded antibodies specific to HBs antigen (Fig. 2b and Supplementary Table 1).
(a) Detection of HBs antigen-specific ASCs. ASCs were detected using HEL (top left, negative control) or HBs antigen (top middle). The cells were then stained with Oregon Green, and the antibody signals (top) and cell signals were merged (bottom). Expanded views are shown for the region marked with yellow boxes (right). The arrowhead indicates the HBs antigen–specific ASC. (b) Antigen specificity of the antibodies derived from single ASCs using ISAAC, as examined by competitive ELISA using varying amounts of HBs antigen. Data represent means ± s.d. (c) Antigen specificities of the antibodies derived from single ASCs using ISAAC, as examined by competitive ELISA using varying doses of influenza virus type B. Data represent means ± s.d. (d) Correlation of 34 independent HBs antigen–specific antibody clones derived from a volunteer. The antibodies in groups I–IV showed similar amino acid sequences in each group. The antibodies in groups V–VIII showed identical amino acid sequences in each group. The rest of the clones showed unrelated amino acid sequences. (e) Correlation of 19 independent InV-specific antibody clones. The antibodies in groups I–III showed corresponding amino acid sequences in each group. The antibodies in group IV showed identical amino acid sequences. The rest of the clones showed unrelated amino acid sequences.
We also produced influenza virus–specific antibodies directly from CD138+ cells in the human peripheral blood of influenza-vaccinated volunteers. We detected influenza virus–specific ASCs at a frequency of 0.69% on day 10 after the boost. We amplified 46 pairs of antibody VH and VL cDNAs from 63 retrieved cells. ELISA and competitive binding assay showed that 19 antibodies specifically bound the virus (Fig. 2c and Supplementary Table 1). In these experiments, not all of the produced antibodies bound the antigens. This might be due to the retrieval of unnecessary cells together with the objective cell and amplification of cDNAs from nonspecific antibody–producing cells, leading to the production of nonspecific antibodies. Especially in the influenza experiments, we observed nonspecific binding of CD138+ cells on the chip surface, which resulted in our retrieving more than two cells.
Characterization of virus-specific human antibodies
We analyzed the affinities and neutralization activities of HBs antigen–specific or influenza virus–specific antibodies (Tables 1 and 2). The affinities, analyzed with surface plasmon resonance, ranged from 1 × 10−6 to 1 × 10−10 M for HBs antigen–specific antibodies and from 1 × 10−6 to 1 × 10−9 M for influenza hemagglutinin-specific antibodies. When we analyzed inhibitory activities of antibodies for virus infection in vitro, half of the HBs antigen–specific antibodies and all of the hemagglutinin-specific antibodies inhibited the virus infection (Tables 1 and 2). We did not observe substantial correlation between their affinities and inhibitory activities. Notably, we obtained one influenza virus–specific antibody for the extracellular domain of matrix protein 2 (M2e)16,17, which reacted to various types of influenza viruses, including H5N1 (Table 2). It is well known that infection with the hepatitis B virus still threatens health across the world, notwithstanding the development of vaccines, and that a pandemic outbreak of H5N1 influenza virus infection is a major concern. Thus, ISAAC could very efficiently contribute to the supply of virus-neutralizing antibodies by detecting antigen-specific ASCs from primary lymphocytes in human peripheral blood and producing human mAbs very rapidly.
When we analyzed the cDNA sequences of HBs antigen– and influenza virus–specific antibodies that we obtained from ASCs in an individual with ISAAC, we could categorize them into several groups with similar sequences, identical sequences and completely distinct sequences (Fig. 2d,e and Supplementary Fig. 3). These results suggest that ISAAC can identify ASC clones whose antibodies are diversified through antigen-driven differentiation or expansion of memory B cells during immune responses. Another group of researchers has reported the pauciclonal expansion of influenza-specific B cells and repertoire diversification with somatic hypermutation after booster vaccination18. Thus, ISAAC could also be useful for lymphocyte repertoire analysis.
Additional applications of ISAAC
We next examined whether ISAAC could detect ASCs to distinct antigens on a chip by using multiple antigens with different fluorophores. We could detect cells secreting HBs antigen–specific antibodies and those secreting HEL-specific antibodies with FITC conjugated to HBs antigen and biotinylated HEL and Cy3-streptavidin, respectively (Fig. 3a). Recently, the number of fluorophores that can be used in microscopy and cytometry has been increasing, which enabled us to detect ASCs for multiple antigens on a chip.
(a) Detection of ASCs specific to multiple antigens on a chip. A mixture of CHO cells secreting HBs-antigen-specific antibody (HB0116) and those secreting HEL-specific antibody (HyHEL10) were cultured on a chip. HBs-antigen-specific ASCs (white arrows) and HEL-specific ASCs (yellow arrows) were then detected with FITC–HBs antigen (left) and biotinylated HEL–Cy3-streptavidin (center), respectively. FITC and Cy3 signals were merged in the right image. (b,c) Differential detection of HBs antigen–specific ASCs with high affinity. The representative ISAAC signals for secreted IgG and HBs antigen binding to secreted antibodies (HB0116, Kd 3.63 × 10−9 M and HB3-68, Kd 2.93 × 10−6 M) are shown in b. The ratios of Cy3 to FITC intensity (y axis) and Kd (x axis) for each of six CHO cells secreting antibodies with different affinity were plotted in c. More than twenty ISAAC spots with the same affinity were examined. Data represent means ± s.e.m.
Selection of antigen-specific antibodies with high affinity is one of the goals for the screening of mAbs. Thus, we investigated the possibility of selecting high-affinity ASCs with ISAAC (Fig. 3b,c). We individually cultured cells secreting antibodies with various dissociation constants (Kd) on the chips. We then detected the amount of secreted antibodies with FITC-conjugated antibody to human IgG and the amount of HBs antigen that bound the secreted antibodies with biotinylated HBs antigen and Cy3-streptavidin. When we calculated the ratio of the amount of bound antigens to the amount of secreted antibodies, the ratio was high for signals for high-affinity antibodies and low for low-affinity antibodies (Supplementary Methods). These results indicate that ISAAC enables direct screening and production of antigen-specific antibodies with high affinity.
Discussion
ISAAC offers several advantages over conventional ASC-screening methods. First, ISAAC enables direct and efficient identification and segregation of antigen-specific ASCs from a polyclonal mixture of primary lymphocytes. Second, early segregation of cells during screening reduces the time and labor required to maintain many individual clones while characterizing the produced antibodies for appropriate reactivity in the immunochemical assays. Third, the ease of carrying and handling the chip allows for analysis of pathogen-specific ASCs on the spot of an infection outbreak and for a rapid response to the outbreak. Fourth, the 5′-rapid amplification of cDNA ends (5′-RACE) procedure19 is combined with ISAAC to produce an antigen-specific antibody, which enables the augmentation of antibody cDNA without the need to design 5′ VH primers. Fifth, ISAAC enables detection of multiple antigens and selection of ASCs secreting high-affinity antibodies on a chip.
Recently, another group reported a microengraving method for the selection of ASCs using an array of wells 50 μm (or 100 μm) in diameter and depth20. Fundamentally, their method is based on ELISA, whereas our ISAAC is based on ELISPOT. In their procedure, they need to prepare the appropriate cell dilution for analyzing ASCs at single-cell levels. Furthermore, constant secretion of large amounts of antibody might be required to reach a sufficient concentration of antibody for detection because of the large well volume of their system. In these contexts, ISAAC seems to offer the more suitable, rapid and efficient protocol for detecting primary ASCs, although the microengraving protocol also enables their detection.
Regarding the frequencies of influenza-specific ASCs in PBL, another study recently reported about 2,500 influenza-specific IgG-secreting cells in a million PBLs from influenza-vaccinated subjects for the highest responder and about 100 influenza-specific IgG-secreting cells for the lowest responder18, whereas we detected about 70 influenza-specific spots in a million PBLs. A recent study showed an average of 50 influenza-specific IgG-secreting cells in a million PBLs21. Taken together, the frequency of influenza-specific ASCs that we detected with ISAAC was not too low, provided that the subject was a low responder. We believe that ISAAC is suitable for detecting objective cells that exist at a low frequency.
In conclusion, ISAAC is applicable for analyzing and isolating ASCs with the desired specificity directly from human PBLs with very high efficiency and accuracy. ISAAC enables us to obtain antigen-specific mAbs within a week. The generation of human mAbs with high affinity from peripheral blood would be markedly enhanced by our method compared with conventional methods8,10,11. ISAAC would be useful for the rapid generation of neutralizing antibodies in a pandemic outbreak of infectious diseases and would contribute to the antibody therapeutics for individual patients in the future.
Methods
Cells and reagents. We grew 110TC (obtained from H. von Boehmer), HyHEL10/110TC (secreting HEL-specific IgG), and HB0116/X63 (secreting HBs antigen–specific IgG)14 cells (established in our laboratory) in RPMI 1640 supplemented with 10% FCS. We cultured CHO cells (Invitrogen) in FreeStyle CHO Expression Medium (Invitrogen). We used HEL (Sigma), HBs antigen (Kaketsuken), inactivated influenza viruses (see Table 2), M2e peptide (MSLLTEVETPIRNEWGCRCNDSSDK-biotin, Operon Biotechnologies) and polypeptides of the extracellular domain of HBs antigen (amino acids 104–120, amino acids 121–137, amino acids 139–148 and amino acids 149–163, Operon Biotechnologies) as antigens.
Immunization of mice and cell preparation. Mouse experiments were approved by the Committee on Animal Experiments at the University of Toyama. We prepared splenocytes from BALB/c mice (Sankyo Labo) immunized with HEL and then isolated CD138+ cells with mouse CD138-specific mAb–conjugated microbeads using autoMACS Separator (Miltenyi Biotec).
Vaccination of human volunteers and cell preparation. Human experiments were performed with the approval of the Ethical Committee at the University of Toyama. We obtained informed consent from all subjects. For HBs antigen–specific antibodies, we immunized healthy volunteers with recombinant adsorbed hepatitis B vaccine (Bimmugen, Kaketsuken) about a year after the three-injection cycle of primary vaccination was completed. For influenza viruses, we inoculated the subjects with influenza 2007–2008 hemagglutinin vaccine (H1N1 A/Solomon Islands/3/2006, H3N2 A/Hiroshima/52/2005 and B/Malaysia/2506/2004). Seven to ten days after the final boost, we isolated PBLs by the Ficoll–Hypaque method and purified CD138+ cells with human CD138-specific mAb–conjugated microbeads (Miltenyi Biotec).
Microwell array chip fabrication. A microwell array chip was manufactured using micromachining techniques at the Toyama Industrial Technology Center, as described previously14. Microwells 10 μm in diameter and 15 μm deep (for primary lymphocytes), or microwells 15 μm in diameter and 20 μm deep (for X63 or CHO cells) were formed on a silicon surface by deep reactive ion etching22.
ISAAC method. We coated the surface of the chip with goat antibody to mouse or human IgG (10 μg ml−1) and incubated it for 1 h. After we removed the antibody solution, we blocked the chip with 0.01% Biolipidure (NOF Corporation) for 15 min and then washed it with the culture medium. We then arrayed cells in culture medium to the chip and removed the residual cells outside the wells by gentle washing. We cultured the cells on the chip for 3 h at 37 °C. After gentle washing, we reacted 2 μg ml−1 biotinylated antigen for 30 min, followed by Cy3-streptavidin (Sigma) for 30 min. Finally, we stained the cells with 1 μM Oregon Green (Molecular Probes) for 5 min. We observed the antigen-specific antibodies released from single cells under a fluorescence microscope (BX51WI, Olympus) or a cell scanner (CRBIO IIe-FITC, Hitachi Software Engineering). We retrieved single antigen-specific ASCs from individual wells using a micromanipulator (TransferMan NK2, Eppendorf) fitted with capillaries (Primetech) under the fluorescence microscope and then expelled them to microtubes for reverse transcription.
Isolation of antibody cDNA and production of antibody. We amplified antibody cDNAs for VH and VL fragments using a single-cell 5′-RACE method19 with primers for γ chain, κ chain and λ chains and inserted them into expression vectors that contained the whole constant region cDNA for heavy or light chains. Thereafter, we transfected both the heavy and light chain expression vectors together into CHO cells to obtain a supernatant containing whole molecules of antibody, as described previously14. We examined the antigen specificity of the recombinant antibodies by ELISA and confirmed the results with competitive ELISA by adding soluble antigen together with antibodies14. In this study, we screened only IgG-secreting cells with ISAAC and produced IgG.
Enzyme-linked immunosorbent spot assay. We coated 96-well filter plates (MultiScreen, Millipore) with 10 μg ml−1 HEL and blocked them with culture medium. Then, we added mouse CD138+ cells to each well and incubated for 36 h at 37 °C. After washing, we added alkaline phosphatase–conjugated mouse IgG-specific antibody. After washing, we added a mixture of 3-bromo-4-chloro-3-indolyl-phosphate toluidine and p-nitroblue tetrazolium chloride (Sigma) to detect immunospots. As a negative control, we used HBs antigen.
Analysis of virus-neutralizing activity for antibodies. We investigated neutralizing activity for hepatitis B virus using HepaRG cells (supplied from BIOPREDIC)23 and for influenza virus using a previously described protocol with slight modifications24 (Supplementary Methods).
References
Little, M., Kipriyanov, S.M., Le Gall, F. & Moldenhauer, G. Of mice and men: hybridoma and recombinant antibodies. Immunol. Today 21, 364–370 (2000).
Boder, E.T. & Wittrup, K.D. Yeast surface display for screening combinatorial polypeptide libraries. Nat. Biotechnol. 15, 553–557 (1997).
McCafferty, J., Griffiths, A.D., Winter, G. & Chiswell, D.J. Phage antibodies: filamentous phage displaying antibody variable domains. Nature 348, 552–554 (1990).
Winter, G., Griffiths, A.D., Hawkins, R.E. & Hoogenboom, H.R. Making antibodies by phage display technology. Annu. Rev. Immunol. 12, 433–455 (1994).
Gustafsson, B. & Hinkula, J. Antibody production of a human EBV-transformed B cell line and its heterohybridoma and trioma cell line descendants in different culture systems. Hum. Antibodies Hybridomas 5, 98–104 (1994).
Kozbor, D. & Roder, J.C. Requirements for the establishment of high-titered human monoclonal antibodies against tetanus toxoid using the Epstein-Barr virus technique. J. Immunol. 127, 1275–1280 (1981).
Traggiai, E. et al. An efficient method to make human monoclonal antibodies from memory B cells: potent neutralization of SARS coronavirus. Nat. Med. 10, 871–875 (2004).
Winter, G. & Milstein, C. Man-made antibodies. Nature 349, 293–299 (1991).
Lagerkvist, A.C., Furebring, C. & Borrebaeck, C.A. Single, antigen-specific B cells used to generate Fab fragments using CD40-mediated amplification or direct PCR cloning. Biotechniques 18, 862–869 (1995).
Babcook, J.S., Leslie, K.B., Olsen, O.A., Salmon, R.A. & Schrader, J.W. A novel strategy for generating monoclonal antibodies from single, isolated lymphocytes producing antibodies of defined specificities. Proc. Natl. Acad. Sci. USA 93, 7843–7848 (1996).
Meijer, P.J. et al. Isolation of human antibody repertoires with preservation of the natural heavy and light chain pairing. J. Mol. Biol. 358, 764–772 (2006).
Ozawa, T. et al. MAC-CCD system: A novel lymphocyte microwell-array chip system equipped with CCD scanner to generate human monoclonal antibodies against influenza virus. Lab Chip 9, 158–163 (2009).
Tajiri, K. et al. Cell-microarray analysis of antigen-specific B-cells: single cell analysis of antigen receptor expression and specificity. Cytometry A 71, 961–967 (2007).
Tokimitsu, Y. et al. Single lymphocyte analysis with a microwell array chip. Cytometry? 71, 1003–1010 (2007).
Yamamura, S. et al. Single-cell microarray for analyzing cellular response. Anal. Chem. 77, 8050–8056 (2005).
Schnell, J.R. & Chou, J.J. Structure and mechanism of the M2 proton channel of influenza A virus. Nature 451, 591–595 (2008).
Stouffer, A.L. et al. Structural basis for the function and inhibition of an influenza virus proton channel. Nature 451, 596–599 (2008).
Wrammert, J. et al. Rapid cloning of high-affinity human monoclonal antibodies against influenza virus. Nature 453, 667–671 (2008).
Ozawa, T., Kishi, H. & Muraguchi, A. Amplification and analysis of cDNA generated from a single cell by 5′-RACE: application to isolation of antibody heavy and light chain variable gene sequences from single B cells. Biotechniques 40, 469–470, 472, 474 passim (2006).
Love, J.C., Ronan, J.L., Grotenbreg, G.M., van der Veen, A.G. & Ploegh, H.L. A microengraving method for rapid selection of single cells producing antigen-specific antibodies. Nat. Biotechnol. 24, 703–707 (2006).
Sasaki, S. et al. Comparison of the influenza virus-specific effector and memory B-cell responses to immunization of children and adults with live attenuated or inactivated influenza virus vaccines. J. Virol. 81, 215–228 (2007).
Laermer, F. & Schilp, A. Method of anisotropically etching silicon. US patent 5,501,893 (1996).
Gripon, P. et al. Infection of a human hepatoma cell line by hepatitis B virus. Proc. Natl. Acad. Sci. USA 99, 15655–15660 (2002).
Okuno, Y. et al. Rapid focus reduction neutralization test of influenza A and B viruses in microtiter system. J. Clin. Microbiol. 28, 1308–1313 (1990).
Acknowledgements
We thank S. Hirota for technical assistance and K. Hata for secretarial work. We are very grateful to M. Isobe and N. Kurosawa, Laboratory of Molecular and Cellular Biology, Graduate School of Science and Engineering, University of Toyama, for their generous supply of an antibody expression system. We thank H. von Boehmer, Dana-Farber Cancer Institute, Harvard Medical School, for 110TC cells. This research was supported by grants from the Toyama Medical Bio-Cluster Project and a Grant-in-Aid of the Ministry of Education, Culture, Sports, Science and Technology, Japan. A.J., K.K. and S.Kadowaki were supported by a postdoctoral fellowship from the Toyama Medical Bio-Cluster Project.
Author information
Authors and Affiliations
Contributions
A.J. established the ISAAC protocol; T. Ozawa analyzed influenza-specific antibodies; K. Tajiri analyzed HBs antigen–specific antibodies; T. Obata developed microwell array chip for ISAAC; S. Kondo and K.K. constructed antibody expression vectors; S. Kadowaki analyzed cells; K. Takahashi analyzed influenza virus neutralization activity; T.S. supervised the project; and H.K. and A.M. organized and conducted the project.
Corresponding author
Ethics declarations
Competing interests
The immunospot array assay on a chip (ISAAC) procedure has been registered for Japan Patent Office patent JP 4148367.
Supplementary information
Supplementary Text and Figures
Supplementary Figs. 1–3, Supplementary Table 1 and Supplementary Methods (PDF 1036 kb)
Rights and permissions
About this article
Cite this article
Jin, A., Ozawa, T., Tajiri, K. et al. A rapid and efficient single-cell manipulation method for screening antigen-specific antibody–secreting cells from human peripheral blood. Nat Med 15, 1088–1092 (2009). https://doi.org/10.1038/nm.1966
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nm.1966
This article is cited by
-
GATS tag system is compatible with biotin labelling methods for protein analysis
Scientific Reports (2023)
-
Identification of druggable regulators of cell secretion via a kinome-wide screen and high-throughput immunomagnetic cell sorting
Nature Biomedical Engineering (2023)
-
Response under pressure: deploying emerging technologies to understand B-cell-mediated immunity in COVID-19
Nature Methods (2022)
-
Rapid cloning of antigen-specific T-cell receptors by leveraging the cis activation of T cells
Nature Biomedical Engineering (2022)
-
Single-cell sorting based on secreted products for functionally defined cell therapies
Microsystems & Nanoengineering (2022)