Abstract
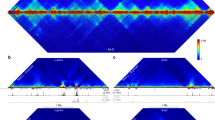
Unlike the well defined T helper type 2 cytokine locus, little is known about the regulatory elements that govern the expression of Ifng, which encodes the 'signature' T helper type 1 cytokine interferon-γ. Here our evolutionary analysis showed that the mouse Ifng locus diverged from the ancestral locus as a result of structural rearrangements producing deletion of the neighboring gene encoding interleukin 26 and disrupting synteny 57 kilobases upstream of Ifng. Proximal to that disruption, we identified by high-resolution mapping many regions with CD4+ T cell subset–specific epigenetic modifications. A subset of those regions represented enhancers, whereas others blocked the activity of upstream enhancers or insulated Ifng from neighboring chromatin. Our findings suggest that proper expression of Ifng is maintained through the collective action of multiple distal regulatory elements present in a region of about 100 kilobases flanking Ifng.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Change history
05 July 2007
In the version of this article initially published, the surname of the penultimate author is misspelled. The correct spelling is Stamatoyannopoulos. The error has been corrected in the HTML and PDF versions of the article.
16 November 2007
In the version of this article initially published, labels in Figures 2 and 4 are incorrect (as is the relevant text for Figure 4), and a reagent is incorrectly identified in Methods. In Figure 2, the label “IfngCNS+54” should be “IfngCNS+55.” For Figure 4, the primers used to analyze the intronic region of Ifng for CpG methylation after bisulfite treatment amplify intron 1; thus, the label above the sixth column in Figure 4a should be “Ifng intron 1” and the accompanying text on page 736, column 1, should read “intron 1” on lines 3 and 13. In the Methods section, page 740 column 2, the end of line 23 should read “rabbit immunoglobulin G.” The errors have been corrected in the PDF version of the article.
References
Ho, I.C. & Glimcher, L.H. Transcription: tantalizing times for T cells. Cell 109 Suppl., S109–S120 (2002).
Murphy, K.M. & Reiner, S.L. The lineage decisions of helper T cells. Nat. Rev. Immunol. 2, 933–944 (2002).
Ansel, K.M., Lee, D.U. & Rao, A. An epigenetic view of helper T cell differentiation. Nat. Immunol. 4, 616–623 (2003).
Szabo, S.J., Sullivan, B.M., Peng, S.L. & Glimcher, L.H. Molecular mechanisms regulating Th1 immune responses. Annu. Rev. Immunol. 21, 713–758 (2003).
Hwang, E.S., Szabo, S.J., Schwartzberg, P.L. & Glimcher, L.H. T helper cell fate specified by kinase-mediated interaction of T-bet with GATA-3. Science 307, 430–433 (2005).
Usui, T. et al. T-bet regulates Th1 responses through essential effects on GATA-3 function rather than on IFNG gene acetylation and transcription. J. Exp. Med. 203, 755–766 (2006).
Mullen, A.C. et al. Role of T-bet in commitment of TH1 cells before IL-12-dependent selection. Science 292, 1907–1910 (2001).
Afkarian, M. et al. T-bet is a STAT1-induced regulator of IL-12R expression in naive CD4+ T cells. Nat. Immunol. 3, 549–557 (2002).
Mullen, A.C. et al. Hlx is induced by and genetically interacts with T-bet to promote heritable TH1 gene induction. Nat. Immunol. 3, 652–658 (2002).
Lee, G.R., Kim, S.T., Spilianakis, C.G., Fields, P.E. & Flavell, R.A. T helper cell differentiation: regulation by cis elements and epigenetics. Immunity 24, 369–379 (2006).
Young, H.A. et al. Expression of human IFN-γ genomic DNA in transgenic mice. J. Immunol. 143, 2389–2394 (1989).
Zhu, H. et al. Unexpected characteristics of the IFN-γ reporters in nontransformed T cells. J. Immunol. 167, 855–865 (2001).
Soutto, M., Zhou, W. & Aune, T.M. Cutting edge: distal regulatory elements are required to achieve selective expression of IFN-γ in Th1/Tc1 effector cells. J. Immunol. 169, 6664–6667 (2002).
Gaszner, M. & Felsenfeld, G. Insulators: exploiting transcriptional and epigenetic mechanisms. Nat. Rev. Genet. 7, 703–713 (2006).
Valenzuela, L. & Kamakaka, R.T. Chromatin insulators. Annu. Rev. Genet. 40, 107–138 (2006).
West, A.G. & Fraser, P. Remote control of gene transcription. Hum. Mol. Genet. 14, R101–11 (2005).
Nardone, J., Lee, D.U., Ansel, K.M. & Rao, A. Bioinformatics for the `bench biologist': how to find regulatory regions in genomic DNA. Nat. Immunol. 5, 768–774 (2004).
Dorschner, M.O. et al. High-throughput localization of functional elements by quantitative chromatin profiling. Nat. Methods 1, 219–225 (2004).
Igawa, D., Sakai, M. & Savan, R. An unexpected discovery of two interferon gamma-like genes along with interleukin (IL)-22 and -26 from teleost: IL-22 and -26 genes have been described for the first time outside mammals. Mol. Immunol. 43, 999–1009 (2006).
Dumoutier, L., Van Roost, E., Ameye, G., Michaux, L. & Renauld, J.C. IL-TIF/IL-22: genomic organization and mapping of the human and mouse genes. Genes Immun. 1, 488–494 (2000).
Vigneau, S., Rohrlich, P.S., Brahic, M. & Bureau, J.F. Tmevpg1, a candidate gene for the control of Theiler's virus persistence, could be implicated in the regulation of gamma interferon. J. Virol. 77, 5632–5638 (2003).
Lee, D.U., Avni, O., Chen, L. & Rao, A. A distal enhancer in the interferon-γ (IFN- γ) locus revealed by genome sequence comparison. J. Biol. Chem. 279, 4802–4810 (2004).
Shnyreva, M. et al. Evolutionarily conserved sequence elements that positively regulate IFN-γ expression in T cells. Proc. Natl. Acad. Sci. USA 101, 12622–12627 (2004).
Bernstein, B.E. et al. Genomic maps and comparative analysis of histone modifications in human and mouse. Cell 120, 169–181 (2005).
Agarwal, S. & Rao, A. Modulation of chromatin structure regulates cytokine gene expression during T cell differentiation. Immunity 9, 765–775 (1998).
Schneider, R. et al. Histone H3 lysine 4 methylation patterns in higher eukaryotic genes. Nat. Cell Biol. 6, 73–77 (2004).
Cao, R. et al. Role of histone H3 lysine 27 methylation in Polycomb-group silencing. Science 298, 1039–1043 (2002).
Bracken, A.P., Dietrich, N., Pasini, D., Hansen, K.H. & Helin, K. Genome-wide mapping of Polycomb target genes unravels their roles in cell fate transitions. Genes Dev. 20, 1123–1136 (2006).
Lee, C.K., Shibata, Y., Rao, B., Strahl, B.D. & Lieb, J.D. Evidence for nucleosome depletion at active regulatory regions genome-wide. Nat. Genet. 36, 900–905 (2004).
Chen, X., Wang, J., Woltring, D., Gerondakis, S. & Shannon, M.F. Histone dynamics on the interleukin-2 gene in response to T-cell activation. Mol. Cell. Biol. 25, 3209–3219 (2005).
Simon, J.A. & Tamkun, J.W. Programming off and on states in chromatin: mechanisms of Polycomb and trithorax group complexes. Curr. Opin. Genet. Dev. 12, 210–218 (2002).
Lee, D.U., Agarwal, S. & Rao, A. Th2 lineage commitment and efficient IL-4 production involves extended demethylation of the IL-4 gene. Immunity 16, 649–660 (2002).
Makar, K.W. et al. Active recruitment of DNA methyltransferases regulates interleukin 4 in thymocytes and T cells. Nat. Immunol. 4, 1183–1190 (2003).
Zhong, X.P. & Krangel, M.S. An enhancer-blocking element between α and δ gene segments within the human T cell receptor α/δ locus. Proc. Natl. Acad. Sci. USA 94, 5219–5224 (1997).
Gombert, W.M. et al. The c-myc insulator element and matrix attachment regions define the c-myc chromosomal domain. Mol. Cell. Biol. 23, 9338–9348 (2003).
Grogan, J.L. & Locksley, R.M. T helper cell differentiation: on again, off again. Curr. Opin. Immunol. 14, 366–372 (2002).
Avni, O. et al. TH cell differentiation is accompanied by dynamic changes in histone acetylation of cytokine genes. Nat. Immunol. 3, 643–651 (2002).
Fields, P.E., Kim, S.T. & Flavell, R.A. Cutting edge: changes in histone acetylation at the IL-4 and IFN-γ loci accompany Th1/Th2 differentiation. J. Immunol. 169, 647–650 (2002).
Spilianakis, C.G., Lalioti, M.D., Town, T., Lee, G.R. & Flavell, R.A. Interchromosomal associations between alternatively expressed loci. Nature 435, 637–645 (2005).
Vigneau, S. et al. Homology between a 173-kb region from mouse chromosome 10, telomeric to the Ifng locus, and human chromosome 12q15. Genomics 78, 206–213 (2001).
Bernstein, B.E. et al. A bivalent chromatin structure marks key developmental genes in embryonic stem cells. Cell 125, 315–326 (2006).
Ansel, K.M., Djuretic, I., Tanasa, B. & Rao, A. Regulation of Th2 differentiation and Il4 locus accessibility. Annu. Rev. Immunol. 24, 607–656 (2006).
Hatton, R.D. et al. A distal conserved sequence element controls Ifng gene expression by T cells and NK cells. Immunity 25, 717–729 (2006).
Chang, S. & Aune, T.M. Histone hyperacetylated domains across the Ifng gene region in natural killer cells and T cells. Proc. Natl. Acad. Sci. USA 102, 17095–17100 (2005).
Biemont, C. & Vieira, C. Genetics: junk DNA as an evolutionary force. Nature 443, 521–524 (2006).
Xie, M.H. et al. Interleukin (IL)-22, a novel human cytokine that signals through the interferon receptor-related proteins CRF2–4 and IL-22R. J. Biol. Chem. 275, 31335–31339 (2000).
Liang, S.C. et al. Interleukin (IL)-22 and IL-17 are coexpressed by Th17 cells and cooperatively enhance expression of antimicrobial peptides. J. Exp. Med. 203, 2271–2279 (2006).
Koyanagi, M. et al. EZH2 and histone 3 trimethyl lysine 27 associated with Il4 and Il13 gene silencing in Th1 cells. J. Biol. Chem. 280, 31470–31477 (2005).
Messi, M. et al. Memory and flexibility of cytokine gene expression as separable properties of human TH1 and TH2 lymphocytes. Nat. Immunol. 4, 78–86 (2003).
Fitzpatrick, D.R. et al. Distinct methylation of the interferon γ (IFN-γ) and interleukin 3 (IL-3) genes in newly activated primary CD8+ T lymphocytes: regional IFN-γ promoter demethylation and mRNA expression are heritable in CD44highCD8+ T cells. J. Exp. Med. 188, 103–117 (1998).
Acknowledgements
We thank C. Surh (Scripps Research Institute) and M. Bevan (University of Washington) for Thy-1.1+ SMARTA TCR-transgenic mice; T. Krumm (University of Washington) and M. Krangel (Duke University) for the Eδ-Pδ-βNeo-scs′ and Pδ-BNeo-scs′ constructs; M. Kaja for LCMV Armstrong; K. Arispe and M. Weaver for technical assistance; S. Taylor for an earlier computational analysis of the Ifng locus; and E. Eichler and M. Orr for comments. Supported by the National Institutes of Health (AI071272 and HD18184; and CA009537 to J.R.S.), the March of Dimes and the Cancer Research Institute (J.R.S.).
Author information
Authors and Affiliations
Contributions
J.R.S. designed and did research, analyzed data and wrote the paper; M.O.D. designed and did research and analyzed data; M. Sekimata did research and analyzed data; D.M.S. did research and analyzed data; M. Shnyreva did the chromatin immunoprecipitation of D10 and AE7 cells; D.R.F. contributed intellectually and helped write the paper; J.A.S. designed research; C.B.W. designed research, analyzed data and wrote the paper; and all authors reviewed the paper before submission.
Corresponding author
Ethics declarations
Competing interests
D.R.F. is a former employee and shareholder of Amgen.
Supplementary information
Supplementary Fig. 1
The rodent Ifng locus contains a complex set of structural rearrangements and segmental duplications beginning 57 kb upstream of Ifng that has disrupted Il26. (PDF 180 kb)
Supplementary Fig. 2
Generation and production of IFN-γ by in vitro TH1 and TH2 cultures or in vivo CD4+ effectors. (PDF 110 kb)
Supplementary Fig. 3
ChIP for total histone H3 in the Ifng locus in naive, TH1 or TH2 CD4+ T cells. (PDF 84 kb)
Supplementary Fig. 4
ChIP for acetylated histone H3 in the Ifng locus in long-term TH1 (AE7) and TH2 (D10) cell lines. (PDF 60 kb)
Supplementary Table 1
Location of DNase HS sites in primary CD4 naive, TH1 and TH2 cells. (PDF 30 kb)
Supplementary Table 2
Chromatin immunoprecipitation primers. (PDF 11 kb)
Supplementary Table 3
Primers used in DNase HS analysis based off of mm8 C57BL/6 sequence (Ifng start at 117844040). (PDF 40 kb)
Supplementary Table 4
BIS-PCR primers for CpG methylation analysis. (PDF 50 kb)
Rights and permissions
About this article
Cite this article
Schoenborn, J., Dorschner, M., Sekimata, M. et al. Comprehensive epigenetic profiling identifies multiple distal regulatory elements directing transcription of the gene encoding interferon-γ. Nat Immunol 8, 732–742 (2007). https://doi.org/10.1038/ni1474
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ni1474
This article is cited by
-
Regulation of antitumor immunity by inflammation-induced epigenetic alterations
Cellular & Molecular Immunology (2022)
-
Low-dose decitabine priming endows CAR T cells with enhanced and persistent antitumour potential via epigenetic reprogramming
Nature Communications (2021)
-
Cancer SLC43A2 alters T cell methionine metabolism and histone methylation
Nature (2020)
-
Extrinsic and Intrinsic Immunometabolism Converge: Perspectives on Future Research and Therapeutic Development for Obesity
Current Obesity Reports (2019)
-
A conserved enhancer regulates Il9 expression in multiple lineages
Nature Communications (2018)