Abstract
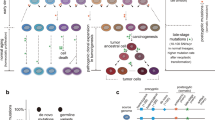
Copy number variants are a recently discovered source of large-scale genomic diversity present in all individuals. We capitalize on these inherent genomic differences, focusing on deletion polymorphisms, to develop informative fluorescence in situ hybridization probes with the ability to unequivocally distinguish between donor and recipient cells in situ. These probes are accurate, specific, highly polymorphic and, notably, can be used to assign genetic identity in situ in a completely gender-independent fashion. We anticipate that these polymorphic deletion probes will be useful in further understanding the dynamics of cellular chimerism after transplantation, including the details of chronic organ rejection, post-transplant lymphoproliferative disorder and graft-versus-host disease, and in optimizing future tissue engineering and pluripotent stem cell therapies.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Krause, D.S. et al. Multi-organ, multi-lineage engraftment by a single bone marrow–derived stem cell. Cell 105, 369–377 (2001).
Tran, S.D. et al. Differentiation of human bone marrow–derived cells into buccal epithelial cells in vivo: a molecular analytical study. Lancet 361, 1084–1088 (2003).
Korbling, M. et al. Hepatocytes and epithelial cells of donor origin in recipients of peripheral-blood stem cells. N. Engl. J. Med. 346, 738–746 (2002).
Jiang, S. et al. Transplanted human bone marrow contributes to vascular endothelium. Proc. Natl. Acad. Sci. USA 101, 16891–16896 (2004).
Minami, E., Laflamme, M.A., Saffitz, J.E. & Murry, C.E. Extracardiac progenitor cells repopulate most major cell types in the transplanted human heart. Circulation 112, 2951–2958 (2005).
Grimm, P.C. et al. Neointimal and tubulointerstitial infiltration by recipient mesenchymal cells in chronic renal-allograft rejection. N. Engl. J. Med. 345, 93–97 (2001).
Iafrate, A.J. et al. Detection of large-scale variation in the human genome. Nat. Genet. 36, 949–951 (2004).
Sebat, J. et al. Large-scale copy number polymorphism in the human genome. Science 305, 525–528 (2004).
Conrad, D.F., Andrews, T.D., Carter, N.P., Hurles, M.E. & Pritchard, J.K. A high-resolution survey of deletion polymorphism in the human genome. Nat. Genet. 38, 75–81 (2006).
Hinds, D.A., Kloek, A.P., Jen, M., Chen, X. & Frazer, K.A. Common deletions and SNPs are in linkage disequilibrium in the human genome. Nat. Genet. 38, 82–85 (2006).
McCarroll, S.A. et al. Common deletion polymorphisms in the human genome. Nat. Genet. 38, 86–92 (2006).
Oliver, D.H., Thompson, R.E., Griffin, C.A. & Eshleman, J.R. Use of single nucleotide polymorphisms (SNP) and real-time polymerase chain reaction for bone marrow engraftment analysis. J. Mol. Diagn. 2, 202–208 (2000).
Mullally, A. & Ritz, J. Beyond HLA: the significance of genomic variation for allogeneic hematopoietic stem cell transplantation. Blood 109, 1355–1362 (2007).
Hochberg, E.P. et al. A novel rapid single nucleotide polymorphism (SNP)-based method for assessment of hematopoietic chimerism after allogeneic stem cell transplantation. Blood 101, 363–369 (2003).
Weber, J.L. & May, P.E. Abundant class of human DNA polymorphisms which can be typed using the polymerase chain reaction. Am. J. Hum. Genet. 44, 388–396 (1989).
Jeffreys, A.J. Genetic fingerprinting. Nat. Med. 11, 1035–1039 (2005).
Litt, M. & Luty, J.A. A hypervariable microsatellite revealed by in vitro amplification of a dinucleotide repeat within the cardiac muscle actin gene. Am. J. Hum. Genet. 44, 397–401 (1989).
Sermon, K., Van Steirteghem, A. & Liebaers, I. Preimplantation genetic diagnosis. Lancet 363, 1633–1641 (2004).
Buckle, V.J. & Kearney, L. New methods in cytogenetics. Curr. Opin. Genet. Dev. 4, 374–382 (1994).
Laflamme, M.A. & Murry, C.E. Regenerating the heart. Nat. Biotechnol. 23, 845–856 (2005).
Rando, T.A. Stem cells, ageing and the quest for immortality. Nature 441, 1080–1086 (2006).
Lindvall, O. & Kokaia, Z. Stem cells for the treatment of neurological disorders. Nature 441, 1094–1096 (2006).
Koike, N. et al. Tissue engineering: creation of long-lasting blood vessels. Nature 428, 138–139 (2004).
Braak, H. & Del Tredici, K. Assessing fetal nerve cell grafts in Parkinson's disease. Nat. Med. 14, 483–485 (2008).
Acknowledgements
We thank M. Han, J. Miller-Batten and J. Reid for excellent technical assistance. We thank C. Lee (Brigham and Women's Hospital) and S. Kantarci (Beth Israel Deaconess Medical Center) for helpful discussions and reagents. We thank S. Ogino for helpful discussions. This work was supported in part by intramural funding from the Department of Pathology, Massachusetts General Hospital to A.J.I.
Author information
Authors and Affiliations
Contributions
D.W. and A.J.I. conceived of the project idea, designed and performed experiments, and wrote the manuscript. L.M.S. designed and performed experiments. Q.V., A.N., H.S. and G.K. performed FISH assays. J.R.S. contributed the autopsy cardiac transplant case and provided helpful discussions. All contributed to the editing of the manuscript.
Corresponding author
Ethics declarations
Competing interests
A provisional application by D.W. and A.J.I. for a patent with the US Patent Office for the use of polymorphic deletion probes is pending and has not yet been licensed.
Supplementary information
Supplementary Text and Figures
Supplementary Methods, Supplementary Figs. 1–4 and Supplementary Table 1 (PDF 403 kb)
Rights and permissions
About this article
Cite this article
Wu, D., Vu, Q., Nguyen, A. et al. In situ genetic analysis of cellular chimerism. Nat Med 15, 215–219 (2009). https://doi.org/10.1038/nm.1862
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nm.1862