Abstract
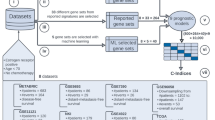
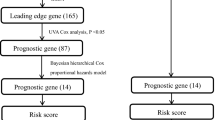
Although prognostic gene expression signatures for survival in early-stage lung cancer have been proposed, for clinical application, it is critical to establish their performance across different subject populations and in different laboratories. Here we report a large, training–testing, multi-site, blinded validation study to characterize the performance of several prognostic models based on gene expression for 442 lung adenocarcinomas. The hypotheses proposed examined whether microarray measurements of gene expression either alone or combined with basic clinical covariates (stage, age, sex) could be used to predict overall survival in lung cancer subjects. Several models examined produced risk scores that substantially correlated with actual subject outcome. Most methods performed better with clinical data, supporting the combined use of clinical and molecular information when building prognostic models for early-stage lung cancer. This study also provides the largest available set of microarray data with extensive pathological and clinical annotation for lung adenocarcinomas.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Jemal, A. et al. Cancer Statistics 2006. CA Cancer J. Clin. 56, 106–130 (2006).
Booth, C.M. & Shepherd, F.A. Adjuvant chemotherapy for resected non-small cell lung cancer. J. Thorac. Oncol. 2, 180–187 (2006).
Gandara, D.R., Wakelee, H., Calhoun, R. & Jablons, D. Adjuvant chemotherapy of stage I non-small cell lung cancer in North America. J. Thorac. Oncol. 7 (suppl. 3), S125–S127 (2007).
Shepherd, F.A. et al. Erlotinib in previously treated non-small-cell lung cancer. N. Engl. J. Med. 353, 123–132 (2005).
Bhattacharjee, A. et al. Classification of human lung carcinomas by mRNA expression profiling reveals distinct adenocarcinoma subclasses. Proc. Natl. Acad. Sci. USA 98, 13790–13795 (2001).
Garber, M.E. et al. Diversity of gene expression in adenocarcinoma of the lung. Proc. Natl. Acad. Sci. USA 98, 13784–13789 (2001).
Beer, D.G. et al. Gene-expression profiles predict survival of subjects with lung adenocarcinoma. Nat. Med. 8, 816–824 (2002).
Wigle, D.A. et al. Molecular profiling of non–small cell lung cancer and correlation with disease-free survival. Cancer Res. 62, 3005–3008 (2002).
Potti, A. et al. A genomic strategy to refine prognosis in early-stage non–small-cell lung cancer. N. Engl. J. Med. 355, 570–580 (2006).
Chen, H.Y. et al. A five-gene signature and clinical outcome in non-small-cell lung cancer. N. Engl. J. Med. 356, 11–20 (2007).
Lu, Y. et al. A gene expression signature predicts survival of subjects with stage I non-small cell lung cancer. PLoS Med. 12, e467 (2006).
Dobbin, K.K. et al. Interlaboratory comparability study of cancer gene expression analysis using oligonucleotide microarrays. Clin. Cancer Res. 11, 565–572 (2005).
Fry, W.A., Phillips, J.L. & Menck, H.R. Ten-year survey of lung cancer treatments and survival in hospitals in the United States. Cancer 86, 1867–1876 (1999).
Li, C. & Wong, W.H. Model-based analysis of oligonucleotide arrays: expression index computation and outlier detection. Proc. Natl. Acad. Sci. USA 98, 31–36 (2001).
Moran, C.J. et al. Rantes expression by lung adenocarcinomas is a predictor of survival in stage I subjects. Clin. Cancer Res. 8, 3803–3812 (2002).
Stephenson, A.J. et al. Integration of gene expression profiling and clinical variables to predict prostate carcinoma recurrence after radical prostatectomy. Cancer 104, 290–298 (2005).
Sotiriou, C. & Piccart, M.J. Taking gene-expression profiling to the clinic: when will molecular signatures become relevant to subject care? Nat. Rev. Cancer 7, 545–553 (2007).
Gonen, M. & Heller, G. Concordance probability and discriminatory power in proportional hazards regression. Biometrika 92, 965–970 (2005).
Acknowledgements
We thank M. Orringer, A. Pickens, F. Taylor, N. Liu, D. Lau, M. Whitehead, L. Chen, L. Vargas, Y. Xiao, M. Maddaus and C. Hoang. We thank M. Heiskanen, L. Liu, D. Reeves and S. Whitley from the US National Cancer Institute Center for Bioinformatics and W. Ricker from Information Management Services for assistance with development of the lung study database and data management. We thank D. Sawyer, J.M. Askew and A. Vaughn of the Cancer and Leukemia Group B Statistical Center, Duke University for quality control of the clinical data. We thank Affymetrix for technical support. This work was supported by US National Cancer Institute grants CA84953, CA84999, CA84995, CA85052 and CA46592 and contracts 263-MQ-319735, 263-MQ-319740, 263-MQ-319746 and 263-MQ-510430 and support from the Canadian Cancer Society.
Author information
Authors and Affiliations
Consortia
Contributions
Writing Committee: K.S., J.M.G.T., S.A.E., M.S.T., T.J.Y., W.L.G., S.E., I.J., V.E.S., M.M., R.K., K.K.D., T.L., J.W.J. and D.G.B. Members of the Writing Committee participated in the planning, initiation, data generation, data analysis and manuscript preparation for the project.
Additional participants: T.J.G., D.E.M., A.C.C. and S.H. participated in aspects of sample collection and preparation, data generation and data analysis at the University of Michigan. C.Q.Z., D.S., F.A.S., K.D. and L.S. participated in aspects of sample collection and preparation, data generation and data analysis at the Ontario Cancer Institute. K.N., N.P., B.W., R.V., C.L.-A and T.G. participated in aspects of sample collection and preparation, data generation and data analysis at the Dana-Farber Cancer Institute and Broad Institute. M.G. assembled the clinical data at the H. Lee Moffitt Cancer Center. J.S., M.Z., V.R., M.K., A.V., N.M., W.T. and A.S. participated in aspects of sample collection and preparation, data generation and data analysis at Memorial Sloan-Kettering Cancer Center. B.C. participated in the planning and initiation of the study.
Corresponding authors
Additional information
The consortium consists of the Writing Committee plus additional participants as detailed in the Author Contributions section.
Supplementary information
Supplementary Text and Figures
Supplementary Figs. 1 and 2 (PDF 2763 kb)
Supplementary Table 1
Supplementary Data (XLS 409 kb)
Rights and permissions
About this article
Cite this article
Director's Challenge Consortium for the Molecular Classification of Lung Adenocarcinoma. Gene expression–based survival prediction in lung adenocarcinoma: a multi-site, blinded validation study. Nat Med 14, 822–827 (2008). https://doi.org/10.1038/nm.1790
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nm.1790
This article is cited by
-
Clonal gene signatures predict prognosis in mesothelioma and lung adenocarcinoma
npj Precision Oncology (2024)
-
Comprehensive analysis of co-expressed genes with TDP-43: prognostic and therapeutic potential in lung adenocarcinoma
Journal of Cancer Research and Clinical Oncology (2024)
-
Identification of four metabolic subtypes and key prognostic markers in lung adenocarcinoma based on glycolytic and glutaminolytic pathways
BMC Cancer (2023)
-
A comparative study of clustering methods on gene expression data for lung cancer prognosis
BMC Research Notes (2023)
-
Immunological role and clinical prognostic significance of P2RY6 in lung adenocarcinoma: a multi-omics studies and single-cell sequencing analysis
World Journal of Surgical Oncology (2023)