Abstract
Interleukin 37 (IL-37) and IL-1R8 (SIGIRR or TIR8) are anti-inflammatory orphan members of the IL-1 ligand family and IL-1 receptor family, respectively. Here we demonstrate formation and function of the endogenous ligand-receptor complex IL-37–IL-1R8–IL-18Rα. The tripartite complex assembled rapidly on the surface of peripheral blood mononuclear cells upon stimulation with lipopolysaccharide. Silencing of IL-1R8 or IL-18Rα impaired the anti-inflammatory activity of IL-37. Whereas mice with transgenic expression of IL-37 (IL-37tg mice) with intact IL-1R8 were protected from endotoxemia, IL-1R8-deficient IL-37tg mice were not. Proteomic and transcriptomic investigations revealed that IL-37 used IL-1R8 to harness the anti-inflammatory properties of the signaling molecules Mer, PTEN, STAT3 and p62(dok) and to inhibit the kinases Fyn and TAK1 and the transcription factor NF-κB, as well as mitogen-activated protein kinases. Furthermore, IL-37–IL-1R8 exerted a pseudo-starvational effect on the metabolic checkpoint kinase mTOR. IL-37 thus bound to IL-18Rα and exploited IL-1R8 to activate a multifaceted intracellular anti-inflammatory program.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Gay, N.J. & Keith, F.J. Drosophila Toll and IL-1 receptor. Nature 351, 355–356 (1991).
Dinarello, C.A. Immunological and inflammatory functions of the interleukin-1 family. Annu. Rev. Immunol. 27, 519–550 (2009).
Sims, J.E. & Smith, D.E. The IL-1 family: regulators of immunity. Nat. Rev. Immunol. 10, 89–102 (2010).
Thomas, C., Bazan, J.F. & Garcia, K.C. Structure of the activating IL-1 receptor signaling complex. Nat. Struct. Mol. Biol. 19, 455–457 (2012).
O'Neill, L.A. & Bowie, A.G. The family of five: TIR-domain-containing adaptors in Toll-like receptor signalling. Nat. Rev. Immunol. 7, 353–364 (2007).
Boraschi, D. & Tagliabue, A. The interleukin-1 receptor family. Semin. Immunol. 25, 394–407 (2013).
Garlanda, C., Dinarello, C.A. & Mantovani, A. The interleukin-1 family: back to the future. Immunity 39, 1003–1018 (2013).
Thomassen, E., Renshaw, B.R. & Sims, J.E. Identification and characterization of SIGIRR, a molecule representing a novel subtype of the IL-1R superfamily. Cytokine 11, 389–399 (1999).
Qin, J., Qian, Y., Yao, J., Grace, C. & Li, X. SIGIRR inhibits interleukin-1 receptor- and toll-like receptor 4-mediated signaling through different mechanisms. J. Biol. Chem. 280, 25233–25241 (2005).
Wald, D. et al. SIGIRR, a negative regulator of Toll-like receptor-interleukin 1 receptor signaling. Nat. Immunol. 4, 920–927 (2003).
Adib-Conquy, M. et al. Up-regulation of MyD88s and SIGIRR, molecules inhibiting Toll-like receptor signaling, in monocytes from septic patients. Crit. Care Med. 34, 2377–2385 (2006).
Garlanda, C. et al. Intestinal inflammation in mice deficient in Tir8, an inhibitory member of the IL-1 receptor family. Proc. Natl. Acad. Sci. USA 101, 3522–3526 (2004).
Lech, M. et al. Different roles of TiR8/Sigirr on toll-like receptor signaling in intrarenal antigen-presenting cells and tubular epithelial cells. Kidney Int. 72, 182–192 (2007).
Polentarutti, N. et al. Unique pattern of expression and inhibition of IL-1 signaling by the IL-1 receptor family member TIR8/SIGIRR. Eur. Cytokine Netw. 14, 211–218 (2003).
Bufler, P. et al. A complex of the IL-1 homologue IL-1F7b and IL-18-binding protein reduces IL-18 activity. Proc. Natl. Acad. Sci. USA 99, 13723–13728 (2002).
Garlanda, C., Anders, H.J. & Mantovani, A. TIR8/SIGIRR: an IL-1R/TLR family member with regulatory functions in inflammation and T cell polarization. Trends Immunol. 30, 439–446 (2009).
Garlanda, C. et al. Intestinal inflammation in mice deficient in Tir8, an inhibitory member of the IL-1 receptor family. Proc. Natl. Acad. Sci. USA 101, 3522–3526 (2004).
Bekker, L.G. et al. Immunopathologic effects of tumor necrosis factor α in murine mycobacterial infection are dose dependent. Infect. Immun. 68, 6954–6961 (2000).
Bulut, Y., Faure, E., Thomas, L., Equils, O. & Arditi, M. Cooperation of Toll-like receptor 2 and 6 for cellular activation by soluble tuberculosis factor and Borrelia burgdorferi outer surface protein A lipoprotein: role of Toll-interacting protein and IL-1 receptor signaling molecules in Toll-like receptor 2 signaling. J. Immunol. 167, 987–994 (2001).
Garlanda, C. et al. Damping excessive inflammation and tissue damage in Mycobacterium tuberculosis infection by Toll IL-1 receptor 8/single Ig IL-1-related receptor, a negative regulator of IL-1/TLR signaling. J. Immunol. 179, 3119–3125 (2007).
Horne, D.J. et al. Common polymorphisms in the PKP3-SIGIRR-TMEM16J gene region are associated with susceptibility to tuberculosis. J. Infect. Dis. 205, 586–594 (2012).
Batliwalla, F.M. et al. Microarray analyses of peripheral blood cells identifies unique gene expression signature in psoriatic arthritis. Mol. Med. 11, 21–29 (2005).
Nold, M.F. et al. IL-37 is a fundamental inhibitor of innate immunity. Nat. Immunol. 11, 1014–1022 (2010).
McNamee, E.N. et al. Interleukin 37 expression protects mice from colitis. Proc. Natl. Acad. Sci. USA 108, 16711–16716 (2011).
Kumar, S. et al. Interleukin-1F7B (IL-1H4/IL-1F7) is processed by caspase-1 and mature IL-1F7B binds to the IL-18 receptor but does not induce IFN-γ production. Cytokine 18, 61–71 (2002).
Pan, G. et al. IL-1H, an interleukin 1-related protein that binds IL-18 receptor/IL-1Rrp. Cytokine 13, 1–7 (2001).
Lewis, E.C. & Dinarello, C.A. Responses of IL-18- and IL-18 receptor-deficient pancreatic islets with convergence of positive and negative signals for the IL-18 receptor. Proc. Natl. Acad. Sci. USA 103, 16852–16857 (2006).
Nold-Petry, C.A. et al. Increased cytokine production in interleukin-18 receptor α-deficient cells is associated with dysregulation of suppressors of cytokine signaling. J. Biol. Chem. 284, 25900–25911 (2009).
O'Neill, L.A. & Hardie, D.G. Metabolism of inflammation limited by AMPK and pseudo-starvation. Nature 493, 346–355 (2013).
Korb, M. et al. The Innate Immune Database (IIDB). BMC Immunol. 9, 7 (2008).
Ward, P.A. An endogenous factor mediates shock-induced injury. Nat. Med. 19, 1368–1369 (2013).
Drexler, S.K. et al. SIGIRR/TIR-8 is an inhibitor of Toll-like receptor signaling in primary human cells and regulates inflammation in models of rheumatoid arthritis. Arthritis Rheum. 62, 2249–2261 (2010).
Bulau, A.M. et al. Role of caspase-1 in nuclear translocation of IL-37, release of the cytokine, and IL-37 inhibition of innate immune responses. Proc. Natl. Acad. Sci. USA 111, 2650–2655 (2014).
Sharma, S. et al. The IL-1 family member 7b translocates to the nucleus and down-regulates proinflammatory cytokines. J. Immunol. 180, 5477–5482 (2008).
Gulen, M.F. et al. The receptor SIGIRR suppresses Th17 cell proliferation via inhibition of the interleukin-1 receptor pathway and mTOR kinase activation. Immunity 32, 54–66 (2010).
De Paula, M.L., Cui, Q.L., Hossain, S., Antel, J. & Almazan, G. The PTEN inhibitor bisperoxovanadium enhances myelination by amplifying IGF-1 signaling in rat and human oligodendrocyte progenitors. Glia 62, 64–77 (2014).
Heindl, M. et al. Autoimmunity, intestinal lymphoid hyperplasia, and defects in mucosal B-cell homeostasis in patients with PTEN hamartoma tumor syndrome. Gastroenterology 142, 1093–1096 (2012).
Ballak, D.B. et al. IL-37 protects against obesity-induced inflammation and insulin resistance. Nat. Commun. 5, 4711 (2014).
Bozza, S. et al. Lack of Toll IL-1R8 exacerbates Th17 cell responses in fungal infection. J. Immunol. 180, 4022–4031 (2008).
Moretti, S. et al. IL-37 inhibits inflammasome activation and disease severity in murine aspergillosis. PLoS Pathog. 10, e1004462 (2014).
Rothlin, C.V., Ghosh, S., Zuniga, E.I., Oldstone, M.B. & Lemke, G. TAM receptors are pleiotropic inhibitors of the innate immune response. Cell 131, 1124–1136 (2007).
Shinohara, H. et al. Dok-1 and Dok-2 are negative regulators of lipopolysaccharide-induced signaling. J. Exp. Med. 201, 333–339 (2005).
Lee, T.W. et al. Fyn deficiency promotes a preferential increase in subcutaneous adipose tissue mass and decreased visceral adipose tissue inflammation. Diabetes 62, 1537–1546 (2013).
Sakurai, H. Targeting of TAK1 in inflammatory disorders and cancer. Trends Pharmacol. Sci. 33, 522–530 (2012).
Oda, K. & Kitano, H. A comprehensive map of the toll-like receptor signaling network. Mol. Syst. Biol. 2, 0015 (2006).
Gross, O. et al. Syk kinase signalling couples to the Nlrp3 inflammasome for anti-fungal host defence. Nature 459, 433–436 (2009).
Wermeling, F., Anthony, R.M., Brombacher, F. & Ravetch, J.V. Acute inflammation primes myeloid effector cells for anti-inflammatory STAT6 signaling. Proc. Natl. Acad. Sci. USA 110, 13487–13491 (2013).
Nold, M. et al. IL-18BPa:Fc cooperates with immunosuppressive drugs in human whole blood. Biochem. Pharmacol. 66, 505–510 (2003).
Nold, M.F. et al. Endogenous IL-32 controls cytokine and HIV-1 production. J. Immunol. 181, 557–565 (2008).
Petry, C., Huwiler, A., Eberhardt, W., Kaszkin, M. & Pfeilschifter, J. Hypoxia increases group IIA phospholipase A2 expression under inflammatory conditions in rat renal mesangial cells. J. Am. Soc. Nephrol. 16, 2897–2905 (2005).
Gersting, S.W., Lotz-Havla, A.S. & Muntau, A.C. Bioluminescence resonance energy transfer: an emerging tool for the detection of protein-protein interaction in living cells. Methods Mol. Biol. 815, 253–263 (2012).
Acknowledgements
We thank J. Gould and L. D'Andrea for technical support; S.-H. Kim for scientific insights; and C. Nowell (Monash Institute of Pharmaceutical Science) for access to and assisting with the use of the Leica GSD microscope. Supported by the National Health and Medical Research Council (1012353 and 1043845 to C.A.N.-P. and M.F.N.; 1022144 to MPG), the National Heart Foundation of Australia (100480 to C.A.N.-P.), Monash University (Larkins Fellowship to M.F.N.), MIMR-PHI (Star Recruitment Fellowship to M.F.N.), Australian Research Council (DP110103616 to N.E.M.), the US National Institutes of Health (AI-15614, AR-45584 to C.A.D.; core facilities of CA-04 6934 to C.A.D.), Deutsche Forschungsgemeinschaft (Bu 1222/3-3 to P.B.), the American Heart Association (12Post12030134 to S.L.) and the Victorian Government's Operational Infrastructure Support Program.
Author information
Authors and Affiliations
Contributions
C.A.N.-P., C.Y.L., S.L., C.A.D. and M.F.N., study design; C.A.N.-P., C.Y.L., I.R., K.D.E., A.S.L.-H., S.W.G., S.X.C., J.C.L., A.M.E., B.R., S.L., T.A., N.E.M., P.B. and M.F.N., experimental work; C.A.N.-P., C.Y.L., I.R., K.D.E., M.P.G., A.S.L.-H., S.W.G., S.X.C., J.C.L., A.M.E., B.R., N.E.M., F.J.R., J.C.W., P.B., C.A.D. and M.F.N., data analysis; C.A.N.-P., C.Y.L., C.A.D. and M.F.N., writing of manuscript; S.L., C.G. and A.M., reagent contribution.
Corresponding author
Ethics declarations
Competing interests
B.R. is an employee of GenXPro.
Integrated supplementary information
Supplementary Figure 1 Additional lung and spleen cytokines in endotoxemia and flow cytometry of IL-1R8 and IL-18Rα in PBMCs.
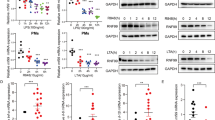
(a, b) Thirty-five wild-type mice (WT; 14 vehicle, 21 LPS), 37 IL‑37tg mice (15 vehicle, 22 LPS), 29 mice deficient in IL-1R8 (IL‑1R8-KO; 10 vehicle, 19 LPS), and 25 mice transgenic for IL‑37 but deficient in IL-1R8 (IL‑37tg x IL-1R8-KO; 8 vehicle, 17 LPS) were injected with 10 mg/kg LPS (a) or vehicle (b). Spleens and lungs were collected 24 h later and cytokines measured by ELISA; mean ± s.e.m. cytokine concentrations normalized to total protein (t.p.) are shown. (c-e) PBMC were freshly isolated from 3 healthy volunteers, stimulated with 100 ng/ml LPS for 20 h and subjected to flow cytometry. (c, d) Results from one representative donor are depicted; (c) IL-1R8 and IL-18Rα in all PBMC, CD3+CD20+ lymphocytes, CD14+ monocytes, CD11c+ conventional DC, CD123+ plasmacytoid DC, and CD56+ NK cells; (d) gating strategies and staining controls. FMO, fluorescence-minus-one. (e) Percentage of PBMC subtypes among IL-1R8+IL-18Rα+ cells.
Supplementary Figure 2 Immunofluorescence microscopy and immunoprecipitation to characterize the interactions of the pairs IL-37–IL-1R8 and IL-37–IL-18Rα.
(a) Immunofluorescence of A549 cells transfected with IL‑37b and stimulated with 10 ng/ml IL-1β for 30 min. The first two panels in each row show the single color channels for IL‑37b-Flag in green and IL‑18Rα in red. The right panels in both rows show the overlays. One set of images out of nine with a similar pattern is shown. (b) Co-localization of IL‑37 with IL-1R8. After transfection with IL‑37b, A549 cells were treated with IL‑1β (10 ng/ml) for the indicated periods of time, followed by assessment of IL‑37 and IL-1R8 by immunofluorescence microscopy. IL‑37-Flag was stained with FITC (green) and IL-1R8 was detected by Cy3 (red). The images show one of n = 5 similar experiments for the 30 min time point, and one of n = 3 for 60 min. (c) Denaturing IL‑37b-Flag-immunoprecipitation in BMDM from IL‑37tg x IL-1R8-KO mice after stimulation with 1 μg/ml LPS for the indicated periods of time; 0 min is immediately before LPS. Staining against IL-1R8 in cells from two mice is shown; n = 3 mice. The 0 and 10 min time points are pooled. (d) Lysates of IL‑37b-transfected A549 cells were immunoprecipitated with an antibody against IL‑1R8 and stained for IL‑37. Duration of IL-1β-stimulation (10 ng/ml) is shown, denaturing SDS-PAGE; one of n = 3 similar results is shown.
Supplementary Figure 3 Specificity of the immunofluorescence experiments: loss of the IL-37–IL-18Rα co-localization signal by silencing of the gene encoding IL-18Rα in RAW cells.
RAW cells were transfected with 100 nM siRNA to Il18r1 or scrambled siRNA (scr), rested, then treated with LPS (100 ng/ml) for 30 min. Confocal microscopy was performed using FITC (green) to detect IL‑37b-Flag and Cy3 (red) to detect IL‑18Rα. Yellow color in the overlay indicates co-localization.
Supplementary Figure 4 Fluorescence lifetime maps of control conditions of the FLIM-FRET experiments.
PBMC were stimulated with 1 μg/ml of LPS or vehicle for the indicated periods of time, stained with IL-37-AlexaFluor555, IL-1R8-AlexaFluor488, or the pertinent IgG control antibodies, and assessed by FLIM-FRET. A decrease in fluorescence lifetime indicates interaction between IL-1R8 and IL‑37 (see bar below last row of images). The data of one of 3 similar experiments are shown.
Supplementary Figure 5 Control conditions for the PLA experiments.
Following treatment with vehicle or LPS (1 μg/ml) for 30 min, PBMC were fixed with 4% paraformaldehyde and incubated with antibodies to IL‑37, IL-1R8, and IL‑18Rα, or the relevant IgG control antibodies. Localization closer than 40 nm was visualized by the PLA reaction, which results in the presence of AlexaFluor568 foci. One representative result of n = 5 donors is shown.
Supplementary Figure 6 Additional data from the super-resolution, BRET, and immunoprecipitation experiments investigating the tripartite complex IL-37–IL-1R8–IL-18Rα.
(a-h) Fluorescence imaging of IL-37, IL-1R8 and IL-18Rα antibodies as well as their respective species IgG staining of PMBC. Scale bars are 3 μm. (a-d) Brightfield (a) and fluorescence (b-d) images of PMBC stained with primary antibodies for IL-37 (b), IL-18Rα (c), and IL-1R8 (d). (e-h) Brightfield (e) and fluorescence (f-h) images of negative control PMBC incubated with mouse IgG (f), rabbit IgG (g), and goat IgG (h) primary antibodies. Primary antibodies were detected with the respective secondary antibodies anti-mouse-AlexaFluor488, anti-rabbit-AlexaFluor568 and anti-goat-AlexaFluor647. (i-l) Variations in shape and structure of the ligand-receptor complex. Individual tripartite clusters show slightly different arrangements of the three proteins with respect to angles and distances. Complexes are shown either as fluorescence-spots overlay (i, j) or just as spots (k, l). The diameter of all depicted spots is 70 nm. Scale bars are 100 nm. (m-p) BRET. (m) HEK293 cells were transfected with C-terminally Rluc-tagged IL‑37 and either IL-1R8 or IL‑18Rα that carried an N-terminal Venus-tag, then incubated with LPS (100 ng/ml) for 3 h and analyzed by BRET. Mean ± s.e.m. BRET ratios are shown; n = 4 independent experiments. (n) BRET titration. After transfection with Rluc-IL‑37 and either Venus-IL-1R8 or Venus-IL-18Rα, the second untagged receptor was added to the HEK293 cultures at the indicated concentrations. The slope of the IL‑37-IL-1R8 + IL-18Rα (dotted line, open symbols) is 0.025, that of IL‑37-IL-18Rα + IL-1R8 (solid line, filled symbols) is 0.001; n ≥ 4. (o, p) BRET saturation and relative affinities in live RAW264.7 macrophages. (o) Cells were transfected with Rluc-IL‑37 (donor) and Venus-IL-1R8 (acceptor) at the indicated concentration ratios, stimulated with 100 ng/ml LPS for 3 h, and assessed by BRET; n = 4. (p) Relative affinities of the IL‑37-IL-1R8 interaction with and without co-transfection of untagged IL‑18Rα. (q) Native PAGE of PBMC treated with LPS (1 μg/ml). Lysates were obtained at the indicated time points. After immunoprecipitation of IL-37, the membranes were stained for IL-18Rα.
Supplementary Figure 7 Activity of IL‑37 in the presence or absence of IL‑18Rα and/or IL-1R8.
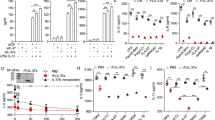
(a)* A549 clones depleted of IL‑18Rα were generated by stable transfection with shRNA; IL‑18Rα protein abundance as assessed by immunoblotting is depicted. (b) Abundance of IL‑37b protein was assessed by immunoblotting in the clones after both transfections and after 20 h of treatment with 10 ng/ml of IL‑1β. Two out of the three sh‑scrambled and both shIl18r1 clones are shown. (c) IL‑18Rα, IL-1R8, or both receptors were stably silenced in THP-1 cells, then each line was transfected with the IL‑37b expression vector or a mock construct, and treated with vehicle for 20 h. Percent changes in IL-1β protein abundance conferred by IL‑37b are depicted. n = 3-8. Differences between shIl18r1, shSigirr, and silencing of both receptors are not significant. (d) siRNA to Sigirr (siSigirr) or scrambled siRNA (scr) was transfected into PBMC and IL-6 abundance was measured in the culture supernatants 20 h after treatment with LPS (1 μg/ml), IL‑12 (20 ng/ml) plus IL‑18 (50 ng/ml), or vehicle. Mean ± s.e.m. absolute cytokine concentrations; n = 6. *, P < 0.05.
*Because the same clones as in were used for this experiment, this panel is identical to panel A of Figure 5 in our previous publication. This research was originally published in the Journal of Biological Chemistry. Nold-Petry C.A., et al. Increased cytokine production in interleukin-18 receptor alpha-deficient cells is associated with dysregulation of suppressors of cytokine signaling. J Biol Chem 2009;284:25900-25911. ©American Society for Biochemistry and Molecular Biology.
Supplementary Figure 8 Additional data on IL-37–IL-1R8–IL-18Rα signaling.
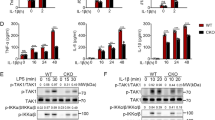
Intracellular mediators regulated by IL‑37 via IL-1R8 (a) or without the contribution of IL-1R8 (b,c). The identical experimental conditions and analyses as in Fig. 5 were performed (stimulation of pooled splenic macrophages and DC with 100 ng/ml LPS, harvest after 8 min, calculation of magnitude of intra-strain changes between vehicle- and LPS-treated conditions in activity of mediators). Results are presented (in arbitrary units, AU) as change in phosphorylation of individual sites in LPS-treated cells relative to their phosphorylation in vehicle-treated cells, with the raw percent change for each mediator multiplied by the average signal intensity of the pertinent antibody, to illustrate relative protein abundance and thus render signal abundance comparable across all markers (Δ phosphorylation x signal intensity). Black phosphosite indicates activating phosphorylation, red phosphosite represents inhibiting phosphorylation. *, P < 0.05; **, P < 0.01. (d-i) Gating strategy for phosphorylation-specific-flow cytometry on transduced and transfected THP-1 cells. To eliminate contaminating noise of dead cells, a transfected, unstimulated sample of all clones pooled was stained with a viability dye (Zombie Aqua, Biolegend). After gating for doublets (d,e), the FSC-A-SSC-A subset gate (f) was examined for viability (g). Further analysis of the dead cells (h) confirmed that most of the remaining contamination was <100k for FSC-A and <130k for SSC-A. Therefore, only cells >100k FSC-A and >130k SSC-A were included in the analysis (final gating in i). One such analysis was performed for every THP-1 phosphorylation-specific-flow cytometry experiment. (j-k) Gating strategy for phosphorylation-specific -flow cytometry on B- and T-cell depleted splenocytes from WT, IL‑37tg, and IL‑37tg x IL-1R8-KO mice. Live cells (j) were assessed for remaining FITC signals from non-depleted CD3+ and CD19+ cells (k). FITC- cells were then divided into a CD11c+ and a CD11c- population, which were subsequently analyzed separately (Fig. 6e-g and h-j, respectively). One such analysis was performed for every spleen.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8 and Supplementary Table 1 (PDF 1111 kb)
Supplementary Table 2: Details of IL-37-mediated gene regulation.
The differentially regulated genes shown in Fig. 8 are listed in the same groups they appear in Fig. 8, and the following information is provided: Column A, Ensembl ID. B, Gene symbol. C, Full gene name. D-F, normalized copy number per million reads, calculated by dividing the individual mRNA gene copy numbers of each strain by the total number of analyzed tags per strain, and multiplying the resulting number by 1 million. G-I, log2 fold change in gene expression between the strains. J-L, P-value, and M-O, false discovery rate (FDR), as calculated by DEG-seq (see Methods); an FDR of 0 indicates a high probability of differential expression. P-W, raw mRNA copy numbers (XLS 84 kb)
Supplementary Animation 1: Three-dimensional animation of Fig. 3a.
Widefield fluorescence image of three PBMC; IL-37, green; IL-18Ra, red; IL-1R8, blue. (MP4 2981 kb)
Rights and permissions
About this article
Cite this article
Nold-Petry, C., Lo, C., Rudloff, I. et al. IL-37 requires the receptors IL-18Rα and IL-1R8 (SIGIRR) to carry out its multifaceted anti-inflammatory program upon innate signal transduction. Nat Immunol 16, 354–365 (2015). https://doi.org/10.1038/ni.3103
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ni.3103