Abstract
Subsets of innate lymphoid cells (ILCs) reside in the mucosa and regulate immune responses to external pathogens. While ILCs can be phenotypically classified into ILC1, ILC2 and ILC3 subsets, the transcriptional control of commitment to each ILC lineage is incompletely understood. Here we report that the transcription factor Runx3 was essential for the normal development of ILC1 and ILC3 cells but not of ILC2 cells. Runx3 controlled the survival of ILC1 cells but not of ILC3 cells. Runx3 was required for expression of the transcription factor RORγt and its downstream target, the transcription factor AHR, in ILC3 cells. The absence of Runx3 in ILCs exacerbated infection with Citrobacter rodentium. Therefore, our data establish Runx3 as a key transcription factor in the lineage-specific differentiation of ILC1 and ILC3 cells.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Diefenbach, A., Colonna, M. & Koyasu, S. Development, differentiation, and diversity of innate lymphoid cells. Immunity 41, 354–365 (2014).
Eberl, G. et al. An essential function for the nuclear receptor RORγ(t) in the generation of fetal lymphoid tissue inducer cells. Nat. Immunol. 5, 64–73 (2004).
Satoh-Takayama, N. et al. Microbial flora drives interleukin 22 production in intestinal NKp46+ cells that provide innate mucosal immune defense. Immunity 29, 958–970 (2008).
Klose, C.S. et al. A T-bet gradient controls the fate and function of CCR6-RORγt+ innate lymphoid cells. Nature 494, 261–265 (2013).
Klose, C.S. et al. Differentiation of type 1 ILCs from a common progenitor to all helper-like innate lymphoid cell lineages. Cell 157, 340–356 (2014).
Yu, X. et al. The basic leucine zipper transcription factor NFIL3 directs the development of a common innate lymphoid cell precursor. eLife 3, e04406 (2014).
Constantinides, M.G., McDonald, B.D., Verhoef, P.A. & Bendelac, A. A committed precursor to innate lymphoid cells. Nature 508, 397–401 (2014).
Fuchs, A. et al. Intraepithelial type 1 innate lymphoid cells are a unique subset of IL-12- and IL-15-responsive IFN-γ-producing cells. Immunity 38, 769–781 (2013).
Yagi, R. et al. The transcription factor GATA3 is critical for the development of all IL-7Rα-expressing innate lymphoid cells. Immunity 40, 378–388 (2014).
Hoyler, T. et al. The transcription factor GATA-3 controls cell fate and maintenance of type 2 innate lymphoid cells. Immunity 37, 634–648 (2012).
Halim, T.Y. et al. Retinoic-acid-receptor-related orphan nuclear receptor α is required for natural helper cell development and allergic inflammation. Immunity 37, 463–474 (2012).
Qiu, J. et al. The aryl hydrocarbon receptor regulates gut immunity through modulation of innate lymphoid cells. Immunity 36, 92–104 (2012).
Lee, J.S. et al. AHR drives the development of gut ILC22 cells and postnatal lymphoid tissues via pathways dependent on and independent of Notch. Nat. Immunol. 13, 144–151 (2012).
Basu, R., Hatton, R.D. & Weaver, C.T. The Th17 family: flexibility follows function. Immunol. Rev. 252, 89–103 (2013).
Serafini, N. et al. Gata3 drives development of RORγt+ group 3 innate lymphoid cells. J. Exp. Med. 211, 199–208 (2014).
Wong, W.F., Kohu, K., Chiba, T., Sato, T. & Satake, M. Interplay of transcription factors in T-cell differentiation and function: the role of Runx. Immunology 132, 157–164 (2011).
Satpathy, A.T. et al. Runx1 and Cbfbeta regulate the development of Flt3+ dendritic cell progenitors and restrict myeloproliferative disorder. Blood 123, 2968–2977 (2014).
Levanon, D. et al. Transcription factor Runx3 regulates interleukin-15-dependent natural killer cell activation. Mol. Cell. Biol. 34, 1158–1169 (2014).
Tachibana, M. et al. Runx1/Cbfβ2 complexes are required for lymphoid tissue inducer cell differentiation at two developmental stages. J. Immunol. 186, 1450–1457 (2011).
Pozner, A. et al. Transcription-coupled translation control of AML1/RUNX1 is mediated by cap- and internal ribosome entry site-dependent mechanisms. Mol. Cell. Biol. 20, 2297–2307 (2000).
Egawa, T., Tillman, R.E., Naoe, Y., Taniuchi, I. & Littman, D.R. The role of the Runx transcription factors in thymocyte differentiation and in homeostasis of naive T cells. J. Exp. Med. 204, 1945–1957 (2007).
Egawa, T. & Littman, D.R. ThPOK acts late in specification of the helper T cell lineage and suppresses Runx-mediated commitment to the cytotoxic T cell lineage. Nat. Immunol. 9, 1131–1139 (2008).
Sojka, D.K. et al. Tissue-resident natural killer (NK) cells are cell lineages distinct from thymic and conventional splenic NK cells. Elife 3, e01659 (2014).
Rankin, L.C. et al. The transcription factor T-bet is essential for the development of NKp46+ innate lymphocytes via the Notch pathway. Nat. Immunol. 14, 389–395 (2013).
Xi, H., Schwartz, R., Engel, I., Murre, C. & Kersh, G.J. Interplay between RORγt, Egr3, and E proteins controls proliferation in response to pre-TCR signals. Immunity 24, 813–826 (2006).
Lai, C.B. & Mager, D.L. Role of runt-related transcription factor 3 (RUNX3) in transcription regulation of natural cytotoxicity receptor 1 (NCR1/NKp46), an activating natural killer (NK) cell receptor. J. Biol. Chem. 287, 7324–7334 (2012).
Cruz-Guilloty, F. et al. Runx3 and T-box proteins cooperate to establish the transcriptional program of effector CTLs. J. Exp. Med. 206, 51–59 (2009).
Sun, Z. et al. Requirement for RORγ in thymocyte survival and lymphoid organ development. Science 288, 2369–2373 (2000).
Sawa, S. et al. Lineage relationship analysis of RORγt+ innate lymphoid cells. Science 330, 665–669 (2010).
van de Pavert, S.A. et al. Maternal retinoids control type 3 innate lymphoid cells and set the offspring immunity. Nature 508, 123–127 (2014).
Lotem, J. et al. Runx3-mediated transcriptional program in cytotoxic lymphocytes. PLoS ONE 8, e80467 (2013).
Ciofani, M. et al. A validated regulatory network for Th17 cell specification. Cell 151, 289–303 (2012).
Shin, J.H. et al. Modulation of natural killer cell antitumor activity by the aryl hydrocarbon receptor. Proc. Natl. Acad. Sci. USA 110, 12391–12396 (2013).
Cooper, M.A. et al. In vivo evidence for a dependence on interleukin 15 for survival of natural killer cells. Blood 100, 3633–3638 (2002).
Gasteiger, G. & Rudensky, A.Y. Interactions between innate and adaptive lymphocytes. Nat. Rev. Immunol. 14, 631–639 (2014).
Hepworth, M.R. & Sonnenberg, G.F. Regulation of the adaptive immune system by innate lymphoid cells. Curr. Opin. Immunol. 27, 75–82 (2014).
Reis, B.S., Rogoz, A., Costa-Pinto, F.A., Taniuchi, I. & Mucida, D. Mutual expression of the transcription factors Runx3 and ThPOK regulates intestinal CD4+ T cell immunity. Nat. Immunol. 14, 271–280 (2013).
Liu, J.Y. et al. Vav proteins are necessary for correct differentiation of mouse cecal and colonic enterocytes. J. Cell Sci. 122, 324–334 (2009).
Simmons, C.P. et al. Impaired resistance and enhanced pathology during infection with a noninvasive, attaching-effacing enteric bacterial pathogen, Citrobacter rodentium, in mice lacking IL-12 or IFN-gamma. J. Immunol. 168, 1804–1812 (2002).
Kohu, K. et al. The Runx3 transcription factor augments Th1 and down-modulates Th2 phenotypes by interacting with and attenuating GATA3. J. Immunol. 183, 7817–7824 (2009).
Yagi, R. et al. The transcription factor GATA3 actively represses RUNX3 protein-regulated production of interferon-γ. Immunity 32, 507–517 (2010).
Lazarevic, V. et al. T-bet represses TH17 differentiation by preventing Runx1-mediated activation of the gene encoding RORγt. Nat. Immunol. 12, 96–104 (2011).
Djuretic, I.M. et al. Transcription factors T-bet and Runx3 cooperate to activate Ifng and silence Il4 in T helper type 1 cells. Nat. Immunol. 8, 145–153 (2007).
Possot, C. et al. Notch signaling is necessary for adult, but not fetal, development of RORγt+ innate lymphoid cells. Nat. Immunol. 12, 949–958 (2011).
Sanos, S.L. et al. RORγt and commensal microflora are required for the differentiation of mucosal interleukin 22-producing NKp46+ cells. Nat. Immunol. 10, 83–91 (2009).
Cherrier, M., Sawa, S. & Eberl, G. Notch, Id2, and RORγt sequentially orchestrate the fetal development of lymphoid tissue inducer cells. J. Exp. Med. 209, 729–740 (2012).
Brenner, O. et al. Loss of Runx3 function in leukocytes is associated with spontaneously developed colitis and gastric mucosal hyperplasia. Proc. Natl. Acad. Sci. USA 101, 16016–16021 (2004).
Weersma, R.K. et al. Runt-related transcription factor 3 is associated with ulcerative colitis and shows epistasis with solute carrier family 22, members 4 and 5. Inflamm. Bowel Dis. 14, 1615–1622 (2008).
Narni-Mancinelli, E. et al. Fate mapping analysis of lymphoid cells expressing the NKp46 cell surface receptor. Proc. Natl. Acad. Sci. USA 108, 18324–18329 (2011).
Brown, J.B. et al. Epithelial phosphatidylinositol-3-kinase signaling is required for β-catenin activation and host defense against Citrobacter rodentium infection. Infect. Immun. 79, 1863–1872 (2011).
Acknowledgements
We thank E. Vivier (Centre d'Immunologie de Marseille-Luminy) for NKp46-Cre mice; D.R. Littman (New York University) for Cbfbf/f, Runx3f/f and Runx3d+/YFP mice; S. Taffner for technical assistance; H. Miyoshi and A. Fuchs for technical advice; and D.K. Sojka, and S. Demehri for discussions. Supported by the Howard Hughes Medical Institute (W.M.Y.) and the US National Institutes of Health (R01AI097244 to T.Eg.; and T32-GM007200 for M.D.B.).
Author information
Authors and Affiliations
Contributions
T.Eb. and W.M.Y. designed experiments and analyzed data; T.Eb., C.S., S.H.R., B.P.-D., L.Y., M.D.B. and T.S.S. performed the experiments; D.L. and Y.G. supplied reagents; T.Eg. analyzed data obtained by chromatin immunoprecipitation followed by deep sequencing; and T.Eb., Y.G., T.S.S., M.C., T.Eg. and W.M.Y. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Absence of NKp46+ ILC3 cells in PPs and reduction in the number of NK cells in tissues of Cbfbf/fNKp46-Cre mice.
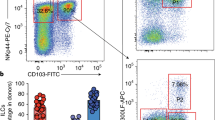
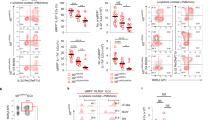
(a) Flow cytometry analyzing NK cells (CD3− CD19− NK1.1+ NKp46+) in liver, skin and salivary gland of Runx3d+/YFP mice for DX5, CD49a and distal Runx3-YFP. Shaded histograms indicate wild-type mice. Upper plots were gated on Live CD45+ CD3− CD19− cells. Arrows in the two upper panels indicate gated cells shown in the lower histogram. Numbers in quadrants and outlined area indicate percent cells in each. (b) Flow cytometry of PP lymphocytes showing ILC3 subsets in Cbfb+/fNKp46-Cre and Cbfbf/fNKp46-Cre mice. Upper plots were gated on Live CD45+ CD3− CD19− cells. (c) Absolute cell numbers of ILC3 subsets in PPs of Cbfb+/fNKp46-Cre and Cbfbf/fNKp46-Cre mice. (d, e) Frequency (d) and absolute cell numbers (e) of indicated NK cells (CD3− CD19− NK1.1+ NKp46+) and their DX5 and CD49a expression (d) in spleen, liver, salivary gland and skin of Cbfb+/fNKp46-Cre and Cbfbf/fNKp46-Cre mice. Upper plots were gated on Live CD45+ CD3− CD19− cells. Arrows in the upper panels indicate gated cells shown in the lower plots (d). Numbers in outlined areas or quadrants indicate percent cells in each (a, b, d). Data are representative of more than three experiments (a, b, d) or from two independent experiments (b and d; mean and s.d. of three mice). *p< 0.05 and **p< 0.01 by Student’s t-test. NS, not significant.
Supplementary Figure 2 Normal differentiation of ILC progenitor cells in Runx3f/fVav1-Cre mice.
Frequency (a, b, c) and absolute number (d) of common lymphoid progenitor cells (CLP: Live CD45+ Lin− cKitlo CD127+ sca1lo Flt3+ α4β7−), α4β7 integrin-expressing CLP (αLP: Live CD45+ Lin− cKitlo CD127+ sca1lo Flt3− α4β7+), common progenitor to all helper-like innate lymphoid cell lineages (CHILP: Live CD45+ Lin− CD127+ α4β7+ Flt3− CD25−), ILC2 precursor (ILC2P: Live CD45+ Lin− CD127+ α4β7+ Flt3− CD25+) and common precursor to ILC (ILCP: Live CD45+ Lin− cKit+ CD127+ α4β7+ PLZF+) in bone marrow of Runx3f/f and Runx3f/fVav1-Cre mice. Plots are gated on Live CD45+ Lin− CD127+ sca1lo cells in a, Live CD45+ Lin− CD127+ cells in b (left plot of each pair), Live CD45+ Lin− CD127+ cells in c. Arrows in the left panels indicate gated cells shown in the right plots. Numbers in quadrants and outlined area indicate percent cells in each. Data are representative of at least two independent experiments (mean and s.d. of three mice in d). NS, not significant.
Supplementary Figure 3 Runx3 is necessary for NK cells and ILC1 cells in tissues.
(a) Flow cytometry of Live CD45+ CD3− Cd19− cells showing NK and ILC1 cells in spleen, intraepithelial lymphocytes (IELs) of small intestine, salivary gland and skin of Runx3+/fNKp46-Cre and Runx3f/fNKp46-Cre mice. (b) Flow cytometry showing frequency of liver NK cells and their DX5 and CD49a expression in Runx3+/fNKp46-Cre and Runx3f/fNKp46-Cre mice. Plots are gated on Live CD45+ CD3− Cd19− cells in a and b (upper). Arrows in the two upper panels indicate gated cells shown in the lower plots in b. (c) Absolute cell numbers of indicated NK cells (CD3− CD19− NK1.1+ NKp46+) and ILC1 cells (CD45+ CD3− CD19− NK1.1+ NKp46+ CD49a+) in Runx3+/fNKp46-Cre and Runx3f/fNKp46-Cre mice. Numbers in outlined areas and quadrants indicate percent cells in each. Data are representative of three experiments (a, b) or from two independent experiments (c; mean and s.d. of three mice). *p < 0.05 and **p < 0.01 by Student’s t-test.
Supplementary Figure 4 CD127+ ILC lineage-negative (ILCLN) cells in RORγt-deficient mice and RORγt expression in Runx3-deficient LTi cells
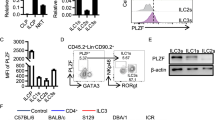
(a) Flow cytometry of CD45+ CD3− CD19− CD127+ NK1.1− LPLs from wild-type (WT) and Rorc(gt)GFP/GFP mice for GATA-3 and RORγt expression as detected by anti-RORγt staining. (Anti-RORγt was necessary for simultaneous staining of GATA-3 in permeabilized cells.) Numbers in outlined areas or quadrants indicate percent cells in each. (b) Flow cytometry (left) for enteric fetal LTi0 (CD45+ CD3− CD11c− CD127+ CD4−) and LTi4 (CD45+ CD3− CD11c− CD127+ CD4+) cells and their RORγt expression (right) in Runx3f/f and Runx3f/fVav1-Cre mice (E17.5). Plots were gated on CD45+ CD3− CD11c− LPLs of intestine. Arrows in the two left panels indicate gated cells shown in the right histogram. (c) Absolute numbers of LTi0 and LTi4 cells in the indicated mice. Data are representative of more than three independent experiments (mean and s.d of 3-5 mice in c). NS, not significant by Student’s t-test.
Supplementary Figure 5 T-bet expression in ILCs is not affected by the absence of Runx3.
(a) Flow cytometry of CD45+ CD3− CD19− (Lin−) CD127+ NK1.1− cells for GATA-3 and RORγt (upper panels) and CD45+ Lin− CD127+ NK1.1− GATA-3int cells for CD4 and NKp46 (lower panels) from large intestinal LPLs of Cbfb+/fNKp46-Cre and Cbfbf/fNKp46-Cre mice. Arrows in the two upper panels indicate gated cells shown in the lower plots. Numbers indicate percent cells in outlined areas or quadrants. (b) Flow cytometry showing T-bet expression in CD45+ Lin− CD127+ NK1.1− GATA-3int NKp46+ or CD4+ cells and CD45+ Lin− NK1.1+ NKp46+ CD127+ (ILC1) or CD127− (NK) cells among LPLs of Cbfb+/fNKp46-Cre and C bfbf/fNKp46-Cre mice. Data are representative of three experiments (a, b).
Supplementary Figure 6 AHR is downstream of RORγt in ILC3 cells.
(a) Expression of indicated genes in CD45+ CD3− CD19− NK1.1− CD127+ LPLs of wild-type (WT) and Rorc(gt)+/GFP mice by qRT-PCR, relative to Actb expression. Data are representative of two independent experiments (mean and s.d. of triplicates). *p < 0.01 by Student’s t-test. (b) Tracks of Runx3 (SRR95607, green) and H3K4me1 (SRR955609, blue) ChIP-seq traces in splenic NK cells, and H3K4me2 (GSM1004811, dark green), p300 (GSM1004849, pink) and RORγt (GSM1004855, green) ChIP-seq traces in Th17 cells.
Supplementary Figure 7 Effect of deficiency in CBF-β or Runx3 on the survival of NK cells in tissues.
(a) Flow cytometry of CD3+ CD19+ cells and indicated NK cells (CD3− CD19− NK1.1+ NKp46+) of Cbfb+/fNKp46-Cre, Cbfbf/fNKp46-Cre, Runx3+/fNKp46-Cre and Runx3f/fNKp46-Cre mice for apoptosis markers. (b) Histograms depicting Ki-67 expression in indicated NK cells of Cbfb+/fNKp46-Cre, Cbfbf/fNKp46-Cre, Runx3+/fNKp46-Cre and Runx3f/fNKp46-Cre mice. Numbers in quadrants indicate percent cells in each. Data are representative of three experiments.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7 (PDF 5973 kb)
Supplementary Table 1
Supplementary Table 1 (XLSX 10 kb)
Supplementary Table 2
Supplementary Table 2 (XLSX 10 kb)
Rights and permissions
About this article
Cite this article
Ebihara, T., Song, C., Ryu, S. et al. Runx3 specifies lineage commitment of innate lymphoid cells. Nat Immunol 16, 1124–1133 (2015). https://doi.org/10.1038/ni.3272
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ni.3272
This article is cited by
-
Proline uptake promotes activation of lymphoid tissue inducer cells to maintain gut homeostasis
Nature Metabolism (2023)
-
RUNX3 inactivates oncogenic MYC through disruption of MYC/MAX complex and subsequent recruitment of GSK3β-FBXW7 cascade
Communications Biology (2023)
-
Regulation of effector and memory CD8 + T cell differentiation: a focus on orphan nuclear receptor NR4A family, transcription factor, and metabolism
Immunologic Research (2023)
-
Whole-genome profiling of DNA methylation and hydroxymethylation identifies distinct regulatory programs among innate lymphocytes
Nature Immunology (2022)
-
Leaving no one behind: tracing every human thymocyte by single-cell RNA-sequencing
Seminars in Immunopathology (2021)