Abstract
Engineered crystallizable fragment (Fc) regions of antibody domains, which assume a unique and unprecedented asymmetric structure within the homodimeric Fc polypeptide, enable completely selective binding to the complement component C1q and activation of complement via the classical pathway without any concomitant engagement of the Fcγ receptor (FcγR). We used the engineered Fc domains to demonstrate in vitro and in mouse models that for therapeutic antibodies, complement-dependent cell-mediated cytotoxicity (CDCC) and complement-dependent cell-mediated phagocytosis (CDCP) by immunological effector molecules mediated the clearance of target cells with kinetics and efficacy comparable to those of the FcγR-dependent effector functions that are much better studied, while they circumvented certain adverse reactions associated with FcγR engagement. Collectively, our data highlight the importance of CDCC and CDCP in monoclonal-antibody function and provide an experimental approach for delineating the effect of complement-dependent effector-cell engagement in various therapeutic settings.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Change history
27 June 2017
In the version of this article initially published online, the labels identifying each plot in Figure 1b were missing. The labels are as follows (left to right): CHO, FcγRI, FcγRIIaR131, FcγRIIaH131, FcγRIIb, FcγRIIIaF158 and FcγRIIIaV158. Also, the reference cited in the accompanying legend (ref. 21) is incorrect. The correct reference is ref. 14. The errors have been corrected in the print, PDF and HTML versions of this article.
References
Weiner, G.J. Building better monoclonal antibody-based therapeutics. Nat. Rev. Cancer 15, 361–370 (2015).
Nimmerjahn, F. & Ravetch, J.V. Translating basic mechanisms of IgG effector activity into next generation cancer therapies. Cancer Immun. 12, 13 (2012).
Pincetic, A. et al. Type I and type II Fc receptors regulate innate and adaptive immunity. Nat. Immunol. 15, 707–716 (2014).
Li, F.J. et al. Emerging roles for the FCRL family members in lymphocyte biology and disease. Curr. Top. Microbiol. Immunol. 382, 29–50 (2014).
van de Donk, N.W.C.J. et al. Monoclonal antibodies targeting CD38 in hematological malignancies and beyond. Immunol. Rev. 270, 95–112 (2016).
Ricklin, D., Hajishengallis, G., Yang, K. & Lambris, J.D. Complement: a key system for immune surveillance and homeostasis. Nat. Immunol. 11, 785–797 (2010).
Dunkelberger, J.R. & Song, W.-C. Complement and its role in innate and adaptive immune responses. Cell Res. 20, 34–50 (2010).
Hess, C. & Kemper, C. Complement-mediated regulation of metabolism and basic cellular processes. Immunity 45, 240–254 (2016).
Sondermann, P. & Szymkowski, D.E. Harnessing Fc receptor biology in the design of therapeutic antibodies. Curr. Opin. Immunol. 40, 78–87 (2016).
Bruhns, P. & Jönsson, F. Mouse and human FcR effector functions. Immunol. Rev. 268, 25–51 (2015).
Taylor, R.P. & Lindorfer, M.A. The role of complement in mAb-based therapies of cancer. Methods 65, 18–27 (2014).
Di Gaetano, N. et al. Complement activation determines the therapeutic activity of rituximab in vivo. J. Immunol. 171, 1581–1587 (2003).
Bruhns, P. et al. Specificity and affinity of human Fcγ receptors and their polymorphic variants for human IgG subclasses. Blood 113, 3716–3725 (2009).
Lux, A., Yu, X., Scanlan, C.N. & Nimmerjahn, F. Impact of immune complex size and glycosylation on IgG binding to human FcγRs. J. Immunol. 190, 4315–4323 (2013).
Golay, J. & Introna, M. Mechanism of action of therapeutic monoclonal antibodies: promises and pitfalls of in vitro and in vivo assays. Arch. Biochem. Biophys. 526, 146–153 (2012).
Vafa, O. et al. An engineered Fc variant of an IgG eliminates all immune effector functions via structural perturbations. Methods 65, 114–126 (2014).
Arduin, E. et al. Highly reduced binding to high and low affinity mouse Fcγ receptors by L234A/L235A and N297A Fc mutations engineered into mouse IgG2a. Mol. Immunol. 63, 456–463 (2015).
Schreiber, A.D. & Frank, M.M. Role of antibody and complement in the immune clearance and destruction of erythrocytes. II. Molecular nature of IgG and IgM complement-fixing sites and effects of their interaction with serum. J. Clin. Invest. 51, 583–589 (1972).
Ackerman, M. & Nimmerjahn, F. Antibody Fc: Linking Adaptive and Innate Immunity (Academic Press, 2013).
Wang, S.-Y., Racila, E., Taylor, R.P. & Weiner, G.J. NK-cell activation and antibody-dependent cellular cytotoxicity induced by rituximab-coated target cells is inhibited by the C3b component of complement. Blood 111, 1456–1463 (2008).
Harvey, B.R. et al. Anchored periplasmic expression, a versatile technology for the isolation of high-affinity antibodies from Escherichia coli-expressed libraries. Proc. Natl. Acad. Sci. USA 101, 9193–9198 (2004).
Jung, S.T. et al. Effective phagocytosis of low Her2 tumor cell lines with engineered, aglycosylated IgG displaying high FcγRIIa affinity and selectivity. ACS Chem. Biol. 8, 368–375 (2013).
Borrok, M.J., Jung, S.T., Kang, T.H., Monzingo, A.F. & Georgiou, G. Revisiting the role of glycosylation in the structure of human IgG Fc. ACS Chem. Biol. 7, 1596–1602 (2012).
Pawluczkowycz, A.W. et al. Binding of submaximal C1q promotes complement-dependent cytotoxicity (CDC) of B cells opsonized with anti-CD20 mAbs ofatumumab (OFA) or rituximab (RTX): considerably higher levels of CDC are induced by OFA than by RTX. J. Immunol. 183, 749–758 (2009).
Teeling, J.L. et al. Characterization of new human CD20 monoclonal antibodies with potent cytolytic activity against non-Hodgkin lymphomas. Blood 104, 1793–1800 (2004).
Boross, P. & Leusen, J.H.W. Mechanisms of action of CD20 antibodies. Am. J. Cancer Res. 2, 676–690 (2012).
Teeling, J.L. et al. The biological activity of human CD20 monoclonal antibodies is linked to unique epitopes on CD20. J. Immunol. 177, 362–371 (2006).
Diebolder, C.A. et al. Complement is activated by IgG hexamers assembled at the cell surface. Science 343, 1260–1263 (2014).
Beum, P.V. et al. Complement activation on B lymphocytes opsonized with rituximab or ofatumumab produces substantial changes in membrane structure preceding cell lysis. J. Immunol. 181, 822–832 (2008).
Gorter, A. & Meri, S. Immune evasion of tumor cells using membrane-bound complement regulatory proteins. Immunol. Today 20, 576–582 (1999).
Min, X. et al. Expression and regulation of complement receptors by human natural killer cells. Immunobiology 219, 671–679 (2014).
Ramos, O.F., Sármay, G., Klein, E., Yefenof, E. & Gergely, J. Complement-dependent cellular cytotoxicity: lymphoblastoid lines that activate complement component 3 (C3) and express C3 receptors have increased sensitivity to lymphocyte-mediated lysis in the presence of fresh human serum. Proc. Natl. Acad. Sci. USA 82, 5470–5474 (1985).
Romain, G. et al. Antibody Fc engineering improves frequency and promotes kinetic boosting of serial killing mediated by NK cells. Blood 124, 3241–3249 (2014).
Bologna, L. et al. Ofatumumab is more efficient than rituximab in lysing B chronic lymphocytic leukemia cells in whole blood and in combination with chemotherapy. J. Immunol. 190, 231–239 (2013).
DiLillo, D.J. & Ravetch, J.V. Differential Fc-receptor engagement drives an anti-tumor vaccinal effect. Cell 161, 1035–1045 (2015).
Klaus, G.G., Pepys, M.B., Kitajima, K. & Askonas, B.A. Activation of mouse complement by different classes of mouse antibody. Immunology 38, 687–695 (1979).
Neuberger, M.S. & Rajewsky, K. Activation of mouse complement by monoclonal mouse antibodies. Eur. J. Immunol. 11, 1012–1016 (1981).
Bruhns, P. Properties of mouse and human IgG receptors and their contribution to disease models. Blood 119, 5640–5649 (2012).
Lim, S.H. et al. Fc gamma receptor IIb on target B cells promotes rituximab internalization and reduces clinical efficacy. Blood 118, 2530–2540 (2011).
Vaughan, A.T. et al. Activatory and inhibitory Fcγ receptors augment rituximab-mediated internalisation of CD20 independent of signalling via the cytoplasmic domain. J. Biol. Chem. jbc.M114.593806 (2015).
Frank, M., Walker, R.C., Lanzilotta, W.N., Prestegard, J.H. & Barb, A.W. Immunoglobulin G1 Fc domain motions: implications for Fc engineering. J. Mol. Biol. 426, 1799–1811 (2014).
Krissinel, E. & Henrick, K. Inference of macromolecular assemblies from crystalline state. J. Mol. Biol. 372, 774–797 (2007).
Gaboriaud, C. et al. The crystal structure of the globular head of complement protein C1q provides a basis for its versatile recognition properties. J. Biol. Chem. 278, 46974–46982 (2003).
Duncan, A.R. & Winter, G. The binding site for C1q on IgG. Nature 332, 738–740 (1988).
Idusogie, E.E. et al. Mapping of the C1q binding site on rituxan, a chimeric antibody with a human IgG1 Fc. J. Immunol. 164, 4178–4184 (2000).
Schneider, S. & Zacharias, M. Atomic resolution model of the antibody Fc interaction with the complement C1q component. Mol. Immunol. 51, 66–72 (2012).
Krapp, S., Mimura, Y., Jefferis, R., Huber, R. & Sondermann, P. Structural analysis of human IgG-Fc glycoforms reveals a correlation between glycosylation and structural integrity. J. Mol. Biol. 325, 979–989 (2003).
Lu, J. et al. Structure of FcγRI in complex with Fc reveals the importance of glycan recognition for high-affinity IgG binding. Proc. Natl. Acad. Sci. USA 112, 833–838 (2015).
Kennedy, A.D. et al. Rituximab infusion promotes rapid complement depletion and acute CD20 loss in chronic lymphocytic leukemia. J. Immunol. 172, 3280–3288 (2004).
Cartron, G. et al. Therapeutic activity of humanized anti-CD20 monoclonal antibody and polymorphism in IgG Fc receptor FcγRIIIa gene. Blood 99, 754–758 (2002).
Mancardi, D.A. et al. FcγRIV is a mouse IgE receptor that resembles macrophage FcɛRI in humans and promotes IgE-induced lung inflammation. J. Clin. Invest. 118, 3738–3750 (2008).
Kelton, W. et al. IgGA: a “cross-isotype” engineered human Fc antibody domain that displays both IgG-like and IgA-like effector functions. Chem. Biol. 21, 1603–1609 (2014).
Lee, C.-F., Paull, T.T. & Person, M.D. Proteome-wide detection and quantitative analysis of irreversible cysteine oxidation using long column UPLC-pSRM. J. Proteome Res. 12, 4302–4315 (2013).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
Winn, M.D. et al. Overview of the CCP4 suite and current developments. Acta Crystallogr. D Biol. Crystallogr. 67, 235–242 (2011).
McCoy, A.J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Adams, P.D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Emsley, P., Lohkamp, B., Scott, W.G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010).
Brünger, A.T. Free R value: a novel statistical quantity for assessing the accuracy of crystal structures. Nature 355, 472–475 (1992).
Chen, V.B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D Biol. Crystallogr. 66, 12–21 (2010).
Terwilliger, T.C. Maximum-likelihood density modification. Acta Crystallogr. D Biol. Crystallogr. 56, 965–972 (2000).
Liadi, I., Roszik, J., Romain, G., Cooper, L.J.N. & Varadarajan, N. Quantitative high-throughput single-cell cytotoxicity assay for T cells. J. Vis. Exp. 72, e50058 (2013).
Merouane, A. et al. Automated profiling of individual cell-cell interactions from high-throughput time-lapse imaging microscopy in nanowell grids (TIMING). Bioinformatics 31, 3189–3197 (2015).
Acknowledgements
We thank Y. Tanno for assistance with protein expression; A. Bui for assistance with liquid chromatography–tandem mass spectrometry; P. Tucker (University of Texas at Austin) for cancer cell lines; D. Lee (MD Anderson Cancer Center) for patient-derived primary acute lymphocytic leukemia cells; the Macromolecular Crystallography Facility of the University of Texas at Austin; the Berkeley Center for Structural Biology; the Advanced Light Source (supported by the Director, Office of Science, Office of Basic Energy Sciences, of the US Department of Energy under contract DE-AC02-05CH11231); the Proteomics Facility at the University of Texas at Austin (supported by grant RP110782 from the Cancer Prevention Research Training Program); and A. Nicola and the Plate-Forme d'Imagerie Dynamique (Institut Pasteur, Paris) for help with the bioluminescence experiments. Supported by the Clayton Foundation, the Institut Pasteur (P.B. laboratory), the Institut National de la Santé et de la Recherche Médicale (P.B. laboratory), the European Research CouncilSeventh Frame-work Program (ERC-2013-CoG 616050 for the P.B. laboratory), the Pasteur–Paris University International PhD program (B.B.), the Cancer Prevention Research Training Program (RP140108 to M.D.; RP160015 to H.T.; and RP130570 to N.V.), the American Cancer Society (123506-PF-13-354-01-CDD to N.M.), Uehara Memorial Foundation (H.T.), Japan Society for the Promotion of Science (H.T.), Deutsche Forschungsgemeinschaft (CRC1181-A07 to F.N.), the US National Institutes of Health (R01CA174385 to N.V.; and R01 GM104896 to Y.J.Z.) and the Welch Foundation (F-1778 to Y.J.Z.). The content is solely the responsibility of the authors and does not necessarily represent the official views of the Cancer Prevention and Research Institute of Texas.
Author information
Authors and Affiliations
Contributions
C.-H.L. and G.G. conceived of and designed the research; C.-H.L., G.R., W.Y., M.W., W.C., B.T., J.L., K.T., M.D., A.L., N.M., M.A.L., O.R.-L.G., B.B., T.H.K., H.T., G.D. and C.A. performed experiments; C.-H.L, G.R., W.Y., O.I.L., R.P.T., F.N., N.V., P.B., Y.J.Z. and G.G. analyzed data; and C.-H.L, G.R., N.V., P.B., Y.J.Z., and G.G. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
G.G. and C.-H.L. are authors of the approved patent PCT/US2016/017100 9 ('Engineered immunoglobulin Fc polypeptides displaying improved complement activation').
Integrated supplementary information
Supplementary Figure 1 Engineering of C1q-specific IgG antibodies.
(a) Schematic illustration of aglycosylated IgG display system for Fc engineering. Soluble C1q-PE competes with non-fluorescent FcγRs-GST for binding to the displayed IgG variants on the surface of bacterial spheroplasts before FACS sorting. (b) Schematic illustration of the construction of the three sub-libraries of mutated Fc domains: S-library: randomization of the focused 15 amino acids; E-library: random mutagenesis on Fc domain by error prone PCR with 1% error rate: SE-library: random mutagenesis of the S-library gene pool by error prone PCR with 1% error rate. The sizes of sub-libraries were 2 × 108 (S-library), 3 × 108 (SE-library), and 1 × 109 (E-library). (c) C1q does not bind to IgG-displaying E.coli spheroplasts in high salt buffer (50 mM phosphate, 330 mM NaCl, pH 7.4). Since E.coli cannot synthesize glycosylated antibodies, E.coli cells were engineered to display an antigen (PA domain 4) on the inner membrane and then the spheroplasted cells were incubated with the very high affinity anti-PA domain 4 antibody, M18. Binding to C1q to the surface-bound C1q was detected by FACS using 10 nM C1q-PE. (d) Flow cytometry analysis of C1q binding onto IgG-displaying E.coli spheroplasts in high salt buffer. (e) Flow cytometry analysis of E.coli library spheroplasts labeled with 10 nM C1q-PE before (Red) and after sorting (Cyan). (f-g) Fluorescent histogram of isolated IgG variants binding to 10 nM of C1q-PE in high salt phosphate buffer (f) or to 10 nM of tetrameric FcγRIIIaV158-PE in PBS (g). (h) MFI and fold increase in MFI following incubation with C1q or tetrameric FcγRIIIaV158 relative to cells unmutated aglycosylated IgG.
Supplementary Figure 2 LC-MS/MS spectra of 801 Fc.
Purified A801 Fc (a-b) or G801 Fc (c-d) was examined without (a & c) or with (b & d) 160 mM of dithiothreitol (DTT). Molecular weight (M.W.) estimated from the LS-MS data are shown; the respective calculated masses were presented at top of each plot. In (c) and (d) multiple predominant species detected are a consequence of glycan heterogeneity.
Supplementary Figure 3 Binding analysis of RA801 or RA802 to human C1q, FcγRs and FcRn.
(a) ELISA analysis of C1q-specific antibody variants binding to human FcγRs. ELISA plates were coated with aglycosylated IgG1 (negative control), glycosylated IgG1 (Rituxan, positive control), RA801 or RA802. 500 nM or 50 nM of his-tagged FcγRI, GST-tagged FcγRIIaH131, GST-tagged FcγRIIaR131, GST-tagged FcγRIIb, GST-tagged FcγRIIIaV158, or GST-tagged FcγRIIIaF158 was added and binding was detected using anti-His IgG conjugated to HRP or anti-GST IgG-HRP, accordingly. Errors bars: standard deviations from triplicate experiments. (b-c) SPR analysis of C1q-specific antibody variants. (b) SPR analysis of antibody binding to purified C1q or to effector Fcγ receptors. Antibody variants were immobilized on CM5 chips. The binding of C1q, or of GST fusion proteins high affinity FcγRI, or low affinity FcγRIIa, FcγRIIb, or FcγRIIIa, all expressed as dimers, to enhance binding, was assayed. (c) pH-dependent binding to human FcRn. In all sensorgrams, x-axis is time (sec) and y-axis is RU (response unit). Data are from one experiment representative of three experiments (a-c).
Supplementary Figure 4 Complement-mediated tumor-cell killing by C1q-specific antibody variants.
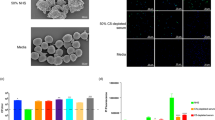
(a) CDC assays with Daudi cells. 10 μg/ml of each antibody was incubated with 50 % of PHS and CD20+ Daudi cells for 15 mins and 30 mins. Lysed cells were detected by TO-PRO-3. (b) C1q deposition on mAb-opsonized CD20+ cells. Time course for C1q deposition on CD20+ Raji cells. Raji cells were incubated in 5% NHS and 10 μg/ml mAb for different time periods at 37°C. Cells were washed twice, incubated with FITC-conjugated anti-C1q and assayed by flow cytometry. MFI were converted to molecules of equivalent soluble fluorochrome (MESF) using calibrated beads (Spherotech). (c-d) CD20+ cancer cells killing activities by PBMC or PMNs. Raji (or Ramos) cells incubated with effector cells in RPMI1640 medium without serum (c) or with 25% of C9-depleted serum (d). (e) Effect of α-CR3 or α-CR4 antibodies in CDCC. Rituximab (or RA801)-opsonized Ramos cells were incubated with: 10 μg/ml of α-CR3 Ab (or α-CR4 Ab)-coated effector cells in RPMI1640 medium supplemented with 25% C9-depleted serum. (f) Cell killing of CD20+ cancer cells by effector cells in the presence of serum. Cell lysis of CD20+ cells with effector cells in RPMI1640 medium supplemented with 25% PHS. In c-f, % of cell lysis was determined 4 hours after the addition of cells and antibody. In all assays, the percent of tumor cell lysis was calculated according to the following formula: 100 × (E-S)/(M-S), where E is the fluorescence of the experimental well, S is the fluorescence in the absence of antibody (tumor cells incubated with medium and complement alone), and M is that of tumor cells with lysis buffer (Triton® X-100 at 2% v/v, SDS at 1% w/v, 100 mM NaCl, and 1 mM EDTA). In all assays, PMN cells were stimulated by incubation with 10 ng/mL GM-CSF and trastuzumab-IgG was used as a negative control. Errors bars indicate the standard deviation (s.d.) from triplicate experiments. Data are from one experiment representative of three experiments.
Supplementary Figure 5 Time-lapse imaging in nanowell grids (TIMING).
Raji tumor cells and NK cells are stained with PKH26 and PKH67 respectively, deposited en masse on a grid containing thousands of nanowells, immersed in fluorescent annexin V-containing medium and imaged for 6 h by high throughput timelapse microscopy. Imaging data were analyzed as described (Merouane Bioinformatics).
Supplementary Figure 6 Mouse complement activation of C1q-specific antibody variants and FcγR-mediated internalization in CD20+FcγRIIb+ cancer cells.
(a) Binding RA801 and RA802 to mouse FcγRs. Microtiter well plates were coated with aglycosylated IgG1 (negative control), glycosylated IgG1 (Rituxan, positive control), mIgG, RA801, or RA802. His-tagged mFcγRs were added and binding was detected using anti-His IgG conjugated to HRP. Errors bars: standard deviations from triplicate experiments. (b) Lack of binding of RA801 immune complexes to mouse FcγRs expressed on CHO cells. The binding activities of performed ICs made of RA801 or Rituxan with PE-conjugated F(ab’)2 goat anti-human IgG F(ab’)2. (c) CDC assays with pooled mouse serum (PMS). CDC assays of CD20+ cells by RA801 with PMS. Experimental conditions for CDC assays of Ramos cells as in Supplementary Fig. 4. Calcein-loaded EL4-hCD20 cells were incubated with 50 % of PMS and serially diluted antibodies for 4 hrs. EC50 values were presented in the plot (Orange: Rituxan, Blue: RA801). (d) α-CD20 antibody internalization in CD20+FcγRIIb+ cancer cells, HBL-1 or TMD8. CD20+FcγRIIb+ cancer cells were incubated with 100 nM of RA801 or RA802 for 0, 2, 4, or 6 h. Surface bound IgG was detected by flow cytometry using FITC conjugated goat anti-human Fc. (Abcam). In all assays, trastuzumab-IgG was used as a negative control and errors bars indicate the s.d. from triplicate experiments. Data are from one experiment representative of three experiments.
Supplementary Figure 7 Structural analysis of C1q-specific Fc variants.
(a) Crystal packing environment for A801-Fc. The crystallographic packing lattice is shown for A801-Fc (PDB ID: 5V43). The dimer from one asymmetric unit is shown in cartoon representation, with Cγ2A in white; Cγ2B in blue; and Cγ3B in pink. Surrounding asymmetric units are shown in ribbon representation (magenta). No contacts or atomic clashes were observed between Cγ2B and molecules from surrounding asymmetric units. (b) B factors distribution fucosylated-WT Fc (PDB ID: 3AVE), G801-Fc (PDB ID: 5V4E), aglycosylated-WT Fc (PDB ID: 3S7G), A801-Fc (PDB ID: 5V43). The coloring of the figure is based on the B factors gradient with lowest (blue) to highest (red). The dash line represents disordered regions. (c) Distance of FcγR-binding motifs of Fc domain to FcγRI (PDB ID: 4X4M). FcγR is shown as a white transparent surface and the secondary structure of the whole complex is shown in ribbon. Two different regions on the Fc are both identified to be important for FcγR and Fc complex formation: LLPP motif (important residue Leu235 is shown in sticks) and C’E loop. (d-e) Overlaid Cγ2 and 'soft' C'E loop of 801-Fc with wild type Fc. (d) Cγ2 of G801-Fc (PDB 5V4E, purple) was superimposed with fucosylated-WT Fc (orange, PDB: 3AVE). (e) G801-Fc (purple) was superimposed with aglycosylated-WT Fc (green, PDB: 3S7G). The 'soft' C'E loop of G801-Fc was highlighted by arrow. The glycan in PDB 3AVE is shown in white.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7 and Supplementary Tables 1–3 (PDF 1541 kb)
Rights and permissions
About this article
Cite this article
Lee, CH., Romain, G., Yan, W. et al. IgG Fc domains that bind C1q but not effector Fcγ receptors delineate the importance of complement-mediated effector functions. Nat Immunol 18, 889–898 (2017). https://doi.org/10.1038/ni.3770
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ni.3770
This article is cited by
-
An engineered Fc fusion protein that targets antigen-specific T cells and autoantibodies mitigates autoimmune disease
Journal of Neuroinflammation (2023)
-
Structural insights into the bi-specific cross-over dual variable antibody architecture by cryo-EM
Scientific Reports (2023)
-
Advances in antibody-based therapy in oncology
Nature Cancer (2023)
-
Unlocking the Power of Complement-Dependent Cytotoxicity: Engineering Strategies for the Development of Potent Therapeutic Antibodies for Cancer Treatments
BioDrugs (2023)
-
Naturally acquired antibodies from Beninese infants promote Plasmodium falciparum merozoite-phagocytosis by human blood leukocytes: implications for control of asymptomatic malaria infections
Malaria Journal (2022)