Abstract
Therapies that boost the anti-tumor responses of cytotoxic T lymphocytes (CTLs) have shown promise; however, clinical responses to the immunotherapeutic agents currently available vary considerably, and the molecular basis of this is unclear. We performed transcriptomic profiling of tumor-infiltrating CTLs from treatment-naive patients with lung cancer to define the molecular features associated with the robustness of anti-tumor immune responses. We observed considerable heterogeneity in the expression of molecules associated with activation of the T cell antigen receptor (TCR) and of immunological-checkpoint molecules such as 4-1BB, PD-1 and TIM-3. Tumors with a high density of CTLs showed enrichment for transcripts linked to tissue-resident memory cells (TRM cells), such as CD103, and CTLs from CD103hi tumors displayed features of enhanced cytotoxicity. A greater density of TRM cells in tumors was predictive of a better survival outcome in lung cancer, and this effect was independent of that conferred by CTL density. Here we define the 'molecular fingerprint' of tumor-infiltrating CTLs and identify potentially new targets for immunotherapy.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Accession codes
References
Pardoll, D.M. The blockade of immune checkpoints in cancer immunotherapy. Nat. Rev. Cancer 12, 252–264 (2012).
Drake, C.G., Lipson, E.J. & Brahmer, J.R. Breathing new life into immunotherapy: review of melanoma, lung and kidney cancer. Nat. Rev. Clin. Oncol. 11, 24–37 (2014).
Garon, E.B. et al. Pembrolizumab for the treatment of non-small-cell lung cancer. N. Engl. J. Med. 372, 2018–2028 (2015).
Sharma, P. & Allison, J.P. Immune checkpoint targeting in cancer therapy: toward combination strategies with curative potential. Cell 161, 205–214 (2015).
Tumeh, P.C. et al. PD-1 blockade induces responses by inhibiting adaptive immune resistance. Nature 515, 568–571 (2014).
Galon, J. et al. Type, density, and location of immune cells within human colorectal tumors predict clinical outcome. Science 313, 1960–1964 (2006).
Yang, Y. Cancer immunotherapy: harnessing the immune system to battle cancer. J. Clin. Invest. 125, 3335–3337 (2015).
Baitsch, L. et al. Exhaustion of tumor-specific CD8+ T cells in metastases from melanoma patients. J. Clin. Invest. 121, 2350–2360 (2011).
Bindea, G. et al. Spatiotemporal dynamics of intratumoral immune cells reveal the immune landscape in human cancer. Immunity 39, 782–795 (2013).
Collins, N. et al. Transcriptional hallmarks of tumor infiltrating lymphocyte responses to melanoma. Blood 122, 3491 (2013).
Tirosh, I. et al. Dissecting the multicellular ecosystem of metastatic melanoma by single-cell RNA-seq. Science 352, 189–196 (2016).
Pauken, K.E. & Wherry, E.J. Overcoming T cell exhaustion in infection and cancer. Trends Immunol. 36, 265–276 (2015).
Brown, I.E., Blank, C., Kline, J., Kacha, A.K. & Gajewski, T.F. Homeostatic proliferation as an isolated variable reverses CD8+ T cell anergy and promotes tumor rejection. J. Immunol. 177, 4521–4529 (2006).
Montes, C.L. et al. Tumor-induced senescent T cells with suppressor function: a potential form of tumor immune evasion. Cancer Res. 68, 870–879 (2008).
Johnston, R.J. et al. The immunoreceptor TIGIT regulates antitumor and antiviral CD8+ T cell effector function. Cancer Cell 26, 923–937 (2014).
Curran, M.A. et al. Systemic 4-1BB activation induces a novel T cell phenotype driven by high expression of Eomesodermin. J. Exp. Med. 210, 743–755 (2013).
Willoughby, J.E. et al. Differential impact of CD27 and 4-1BB costimulation on effector and memory CD8 T cell generation following peptide immunization. J. Immunol. 193, 244–251 (2014).
Topalian, S.L., Drake, C.G. & Pardoll, D.M. Immune checkpoint blockade: a common denominator approach to cancer therapy. Cancer Cell 27, 450–461 (2015).
Gros, A. et al. PD-1 identifies the patient-specific CD8+ tumor-reactive repertoire infiltrating human tumors. J. Clin. Invest. 124, 2246–2259 (2014).
Fourcade, J. et al. Upregulation of Tim-3 and PD-1 expression is associated with tumor antigen-specific CD8+ T cell dysfunction in melanoma patients. J. Exp. Med. 207, 2175–2186 (2010).
Wolfl, M. et al. Activation-induced expression of CD137 permits detection, isolation, and expansion of the full repertoire of CD8+ T cells responding to antigen without requiring knowledge of epitope specificities. Blood 110, 201–210 (2007).
Wherry, E.J. et al. Molecular signature of CD8+ T cell exhaustion during chronic viral infection. Immunity 27, 670–684 (2007).
Mackay, L.K. et al. The developmental pathway for CD103+CD8+ tissue-resident memory T cells of skin. Nat. Immunol. 14, 1294–1301 (2013).
Mueller, S.N., Gebhardt, T., Carbone, F.R. & Heath, W.R. Memory T cell subsets, migration patterns, and tissue residence. Annu. Rev. Immunol. 31, 137–161 (2013).
Mackay, L.K. et al. Hobit and Blimp1 instruct a universal transcriptional program of tissue residency in lymphocytes. Science 352, 459–463 (2016).
Sathaliyawala, T. et al. Distribution and compartmentalization of human circulating and tissue-resident memory T cell subsets. Immunity 38, 187–197 (2013).
Lee, L.N. et al. CXCR6 is a marker for protective antigen-specific cells in the lungs after intranasal immunization against Mycobacterium tuberculosis. Infect. Immun. 79, 3328–3337 (2011).
Tse, S.W., Radtke, A.J., Espinosa, D.A., Cockburn, I.A. & Zavala, F. The chemokine receptor CXCR6 is required for the maintenance of liver memory CD8+ T cells specific for infectious pathogens. J. Infect. Dis. 210, 1508–1516 (2014).
Skon, C.N. et al. Transcriptional downregulation of S1pr1 is required for the establishment of resident memory CD8+ T cells. Nat. Immunol. 14, 1285–1293 (2013).
Ariotti, S. et al. T cell memory. Skin-resident memory CD8+ T cells trigger a state of tissue-wide pathogen alert. Science 346, 101–105 (2014).
Schenkel, J.M. et al. T cell memory. Resident memory CD8 T cells trigger protective innate and adaptive immune responses. Science 346, 98–101 (2014).
Best, J.A. et al. Transcriptional insights into the CD8+ T cell response to infection and memory T cell formation. Nat. Immunol. 14, 404–412 (2013).
Bono, M.R., Fernández, D., Flores-Santibáñez, F., Rosemblatt, M. & Sauma, D. CD73 and CD39 ectonucleotidases in T cell differentiation: beyond immunosuppression. FEBS Lett. 589, 3454–3460 (2015).
Kuhny, M., Hochdörfer, T., Ayata, C.K., Idzko, M. & Huber, M. CD39 is a negative regulator of P2X7-mediated inflammatory cell death in mast cells. Cell Comm. Signal. 12, 40 (2014).
Ohta, A. A metabolic immune checkpoint: adenosine in tumor microenvironment. Front. Immunol. 7, 109 (2016).
Gupta, P.K. et al. CD39 expression identifies terminally exhausted CD8+ T cells. PLoS Pathog. 11, e1005177 (2015).
Sandoval-Montes, C. & Santos-Argumedo, L. CD38 is expressed selectively during the activation of a subset of mature T cells with reduced proliferation but improved potential to produce cytokines. J. Leukoc. Biol. 77, 513–521 (2005).
Muñoz, P. et al. Antigen-induced clustering of surface CD38 and recruitment of intracellular CD38 to the immunologic synapse. Blood 111, 3653–3664 (2008).
Hombrink, P. et al. Programs for the persistence, vigilance and control of human CD8+ lung-resident memory T cells. Nat. Immunol. 17, 1467–1478 (2016).
Faure, M. & Long, E.O. KIR2DL4 (CD158d), an NK cell-activating receptor with inhibitory potential. J. Immunol. 168, 6208–6214 (2002).
Rajagopalan, S. & Long, E.O. KIR2DL4 (CD158d): an activation receptor for HLA-G. Front. Immunol. 3, 258 (2012).
Wiśniewski, A. et al. Genetic polymorphisms and expression of HLA-G and its receptors, KIR2DL4 and LILRB1, in non-small cell lung cancer. Tissue Antigens 85, 466–475 (2015).
Brooke, G., Holbrook, J.D., Brown, M.H. & Barclay, A.N. Human lymphocytes interact directly with CD47 through a novel member of the signal regulatory protein (SIRP) family. J. Immunol. 173, 2562–2570 (2004).
Piccio, L. et al. Adhesion of human T cells to antigen-presenting cells through SIRPbeta2-CD47 interaction costimulates T-cell proliferation. Blood 105, 2421–2427 (2005).
Wang, Z.Q. et al. CD103 and intratumoral immune response in breast cancer. Clinical Cancer Res. 22, 6290–6297 (2016).
Webb, J.R., Milne, K. & Nelson, B.H. Location, location, location: CD103 demarcates intraepithelial, prognostically favorable CD8+ tumor-infiltrating lymphocytes in ovarian cancer. OncoImmunology 3, e27668 (2014).
Djenidi, F. et al. CD8+CD103+ tumor-infiltrating lymphocytes are tumor-specific tissue-resident memory T cells and a prognostic factor for survival in lung cancer patients. J. Immunol. 194, 3475–3486 (2015).
Kurachi, M. et al. The transcription factor BATF operates as an essential differentiation checkpoint in early effector CD8+ T cells. Nat. Immunol. 15, 373–383 (2014).
Xin, G. et al. A critical role of IL-21-induced BATF in sustaining CD8-T-cell-mediated chronic viral control. Cell Rep. 13, 1118–1124 (2015).
Wolkers, M.C. et al. Nab2 regulates secondary CD8+ T-cell responses through control of TRAIL expression. Blood 119, 798–804 (2012).
Vijayanand, P. et al. Invariant natural killer T cells in asthma and chronic obstructive pulmonary disease. N. Engl. J. Med. 356, 1410–1422 (2007).
Engel, I. et al. Innate-like functions of natural killer T cell subsets result from highly divergent gene programs. Nat. Immunol. 17, 728–739 (2016).
Marsh, D. et al. Stromal features are predictive of disease mortality in oral cancer patients. J. Pathol. 223, 470–481 (2011).
Ward, M.J. et al. Tumour-infiltrating lymphocytes predict for outcome in HPV-positive oropharyngeal cancer. Br. J. Cancer 110, 489–500 (2014).
Seumois, G. et al. Transcriptional profiling of Th2 cells identifies pathogenic features associated with asthma. J. Immunol. 197, 655–664 (2016).
Picelli, S. et al. Full-length RNA-seq from single cells using Smart-seq2. Nat. Protoc. 9, 171–181 (2014).
Trapnell, C., Pachter, L. & Salzberg, S.L. TopHat: discovering splice junctions with RNA-Seq. Bioinformatics 25, 1105–1111 (2009).
Love, M.I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Bolotin, D.A. et al. MiXCR: software for comprehensive adaptive immunity profiling. Nat. Methods 12, 380–381 (2015).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. USA 102, 15545–15550 (2005).
Safford, M. et al. Egr-2 and Egr-3 are negative regulators of T cell activation. Nat. Immunol. 6, 472–480 (2005).
Fridman, A.L. & Tainsky, M.A. Critical pathways in cellular senescence and immortalization revealed by gene expression profiling. Oncogene 27, 5975–5987 (2008).
Acknowledgements
We thank M. Chamberlain, K. Amer, C. Fixmer and B. Johnson for assistance with recruitment of study subjects and processing of samples; A. Easton for help with the assignment of scores to TILs; Z. Fu and J. Greenbaum for help with the processing and analysis of sequencing data; and H. Cheroutre, M. Kronenberg and J. Moore for reviewing the manuscript and providing insight. Supported by the Wessex Clinical Research Network and the National Institute of Health Research, UK (sample collection), Cancer Research UK (O.W., E.V.K., C.H.O.; and C11512/A20256, for CD103 pathology analysis), the William K. Bowes Jr Foundation (P.V. and A.-P.G.) and the Faculty of Medicine of the University of Southampton (P.V., T.S.-E. and C.H.O.).
Author information
Authors and Affiliations
Contributions
A.-P.G., P.S.F., T.S.-E., P.V. and C.H.O. conceived of the work and designed, performed and analyzed experiments; A.-P.G., D.S-C. and D.S. performed micro-scaled RNA-Seq experiments and analysis under the supervision of G.S., P.V. and C.H.O.; A.-P.G. performed data analysis and wrote the manuscript under the supervision of P.V. and C.H.O.; J.C., O.W., E.M.G.-M. and T.M. performed the cell isolations and immunohistochemistry data analysis under the supervision of G.J.T. and C.H.O.; and S.J.C., A.A., E.W. and E.V.K. assisted in patient recruitment, obtaining consent and sample collection.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 RNA-Seq analysis of CD8+ TILs and N-TILs.
(a) Schematic representation of the tumor and cell types used for the study. HNSCC, head and neck squamous cell cancer; NSCLC, non-small cell lung cancer; TIL, tumor-infiltrating lymphocyte; N-TIL, non-tumor-infiltrating lymphocyte. (b) ‘Minus-average’ (MA) plots illustrate differentially expressed genes (red dots) for pairwise comparison of lung CD8+ N-TILs versus NSCLC CD8+ TILs (left), comparison of CD8+ TILs from NSCLC adenocarcinoma versus NSCLC squamous carcinoma (center), comparison of CD8+ TILs from HNSCC HPV-positive versus HPV-negative tumors (right) (DESeq2 analysis, Benjamini-Hochberg adjusted P < 0.05) (Online Methods and Supplementary Table 3). (c) Unsupervised hierarchical clustering of the transcriptomes of all CD8+ N-TILs (grey) and CD8+ TILs from NSCLC adenocarcinoma (red) and squamous carcinoma (pink), HPV-negative (light blue) and HPV-positive (dark blue) HNSCC, based on the expression of genes with the highest variance (n = 1000); each line represents an independent sample. (d) t-SNE plots of CD8+ T cell core transcriptome (symbol) from N-TILs and TILs of indicated cancer subtypes. Color scheme is identical to (c). Data are from one experiment (b-d).
Supplementary Figure 2 Heterogeneity in the expression of molecules that are targets of immunotherapy.
RNA-Seq analysis (row-wise normalized counts; bottom key) of various transcripts (left margin; one per row) in CD8+ TIL from patients with HNSCC (one per column); above, CD8+ TIL density (top row; top left key) and tumor stage (bottom row; far right key) for each patient. Data are from one experiment.
Supplementary Figure 3 Correlation of the abundance of PDCD1 and 4-1BB transcripts with clinical and pathological characteristics of patients with NSCLC.
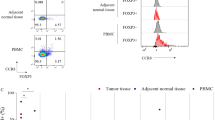
(a) Correlation of PDCD1 transcript expression (log2 normalized counts) in NSCLC CD8+ TIL and the average number of tumor-infiltrating PD-1+ cells (quantified by immunohistochemistry). Each symbol represents an individual patient (n = 10) (r value and P value as in Fig. 3c,d). (b) Flow-cytometry analysis of the expression of PD-1 versus that of CD8 in live and singlet-gated CD45+CD3+ T cells obtained from matched PBMC, lung N-TILs and NSCLC TILs (above plots) from the same patient. (c) Correlation of age of NSCLC patients with PDCD1 or 4-1BB transcript expression (log2 normalized counts) in CD8+ TILs (left) (r value as in Fig. 3c,d). Bar graphs show PDCD1 or 4-1BB transcript expression in CD8+ TILs from patient groups categorized based on gender (first), tumor histology (second), tumor stage (third), performance status (fourth) and smoking status (fifth) of NSCLC patients; adenocarcinoma (Adeno), squamous carcinoma (Sq). Each symbol represents an individual patient (a,c); small horizontal lines are the mean (± s.e.m.). P value insignificant (Kolmogorov-Smirnov test). Data are from ten experiments (a) or one experiment (c) or representative of six experiments (b).
Supplementary Figure 4 TIL status of patients with NSCLC.
Graph shows the average number of CD8α+ cells per high power field (HPF) in tumor samples from each NSCLC patient (Online Methods).
Supplementary Figure 5 Pathways for which CD8+ TILs from NSCLC TILhi tumors show enrichment, and phenotype of CD8+CD103+ TILs.
(a) Ingenuity pathway analysis of genes downregulated in CD8+ TILs from NSCLC TILhi tumors relative to their expression in TILlo tumors (blue), encoding molecules associated with tissue egress (shape indicates function (key)). (b) Flow-cytometry analysis of the expression of CD69, CD49a, KLRG1, CD62L or CCR7 versus that of CD103 in live and singlet-gated CD45+CD3+CD8+ T cells obtained from NSCLC TILs (left); frequency of CD103+CD8+ or CD103−CD8+ TILs (n = 6) that express the indicated surface marker (right). * P = 0.0025 (CD69), P = 0.0025 (CD49a), P = 0.0016 (KLRG1), P = 0.0021 (CD62L) (paired Student’s two-tailed t-test). (c) Analysis of canonical pathways from the Ingenuity pathway analysis database (horizontal axis; bars in plot) for which CD8+ TILs from NSCLC TILhi tumors show enrichment (presented as in Fig. 2a) relative to their expression in TILlo tumors (P values as in Fig. 2a). Each symbol (b) represents an individual sample; small horizontal lines indicate the mean (± s.e.m.). Data are from one experiment (a,c) or from six experiments (b).
Supplementary Figure 6 CD103 (ITGAE) status of CD8+ TILs.
(a) Correlation of ITGAE transcript expression (log2 normalized counts) in NSCLC CD8+ TIL and the average number of tumor-infiltrating CD103+ cells (quantified by immunohistochemistry). Each symbol represents an individual patient (n = 10) (r value and P value as in Fig. 3c,d). (b) Classification of tumors on the basis of the expression of ITGAE (CD103) transcripts (normalized counts) in CD8+ TILs from each NSCLC patient (Online Methods). Data are from ten experiments (a) or one experiment (b).
Supplementary Figure 7 Density of CD103+ cells is predictive of survival in lung cancer.
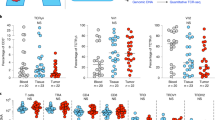
a) Correlation of GZMB transcript expression (log2 normalized counts) in NSCLC CD8+ TIL and the average number of tumor-infiltrating GZMB+ cells (quantified by immunohistochemistry, n = 10) (r value and P value as in Fig. 3c,d). (b) Immunohistochemistry microscopy of cytokeratin, CD103, CD8α, PD-1 and GZMB (above images) in CD103lo and CD103hi NSCLC tumors (left margin). Scale bars, 100 μm. (c) Frequency of CD8+TILs expressing the indicated molecules that are either CD103+ or CD103- (n = 9-19). * P = 0.003 for GZMA and P = 0.0029 for GZMB (paired Student’s two-tailed t-test). (d) Frequency of CD103+CD8+ and CD103−CD8+ TILs (n = 9-19) that express the indicated molecules. * P = 0.047 (paired Student’s two-tailed t-test). (e) Expression (gMFI) of granzyme A or perforin in CD8+ TILs from CD103lo tumors (n = 3-5) or CD103hi (n = 7-8) tumors. NS, P value insignificant (Mann-Whitney test). (f) Survival of patients with lung adenocarcinoma from the TCGA dataset, classified on the basis of expression of ITGAE transcripts into CD103hi tumors (upper 10th percentile, n = 49) or CD103lo tumors (lower 10th percentile, n = 49). P = 0.0148 (log-rank test). Each symbol represents an individual patient (a,e) or sample (c,d); small horizontal line indicates the mean (± s.e.m.). Data are from ten experiments (a) or nine to nineteen experiments (c,d) or thirteen experiments (e) or are representative of 10 experiments (b).
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7 (PDF 1604 kb)
Supplementary Table 1
Demographic, clinical and histopathological characteristics of cancer patients (XLSX 50 kb)
Supplementary Table 2
Details of libraries run for RNA sequencing (XLSX 71 kb)
Supplementary Table 3
List of differentially expressed genes in CD8+ TILs from NSCLC (XLSX 1333 kb)
Supplementary Table 4
Pathway analysis of differentially expressed genes (DEGs) in CD8+ TILs from NSCLC (XLS 40 kb)
Supplementary Table 5
Analysis of TCR beta chain sequences from RNA-Seq data of CD8+ N-TIL versus NSCLC CD8+ TILs (XLSX 43 kb)
Supplementary Table 6
List of differentially expressed genes in NSCLC CD8+ TILs from TILhi versus TILlo tumors. (XLSX 1449 kb)
Supplementary Table 7
List of differentially expressed genes in NSCLC CD8+ TILs from CD103hi versus CD103lo tumors (XLSX 1511 kb)
Supplementary Table 8
Pathway analysis of differentially expressed genes in CD103hi TILs from NSCLC (XLS 51 kb)
Supplementary Table 9
Disease-specific survival in NSCLC patients based on CD8α and CD103 density in tumor tissue (XLSX 75 kb)
Supplementary Table 10
Gene lists utilized for GSEA analysis (XLSX 23 kb)
Rights and permissions
About this article
Cite this article
Ganesan, AP., Clarke, J., Wood, O. et al. Tissue-resident memory features are linked to the magnitude of cytotoxic T cell responses in human lung cancer. Nat Immunol 18, 940–950 (2017). https://doi.org/10.1038/ni.3775
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ni.3775
This article is cited by
-
Tissue-resident memory T cells: decoding intra-organ diversity with a gut perspective
Inflammation and Regeneration (2024)
-
ImmunoPET provides a novel way to visualize the CD103+ tissue-resident memory T cell to predict the response of immune checkpoint inhibitors
EJNMMI Research (2024)
-
Spatial features of specific CD103+CD8+ tissue-resident memory T cell subsets define the prognosis in patients with non-small cell lung cancer
Journal of Translational Medicine (2024)
-
Transient synaptic CD61 pairing with CD103 increases the cytotoxicity of antigen-specific T cells
Nature Immunology (2024)
-
Cellular metabolism regulates the differentiation and function of T-cell subsets
Cellular & Molecular Immunology (2024)