Abstract
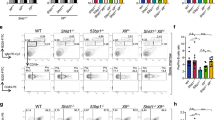
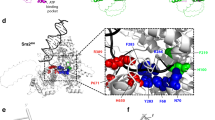
Activation-induced cytidine deaminase (AID) is essential for class-switch recombination (CSR) and somatic hypermutation (SHM). Mutants with changes in the C-terminal region of AID retain SHM but lose CSR activity. Here we describe five mutants with alterations in the N-terminal region of AID that caused selective deficiency in SHM but retained CSR, suggesting that the CSR and SHM activities of AID may dissociate via interaction of CSR- or SHM-specific cofactors with different domains of AID. Unlike cells expressing C-terminal AID mutants, B cells expressing N-terminal AID mutants had mutations in the switch μ region, indicating that such mutations are generated by reactions involved in CSR but not SHM. Thus, we propose that separate domains of AID interact with specific cofactors to regulate these two distinct genetic events in a target-specific way.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Change history
20 June 2004
added erratum PDF to AOP PDF, added note to XML, and corrected online date will be included in the issue PDF
Notes
*Note: In the version of this article originally published online, the values for "Total base number" in Table 4 were incomplete. The correct numbers are as follows: 24,487; 25,008; 23,445; 10,420; and 20,840. Line 3 of the legend for Figure 1 should read "mAID, mouse AID," and line 1 of the legend to Table 2 legend should begin "DNA". This error has been corrected for the HTML and print versions of this article.
References
Honjo, T., Kinoshita, K. & Muramatsu, M. Molecular mechanism of class switch recombination: linkage with somatic hypermutation. Annu. Rev. Immunol. 20, 165–196 (2002).
Luby, T.M., Schrader, C.E., Stavnezer, J. & Selsing, E. The μ switch region tandem repeats are important, but not required, for antibody class switch recombination. J. Exp. Med. 193, 159–168 (2001).
Shinkura, R. et al. The influence of transcriptional orientation on endogenous switch region function. Nat. Immunol. 4, 435–441 (2003).
Rogozin, I.B. & Diaz, M. Cutting edge: DGYW/WRCH is a better predictor of mutability at G:C bases in Ig hypermutation than the widely accepted RGYW/WRCY motif and probably reflects a two-step activation-induced cytidine deaminase-triggered process. J. Immunol. 172, 3382–3384 (2004).
Milstein, C., Neuberger, M.S. & Staden, R. Both DNA strands of antibody genes are hypermutation targets. Proc. Natl. Acad. Sci. USA 95, 8791–8794 (1998).
Muramatsu, M. et al. Class switch recombination and hypermutation require activation-induced cytidine deaminase (AID), a potential RNA editing enzyme. Cell 102, 553–563 (2000).
Revy, P. et al. Activation-induced cytidine deaminase (AID) deficiency causes the autosomal recessive form of the Hyper-IgM syndrome (HIGM2). Cell 102, 565–575 (2000).
Yoshikawa, K. et al. AID enzyme-induced hypermutation in an actively transcribed gene in fibroblasts. Science 296, 2033–2036 (2002).
Okazaki, I.M., Kinoshita, K., Muramatsu, M., Yoshikawa, K. & Honjo, T. The AID enzyme induces class switch recombination in fibroblasts. Nature 416, 340–345 (2002).
Martin, A. & Scharff, M.D. Somatic hypermutation of the AID transgene in B and non-B cells. Proc. Natl. Acad. Sci. USA 99, 12304–12308 (2002).
Martin, A. et al. Activation-induced cytidine deaminase turns on somatic hypermutation in hybridomas. Nature 415, 802–806 (2002).
Petersen, S. et al. AID is required to initiate Nbs1/γ-H2AX focus formation and mutations at sites of class switching. Nature 414, 660–665 (2001).
Doi, T., Kinoshita, K., Ikegawa, M., Muramatsu, M. & Honjo, T. De novo protein synthesis is required for the activation-induced cytidine deaminase function in class-switch recombination. Proc. Natl. Acad. Sci. USA 100, 2634–2638 (2003).
Dickerson, S.K., Market, E., Besmer, E. & Papavasiliou, F.N. AID mediates hypermutation by deaminating single stranded DNA. J. Exp. Med. 197, 1291–1296 (2003).
Peterson-Mahrt, S.K. & Neuberger, M.S. In vitro deamination of cytosine to uracil in single-stranded DNA by apolipoprotein B editing complex catalytic subunit 1 (APOBEC1). J. Biol. Chem. 278, 19583–19586 (2003).
Ramiro, A.R., Stavropoulos, P., Jankovic, M. & Nussenzweig, M.C. Transcription enhances AID-mediated cytidine deamination by exposing single-stranded DNA on the nontemplate strand. Nat. Immunol. 4, 452–456 (2003).
Bransteitter, R., Pham, P., Scharff, M.D. & Goodman, M.F. Activation-induced cytidine deaminase deaminates deoxycytidine on single-stranded DNA but requires the action of RNase. Proc. Natl. Acad. Sci. USA 100, 4102–4107 (2003).
Chaudhuri, J. et al. Transcription-targeted DNA deamination by the AID antibody diversification enzyme. Nature 422, 726–730 (2003).
Sohail, A., Klapacz, J., Samaranayake, M., Ullah, A. & Bhagwat, A.S. Human activation-induced cytidine deaminase causes transcription-dependent, strand-biased C to U deaminations. Nucleic Acids Res. 31, 2990–2994 (2003).
Petersen-Mahrt, S.K., Harris, R.S. & Neuberger, M.S. AID mutates E. coli suggesting a DNA deamination mechanism for antibody diversification. Nature 418, 99–103 (2002).
Di Noia, J. & Neuberger, M.S. Altering the pathway of immunoglobulin hypermutation by inhibiting uracil-DNA glycosylase. Nature 419, 43–48 (2002).
Rada, C. et al. Immunoglobulin isotype switching is inhibited and somatic hypermutation perturbed in UNG-deficient mice. Curr. Biol. 12, 1748–1755 (2002).
Imai, K. et al. Human uracil-DNA glycosylase deficiency associated with profoundly impaired immunoglobulin class-switch recombination. Nat. Immunol. 4, 1023–1028 (2003).
Ta, V.T. et al. AID mutant analyses indicate requirement for class-switch-specific cofactors. Nat. Immunol. 4, 843–848 (2003).
Barreto, V., Reina-San-Martin, B., Ramiro, A.R., McBride, K.M. & Nussenzweig, M. C-terminal deletion of AID uncouples class switch recombination from somatic hypermutation and gene conversion. Mol. Cell 12, 501–508 (2003).
Fagarasan, S., Kinoshita, K., Muramatsu, M., Ikuta, K. & Honjo, T. In situ class switching and differentiation to IgA-producing cells in the gut lamina propria. Nature 413, 639–643 (2001).
Nagaoka, H., Muramatsu, M., Yamamura, N., Kinoshita, K. & Honjo, T. Activation-induced deaminase (AID)-directed hypermutation in the immunoglobulin Sμ region: implication of AID involvement in a common step of class switch recombination and somatic hypermutation. J. Exp. Med. 195, 529–534 (2002).
Ito, S. et al. Activation-induced cytidine deaminase shuttles between nucleus and cytoplasm like apolipoprotein B mRNA editing catalytic polypeptide 1. Proc. Natl. Acad. Sci. USA 101, 1975–1980 (2004).
Woo, C.J., Martin, A. & Scharff, M.D. Induction of somatic hypermutation is associated with modifications in immunoglobulin variable region chromatin. Immunity 19, 479–489 (2003).
Chester, A. et al. The apolipoprotein B mRNA editing complex performs a multifunctional cycle and suppresses nonsense-mediated decay. EMBO J. 22, 3971–3982 (2003).
Kinoshita, K., Harigai, M., Fagarasan, S., Muramatsu, M. & Honjo, T. A hallmark of active class switch recombination: Transcript directed by I promoters on looped-out circular DNAs. Proc. Natl. Acad. Sci. USA 98, 12620–12633 (2001).
Acknowledgements
We thank J.C. Weill and C.A. Reynaud for providing BL2 cells; S. Fagarasan for critical comments and reading the manuscript; E. Inoue, Y. Sasaki and Y. Hosoe for technical assistance; and Y. Shiraki for preparation of the manuscript. Supported by Ministry of Education, Science, Sports, and Culture of Japan (Center of Excellence Grant).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Shinkura, R., Ito, S., Begum, N. et al. Separate domains of AID are required for somatic hypermutation and class-switch recombination. Nat Immunol 5, 707–712 (2004). https://doi.org/10.1038/ni1086
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ni1086
This article is cited by
-
A Novel AICDA Splice-Site Mutation in Two Siblings with HIGM2 Permits Somatic Hypermutation but Abrogates Mutational Targeting
Journal of Clinical Immunology (2022)
-
Diversity of Immunoglobulin (Ig) Isotypes and the Role of Activation-Induced Cytidine Deaminase (AID) in Fish
Molecular Biotechnology (2018)
-
Immuno-engineered organoids for regulating the kinetics of B-cell development and antibody production
Nature Protocols (2017)
-
Mutations in human AID differentially affect its ability to deaminate cytidine and 5-methylcytidine in ssDNA substrates in vitro
Scientific Reports (2017)
-
Targeted AID-mediated mutagenesis (TAM) enables efficient genomic diversification in mammalian cells
Nature Methods (2016)