Abstract
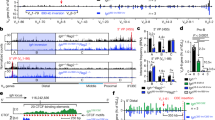
Reversible contraction of immunoglobulin loci juxtaposes the variable (V) genes next to the (diversity)-joining-constant ((D)JC) gene domain, thus facilitating V-(D)J recombination. Here we show that the T cell receptor β (Tcrb) and T cell receptor αδ (Tcra-Tcrd) loci also underwent long-range interactions by looping in double-negative and double-positive thymocytes, respectively. Contraction of the Tcrb and Tcra loci occurred in rearranging thymocytes and was reversed at the next developmental stage. Decontraction of the Tcrb locus probably prevented further Vβ-DJβ rearrangements in double-positive thymocytes by separating the Vβ genes from the DJCβ domain. In most double-negative cells, one Tcrb allele was recruited to pericentromeric heterochromatin. Such allelic positioning may facilitate asynchronous Vβ-DJβ recombination. Hence, pericentromeric recruitment and locus 'decontraction' seem to contribute to the initiation and maintenance of allelic exclusion at the Tcrb locus.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Bassing, C.H., Swat, W. & Alt, F.W. The mechanism and regulation of chromosomal V(D)J recombination. Cell 109 Suppl., S45–S55 (2002).
Melchers, F. The pre–B-cell receptor: selector of fitting immunoglobulin heavy chains for the B-cell repertoire. Nat. Rev. Immunol. 5, 578–584 (2005).
von Boehmer, H. Unique features of the pre–T-cell receptor α-chain: not just a surrogate. Nat. Rev. Immunol. 5, 571–577 (2005).
Yancopoulos, G.D. & Alt, F.W. Developmentally controlled and tissue-specific expression of unrearranged VH gene segments. Cell 40, 271–281 (1985).
Stanhope-Baker, P., Hudson, K.M., Shaffer, A.L., Constantinescu, A. & Schlissel, M.S. Cell type–specific chromatin structure determines the targeting of V(D)J recombinase activity in vitro. Cell 85, 887–897 (1996).
McMurry, M.T. & Krangel, M.S. A role for histone acetylation in the developmental regulation of V(D)J recombination. Science 287, 495–498 (2000).
Kosak, S.T. et al. Subnuclear compartmentalization of immunoglobulin loci during lymphocyte development. Science 296, 158–162 (2002).
Mostoslavsky, R. et al. κ chain monoallelic demethylation and the establishment of allelic exclusion. Genes Dev. 12, 1801–1811 (1998).
Kwon, J., Morshead, K.B., Guyon, J.R., Kingston, R.E. & Oettinger, M.A. Histone acetylation and hSWI/SNF remodeling act in concert to stimulate V(D)J cleavage of nucleosomal DNA. Mol. Cell 6, 1037–1048 (2000).
Chowdhury, D. & Sen, R. Stepwise activation of the immunoglobulin μ heavy chain gene locus. EMBO J. 20, 6394–6403 (2001).
Tripathi, R., Jackson, A. & Krangel, M.S. A change in the structure of Vβ chromatin associated with TCR β allelic exclusion. J. Immunol. 168, 2316–2324 (2002).
Bolland, D.J. et al. Antisense intergenic transcription in V(D)J recombination. Nat. Immunol. 5, 630–637 (2004).
Johnston, C.M., Wood, A.L., Bolland, D.J. & Corcoran, A.E. Complete sequence assembly and characterization of the C57BL/6 mouse Ig heavy chain V region. J. Immunol. 176, 4221–4234 (2006).
Thiebe, R. et al. The variable genes and gene families of the mouse immunoglobulin κ locus. Eur. J. Immunol. 29, 2072–2081 (1999).
Fuxa, M. et al. Pax5 induces V-to-DJ rearrangements and locus contraction of the immunoglobulin heavy-chain gene. Genes Dev. 18, 411–422 (2004).
Roldán, E. et al. Locus 'decontraction' and centromeric recruitment contribute to allelic exclusion of the immunoglobulin heavy-chain gene. Nat. Immunol. 6, 31–41 (2005).
Sayegh, C., Jhunjhunwala, S., Riblet, R. & Murre, C. Visualization of looping involving the immunoglobulin heavy-chain locus in developing B cells. Genes Dev. 19, 322–327 (2005).
Bosc, N. & Lefranc, M.P. The mouse (Mus musculus) T cell receptor β variable (TRBV), diversity (TRBD) and joining (TRBJ) genes. Exp. Clin. Immunogenet. 17, 216–228 (2000).
Jackson, A.M. & Krangel, M.S. Turning T-cell receptor β recombination on and off: more questions than answers. Immunol. Rev. 209, 129–141 (2006).
Bosc, N. & Lefranc, M.P. The mouse (Mus musculus) T cell receptor α (TRA) and δ (TRD) variable genes. Dev. Comp. Immunol. 27, 465–497 (2003).
Krangel, M.S., Carabana, J., Abbarategui, I., Schlimgen, R. & Hawwari, A. Enforcing order within a complex locus: current perspectives on the control of V(D)J recombination at the murine T-cell receptor α/δ locus. Immunol. Rev. 200, 224–232 (2004).
Schmitt, T.M. & Zúñiga-Pflücker, J.C. Induction of T cell development from hematopoietic progenitor cells by delta-like-1 in vitro. Immunity 17, 749–756 (2002).
Höflinger, S. et al. Analysis of Notch1 function by in vitro T cell differentiation of Pax5 mutant lymphoid progenitors. J. Immunol. 173, 3935–3944 (2004).
Skok, J.A. et al. Nonequivalent nuclear location of immunoglobulin alleles in B lymphocytes. Nat. Immunol. 2, 848–854 (2001).
Dekker, J., Rippe, K., Dekker, M. & Kleckner, N. Capturing chromosome conformation. Science 295, 1306–1311 (2002).
Tolhuis, B., Palstra, R-J., Splinter, E., Grosveld, F. & de Laat, W. Looping and interaction between hypersensitive sites in the active β-globin locus. Mol. Cell 10, 1453–1465 (2002).
Splinter, E., Grosveld, F. & de Laat, W. 3C technology: analyzing the spatial organization of genomic loci in vivo. Methods Enzymol. 375, 493–507 (2004).
Shinkai, Y. et al. Restoration of T cell development in RAG-2–deficient mice by functional TCR transgenes. Science 259, 822–825 (1993).
Goldmit, M. et al. Epigenetic ontogeny of the Igk locus during B cell development. Nat. Immunol. 6, 198–203 (2005).
Bories, J.-C., Demengeot, J., Davidson, L. & Alt, F.W. Gene-targeted deletion and replacement mutations of the T-cell receptor β-chain enhancer: the role of enhancer elements in controlling V(D)J recombination accessibility. Proc. Natl. Acad. Sci. USA 93, 7871–7876 (1996).
Whitehurst, C.E., Chattopadhyay, S. & Chen, J. Control of V(D)J recombinational accessibility of the Dβ1 gene segment at the TCRβ locus by a germline promoter. Immunity 10, 313–322 (1999).
Mathieu, N., Hempel, W.M., Spicuglia, S., Verthuy, C. & Ferrier, P. Chromatin remodeling by the T cell receptor (TCR)-β gene enhancer during early T cell development: implications for the control of TCR-β locus recombination. J. Exp. Med. 192, 625–636 (2000).
Hawwari, A. & Krangel, M.S. Regulation of TCR δ and α repertoires by local and long-distance control of variable gene segment chromatin structure. J. Exp. Med. 202, 467–472 (2005).
Oestreich, K.J. et al. Regulation of TCRβ gene assembly by a promoter/enhancer holocomplex. Immunity 24, 381–391 (2006).
Glusman, G. et al. Comparative genomics of the human and mouse T cell receptor loci. Immunity 15, 337–349 (2001).
Patrinos, G.P. et al. Multiple interactions between regulatory regions are required to stabilize an active chromatin hub. Genes Dev. 18, 1495–1509 (2004).
Lee, G.R., Spilianakis, C.G. & Flavell, R.A. Hypersensitive site 7 of the TH2 locus control region is essential for expressing TH2 cytokine genes and for long-range intrachromosomal interactions. Nat. Immunol. 6, 42–48 (2005).
Ryu, C.J. et al. The T-cell receptor β variable gene promoter is required for efficient Vβ rearrangement but not allelic exclusion. Mol. Cell. Biol. 24, 7015–7023 (2004).
Wilson, A., Held, W. & MacDonald, H.R. Two waves of recombinase gene expression in developing thymocytes. J. Exp. Med. 179, 1355–1360 (1994).
Mathieu, N. et al. Assessing the role of the T cell receptor β gene enhancer in regulating coding joint formation during V(D)J recombination. J. Biol. Chem. 278, 18101–18109 (2003).
Chowdhury, D. & Sen, R. Transient IL-7/IL-7R signaling provides a mechanism for feedback inhibition of immunoglobulin heavy chain gene rearrangements. Immunity 18, 229–241 (2003).
Wang, L., Senoo, M. & Habu, S. Differential regulation between gene expression and histone H3 acetylation in the variable regions of the TCRβ locus. Biochem. Biophys. Res. Commun. 298, 420–426 (2002).
Jackson, A., Kondilis, H.D., Khor, B., Sleckman, B.P. & Krangel, M.S. Regulation of T cell receptor β allelic exclusion at a level beyond accessibility. Nat. Immunol. 6, 189–197 (2005).
Sieh, P. & Chen, J. Distinct control of the frequency and allelic exclusion of the Vβ gene rearrangement at the TCRβ locus. J. Immunol. 167, 2121–2129 (2001).
Senoo, M. et al. Increase of TCR Vβ accessibility within Eβ regulatory region influences its recombination frequency but not allelic exclusion. J. Immunol. 171, 829–835 (2003).
Mostoslavsky, R. et al. Asynchronous replication and allelic exclusion in the immune system. Nature 414, 221–225 (2001).
Liang, H.E., Hsu, L-Y., Cado, D. & Schlissel, M.S. Variegated transcriptional activation of the immunoglobulin κ locus in pre-B cells contributes to the allelic exclusion of light-chain expression. Cell 118, 19–29 (2004).
Sleckman, B.P., Khor, B., Monroe, R. & Alt, F.W. Assembly of productive T cell receptor δ variable region genes exhibits allelic inclusion. J. Exp. Med. 188, 1465–1471 (1998).
Padovan, E. et al. Expression of two T cell receptor α chains: dual receptor T cells. Science 262, 422–424 (1993).
Pasqual, N. et al. Quantitative and qualitative changes in V-J α rearrangements during mouse thymocytes differentiation: implication for a limited T cell receptor α chain repertoire. J. Exp. Med. 196, 1163–1173 (2002).
Su, I-H. et al. Ezh2 controls B cell development through histone H3 methylation and Igh rearrangement. Nat. Immunol. 4, 124–131 (2003).
Francis, N.J., Kingston, R.E. & Woodcock, C.L. Chromatin compaction by a polycomb group protein complex. Science 306, 1574–1577 (2004).
Su, I-H. et al. Polycomb group protein Ezh2 controls actin polymerization and cell signaling. Cell 121, 425–436 (2005).
Wolfer, A., Wilson, A., Nemir, M., MacDonald, H.R. & Radtke, F. Inactivation of Notch1 impairs VDJβ rearrangement and allows pre-TCR-independent survival of early αβ lineage thymocytes. Immunity 16, 869–879 (2002).
Bender, T.P., Kremer, C.S., Kraus, M., Buch, T. & Rajewsky, K. Critical functions for c-Myb at three checkpoints during thymocyte development. Nat. Immunol. 5, 721–729 (2004).
Pircher, H. et al. T cell receptor (TcR) β chain transgenic mice: studies on allelic exclusion and on the TcR+ γ/δ population. Eur. J. Immunol. 20, 417–424 (1990).
Klein, L., Klein, T., Rüther, U. & Kyewski, B. CD4 T cell tolerance to human C-reactive protein, an inducible serum protein, is mediated by medullary thymic epithelium. J. Exp. Med. 188, 5–16 (1998).
Acknowledgements
We thank P. Ferrier (Centre d'Immunologie de Marseille-Luminy) for Tcra-transgenic Rag2−/− mice; L. Klein (Research Institute of Molecular Pathology) for Tcra-Tcrb-transgenic Rag2−/− mice; A. Souabni for help with the OP9-DL1 differentiation system; and G. Stengl for flow cytometry sorting. Supported by Boehringer Ingelheim, the Austrian GEN-AU initiative (financed by the Bundesminsterium für Bildung, Wissenschaft und Kultur), the Wellcome Trust (J.A.S.) and the Swedish Research Council (R.G.).
Author information
Authors and Affiliations
Contributions
J.A.S. did most of the FISH experiments and did all confocal analyses of the FISH data; D.F. contributed to some FISH experiments and prepared DNA-FISH probes; R.G. did 3C analyses, in vitro differentiation experiments and flow cytometry sorting; W.d.L. provided training and supervision for the 3C experiments; M.N. did the bioinformatical analyses of the TCR loci; and M.B. planned the experimental outline and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Fig. 1
FACS analysis of the lymphoid cells used for 3C analysis. (PDF 217 kb)
Supplementary Fig. 2
Extended configuration of the Tcrb and Tcra/d loci in pre-B cells. (PDF 292 kb)
Supplementary Fig. 3
Additional pictures with looping configurations of the Tcrb and Tcra/d alleles. (PDF 500 kb)
Supplementary Fig. 4
Representative PCR analysis of a 3C experiment. (PDF 217 kb)
Supplementary Fig. 5
Linearity of the PCR assay and BglII digestion of the crosslinked chromatin. (PDF 243 kb)
Supplementary Fig. 6
Interactions of the proximal Dβ1 region with distal Vβ genes of the Tcrb locus in DN pro-T cells. (PDF 181 kb)
Supplementary Fig. 7
Peripheral and pericentromeric location of the Tcrb and Tcra/d loci. (PDF 191 kb)
Supplementary Table 1
Separation of Tcrb and Tcra/d probe signals in developing T cells. (PDF 32 kb)
Supplementary Table 2
Subnuclear location and monoallelic pericentromeric recruitment of the Tcrb and Tcra/d loci in thymocytes and peripheral T cells (PDF 28 kb)
Supplementary Table 3
Predominant recruitment of the Tcrb and Tcra/d loci to pericentromeric heterochromatin. (PDF 74 kb)
Supplementary Table 4
PCR primers used for 3C analysis of the Tcrb locus. (PDF 27 kb)
Rights and permissions
About this article
Cite this article
Skok, J., Gisler, R., Novatchkova, M. et al. Reversible contraction by looping of the Tcra and Tcrb loci in rearranging thymocytes. Nat Immunol 8, 378–387 (2007). https://doi.org/10.1038/ni1448
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ni1448
This article is cited by
-
T cell receptor repertoires of ex vivo–expanded tumor-infiltrating lymphocytes from breast cancer patients
Immunologic Research (2020)
-
Genome organization in immune cells: unique challenges
Nature Reviews Immunology (2019)
-
Nuclear positioning and pairing of X-chromosome inactivation centers are not primary determinants during initiation of random X-inactivation
Nature Genetics (2019)
-
A discrete chromatin loop in the mouse Tcra-Tcrd locus shapes the TCRδ and TCRα repertoires
Nature Immunology (2015)
-
Highly diverse TCRα chain repertoire of pre-immune CD8+T cells reveals new insights in gene recombination
The EMBO Journal (2012)