Abstract
The transcription factor ThPOK promotes CD4+ T cell differentiation in the thymus. Here, using a mouse strain that allows post-thymic gene deletion, we show that ThPOK maintains CD4+ T lineage integrity and couples effector differentiation to environmental cues after antigenic stimulation. ThPOK preserved the integrity and amplitude of effector responses and was required for proper differentiation of types 1 and 2 helper T cells in vivo by restraining the expression and function of Runx3, a nuclear factor crucial for cytotoxic T cell differentiation. The transcription factor LRF acts redundantly with ThPOK to prevent the transdifferentiation of mature CD4+ T cells into CD8+ T cells. As such, the ThPOK-LRF transcriptional module was essential for CD4+ T cell integrity and responses.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Accession codes
References
O'Shea, J.J. & Paul, W.E. Mechanisms underlying lineage commitment and plasticity of helper CD4+ T cells. Science 327, 1098–1102 (2010).
Zhu, J., Yamane, H. & Paul, W.E. Differentiation of effector CD4 T cell populations. Annu. Rev. Immunol. 28, 445–489 (2010).
Masopust, D., Vezys, V., Wherry, E.J. & Ahmed, R. A brief history of CD8 T cells. Eur. J. Immunol. 37 (suppl. 1), S103–S110 (2007).
Carpenter, A.C. & Bosselut, R. Decision checkpoints in the thymus. Nat. Immunol. 11, 666–673 (2010).
Taniuchi, I. et al. Differential requirements for Runx proteins in CD4 repression and epigenetic silencing during T lymphocyte development. Cell 111, 621–633 (2002).
Woolf, E. et al. Runx3 and Runx1 are required for CD8 T cell development during thymopoiesis. Proc. Natl. Acad. Sci. USA 100, 7731–7736 (2003).
He, X. et al. The zinc finger transcription factor Th-POK regulates CD4 versus CD8 T-cell lineage commitment. Nature 433, 826–833 (2005).
Sun, G. et al. The zinc finger protein cKrox directs CD4 lineage differentiation during intrathymic T cell positive selection. Nat. Immunol. 6, 373–381 (2005).
Wang, L. & Bosselut, R. CD4–CD8 lineage differentiation: Thpok-ing into the nucleus. J. Immunol. 183, 2903–2910 (2009).
Wang, L. et al. The zinc finger transcription factor Zbtb7b represses CD8-lineage gene expression in peripheral CD4+ T cells. Immunity 29, 876–887 (2008).
Muroi, S. et al. Cascading suppression of transcriptional silencers by ThPOK seals helper T cell fate. Nat. Immunol. 9, 1113–1121 (2008).
Egawa, T. & Littman, D.R. ThPOK acts late in specification of the helper T cell lineage and suppresses Runx-mediated commitment to the cytotoxic T cell lineage. Nat. Immunol. 9, 1131–1139 (2008).
Setoguchi, R. et al. Repression of the transcription factor Th-POK by Runx complexes in cytotoxic T cell development. Science 319, 822–825 (2008).
Davies, J.M. et al. Novel BTB/POZ domain zinc-finger protein, LRF, is a potential target of the LAZ-3/BCL-6 oncogene. Oncogene 18, 365–375 (1999).
Foy, T.M., Aruffo, A., Bajorath, J., Buhlmann, J.E. & Noelle, R.J. Immune regulation by CD40 and its ligand GP39. Annu. Rev. Immunol. 14, 591–617 (1996).
Carpenter, A.C. et al. The transcription factors Thpok and LRF are necessary and partly redundant for T helper cell differentiation. Immunity 37, 622–633 (2012).
Reis, B.S., Rogoz, A., Costa-Pinto, F.A., Taniuchi, I. & Mucida, D. Mutual expression of the transcription factors Runx3 and ThPOK regulates intestinal CD4+ T cell immunity. Nat. Immunol. 14, 271–280 (2013).
Djuretic, I.M. et al. Transcription factors T-bet and Runx3 cooperate to activate Ifng and silence Il4 in T helper type 1 cells. Nat. Immunol. 8, 145–153 (2007).
Naoe, Y. et al. Repression of interleukin-4 in T helper type 1 cells by Runx/Cbf β binding to the Il4 silencer. J. Exp. Med. 204, 1749–1755 (2007).
Lucas, B., Vasseur, F. & Penit, C. Normal sequence of phenotypic transitions in one cohort of 5-bromo-2′-deoxyuridine-pulse-labeled thymocytes. Correlation with T cell receptor expression. J. Immunol. 151, 4574–4582 (1993).
McCaughtry, T.M., Wilken, M.S. & Hogquist, K.A. Thymic emigration revisited. J. Exp. Med. 204, 2513–2520 (2007).
Wang, L. et al. Distinct functions for the transcription factors GATA-3 and ThPOK during intrathymic differentiation of CD4+ T cells. Nat. Immunol. 9, 1122–1130 (2008).
Guy-Grand, D. & Vassalli, P. Gut intraepithelial T lymphocytes. Curr. Opin. Immunol. 5, 247–252 (1993).
Taniuchi, I. & Ellmeier, W. Transcriptional and epigenetic regulation of CD4/CD8 lineage choice. Adv. Immunol. 110, 71–110 (2011).
Mangan, P.R. et al. Transforming growth factor-β induces development of the TH17 lineage. Nature 441, 231–234 (2006).
Oswald, I.P. et al. IL-12 inhibits Th2 cytokine responses induced by eggs of Schistosoma mansoni. J. Immunol. 153, 1707–1713 (1994).
Ho, I.C., Tai, T.S. & Pai, S.Y. GATA3 and the T-cell lineage: essential functions before and after T-helper-2-cell differentiation. Nat. Rev. Immunol. 9, 125–135 (2009).
Pearce, E.L. et al. Control of effector CD8+ T cell function by the transcription factor Eomesodermin. Science 302, 1041–1043 (2003).
Cruz-Guilloty, F. et al. Runx3 and T-box proteins cooperate to establish the transcriptional program of effector CTLs. J. Exp. Med. 206, 51–59 (2009).
Yagi, R. et al. The transcription factor GATA3 actively represses RUNX3 protein-regulated production of interferon-γ. Immunity 32, 507–517 (2010).
Betts, M.R. et al. Sensitive and viable identification of antigen-specific CD8+ T cells by a flow cytometric assay for degranulation. J. Immunol. Methods 281, 65–78 (2003).
Wildt, K.F. et al. The transcription factor zbtb7b promotes CD4 expression by antagonizing runx-mediated activation of the CD4 silencer. J. Immunol. 179, 4405–4414 (2007).
Grover, H.S. et al. The Toxoplasma gondii peptide AS15 elicits CD4 T cells that can control parasite burden. Infect. Immun. 80, 3279–3288 (2012).
Zhang, D.J. et al. Selective expression of the Cre recombinase in late-stage thymocytes using the distal promoter of the Lck gene. J. Immunol. 174, 6725–6731 (2005).
Wang, Q., Strong, J. & Killeen, N. Homeostatic competition among T cells revealed by conditional inactivation of the mouse Cd4 gene. J. Exp. Med. 194, 1721–1730 (2001).
Jenkinson, S.R. et al. Expression of the transcription factor cKrox in peripheral CD8 T cells reveals substantial postthymic plasticity in CD4–CD8 lineage differentiation. J. Exp. Med. 204, 267–272 (2007).
Rui, J., Liu, H., Zhu, X., Cui, Y. & Liu, X. Epigenetic silencing of Cd8 genes by ThPOK-mediated deacetylation during CD4 T cell differentiation. J. Immunol. 189, 1380–1390 (2012).
Mucida, D. et al. Transcriptional reprogramming of mature CD4+ helper T cells generates distinct MHC class II–restricted cytotoxic T lymphocytes. Nat. Immunol. 14, 281–289 (2013).
Zou, Y.R. et al. Epigenetic silencing of CD4 in T cells committed to the cytotoxic lineage. Nat. Genet. 29, 332–336 (2001).
Dupont, C.D., Christian, D.A. & Hunter, C.A. Immune response and immunopathology during toxoplasmosis. Semin. Immunopathol. 34, 793–813 (2012).
Reichmann, G. et al. The CD40/CD40 ligand interaction is required for resistance to toxoplasmic encephalitis. Infect. Immun. 68, 1312–1318 (2000).
Engel, I., Zhao, M., Kappes, D., Taniuchi, I. & Kronenberg, M. The transcription factor Th-POK negatively regulates Th17 differentiation in Vα14i NKT cells. Blood 120, 4524–4532 (2012).
Liu, Z. et al. Prevention of experimental colitis in SCID mice reconstituted with CD45RBhigh CD4+ T cells by blocking the CD40–CD154 interactions. J. Immunol. 164, 6005–6014 (2000).
Strober, W. & Fuss, I.J. Proinflammatory cytokines in the pathogenesis of inflammatory bowel diseases. Gastroenterology 140, 1756–1767 (2011).
Sakaguchi, S. et al. The zinc-finger protein MAZR is part of the transcription factor network that controls the CD4 versus CD8 lineage fate of double-positive thymocytes. Nat. Immunol. 11, 442–448 (2010).
Xiong, Y. & Bosselut, R. CD4–CD8 differentiation in the thymus: connecting circuits and building memories. Curr. Opin. Immunol. 24, 139–145 (2012).
Sattentau, Q.J. & Weiss, R.A. The CD4 antigen: physiological ligand and HIV receptor. Cell 52, 631–633 (1988).
Beaumier, C.M. et al. CD4 downregulation by memory CD4+ T cells in vivo renders African green monkeys resistant to progressive SIVagm infection. Nat. Med. 15, 879–885 (2009).
Vinton, C. et al. CD4-like immunological function by CD4− T cells in multiple natural hosts of simian immunodeficiency virus. J. Virol. 85, 8702–8708 (2011).
Maeda, T. et al. Role of the proto-oncogene Pokemon in cellular transformation and ARF repression. Nature 433, 278–285 (2005).
Intlekofer, A.M. et al. Anomalous type 17 response to viral infection by CD8+ T cells lacking T-bet and eomesodermin. Science 321, 408–411 (2008).
Srinivas, S. et al. Cre reporter strains produced by targeted insertion of EYFP and ECFP into the ROSA26 locus. BMC Dev. Biol. 1, 4 (2001).
Barnden, M.J., Allison, J., Heath, W.R. & Carbone, F.R. Defective TCR expression in transgenic mice constructed using cDNA-based α- and β-chain genes under the control of heterologous regulatory elements. Immunol. Cell Biol. 76, 34–40 (1998).
Sun, C.M. et al. Small intestine lamina propria dendritic cells promote de novo generation of Foxp3 T reg cells via retinoic acid. J. Exp. Med. 204, 1775–1785 (2007).
Spencer, S.P. et al. Adaptation of innate lymphoid cells to a micronutrient deficiency promotes type 2 barrier immunity. Science 343, 432–437 (2014).
Oldenhove, G. et al. Decrease of Foxp3+ Treg cell number and acquisition of effector cell phenotype during lethal infection. Immunity 31, 772–786 (2009).
Acknowledgements
We thank T. Ciucci for experimental assistance and discussions; T.-A. Lewis for animal care and genotyping; M. McGinty and Q. Xiao for technical assistance; X. Wu for microarray analyses; D. Littman, P.P. Pandolfi and S. Reiner for mice; M. Grigg for the T. gondii strain; J. Grainger, T. Hand and S. Spencer for discussions; and J. Ashwell and J. Brenchley for reading the manuscript. Supported by the US National Institutes of Health Intramural Research Programs of the National Cancer Institute, Center for Cancer Research, the National Institute of Allergy and Infectious Diseases and the Eunice Kennedy Shriver National Institute of Child Health and Human Development.
Author information
Authors and Affiliations
Contributions
M.S.V., L.W., Y.B. and R.B. designed the research; M.S.V., L.W., N.B, A.C.C., Y.X., L.C.W. and E.W. performed and analyzed experiments; K.-D.S. and P.E.L. constructed and provided CD2-Cre mice; R.B. supervised the research; M.S.V. and R.B. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Analyses of resting cell populations in Thpokpd mice.
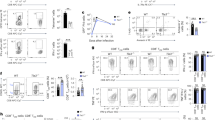
(a) Percentage of Rosa26YFP-expressing cells among CD4 or CD8SP thymocytes (Th) or spleen T cells (Spl) from CD2-Cre transgenic mice. (b,c) Plots indicate numbers (× 106) of total thymocytes (b, top), gated thymocytes (b, bottom three rows) and gated splenocytes (c). Each symbol represents one 6–8 week-old mouse. (a–c) Data are representative of four independent experiments. Thpok +/+ and Thpokpd mice are littermates to control for background effects. Significance was determined by Student’s t-test (*, P < 0.05, ** P < 0.01, *** P < 0.001, **** P < 0.0001, not significant in other cases). (d) Percent of Rosa26YFP-expressing cells among Foxp3-expressing CD4SP thymocytes or CD4+ spleen T cells from Thpokpd mice. Each symbol represents one mouse; data are from three experiments. (e) Immunoblot analysis of Runx protein expression in sorted spleen naïve CD4+ or CD8+ T from Thpokpd or Thpok+/+ littermate mice. The same membrane was subsequently stripped and reprobed for β-actin expression to evaluate lane loading. Representative of three independent experiments. (f) Contours plots show CD4 versus CD8 expression on total (left) or TCRβhiCD24lo (right) thymocytes from 6–9 week-old littermate mice of the indicated genotype. Representative of three experiments. (g) Absolute numbers of mature (TCRhi CD24lo) CD4SP thymocytes in mice of the indicated genotype. Each symbol represents one mouse. No significant difference among genotypes.
Supplementary Figure 2 TH17 differentiation of Thpokpd cells.
(a,b) Graphs show cumulative results for colon length (a) and mouse weight (b) from two independent C. rodentium infection experiments. Mean ± s.e.m. is shown. Each symbol represents a mouse. No significance by t-test. (c) ELISA quantification of IL-17 secretion by Thpokpd and Thpok+/+ cells activated under non-polarizing (THN), TH1 or TH17 conditions. Each symbol represents a separate culture. Data are from three mice of each genotype analyzed separately. Bars indicate mean ± s.e.m. No significance by t-test between Thpok+/+ and Thpokpd. (d) Histograms show GzmB expression in T effector cells obtained after 5-day cultures in TH17 conditions of sorted CD4+CD44lo spleen T cells from Thpok+/+ (solid grey) or Thpokpd (plain line) mice. The dashed line histogram shows GzmB expression on wild-type CD8+ effector cells cultured under THN conditions. Representative of three independent experiments.
Supplementary Figure 3 TH2 differentiation of Thpokpd cells.
(a) ELISA quantification of IL-4 secretion by Thpokpd and Thpok+/+ cells activated under nonpolarizing conditions as in Fig. 3e. Each symbol represents a separate culture. Data are from three distinct sorting experiments, each with a distinct mouse from each genotype. The average IL-4 concentrations were 0.17 ng/ml for Thpok+/+ cells and <0.01 ng/ml for Thpokpd cells (*P < 0.005 by paired t-test). (b) Scatter plots show the percentage of IL-4 or IFNγ producers among Thpokpd or Thpok+/+ cells cultured under non polarizing conditions as in Fig. 3e. Each symbol represents a culture derived from a single mouse. Statistical significance was determined by t-test (*P < 0.005; **P < 10–4).
Supplementary Figure 4 Functional relationships between ThPOK and Eomes and Cbfβ.
(a) Scatter plots show the percentage of IL 4- or IFNγ-producing cells among Thpok+/+, Thpokpd, Thpokpd Cbfbpd or Thpokpd Eomespd cells cultured under nonpolarizing conditions as in Fig. 3e. Each symbol represents a culture derived from a single mouse. Statistical significance was determined by t-test (*P <0.05; **P < 0.025; ***P < 0.01) on pooled data from three independent experiments. Contours plots (b) show CD4 versus CD8 expression on total (left) or TCRβhiCD24lo (center) thymocytes, or on TCRβ+ spleen cells (right) from 5 week old Thpok+/+Eomes+/+, Thpokpd Eomes+/+ or ThpokpdEomespd mice. Representative of three or four mice for each genotype from two independent experiments. (c) Graphs show the absolute numbers of CD4+CD8– and CD8+ T cells in the spleens from mice shown in b. There was no significant difference in the size of either subset among the three genotypes.
Supplementary Figure 5 Analyses of gene expression in Thpokpd effector cells.
(a) Contour plots show CD8 versus CD4 expression in Thpok+/+ and Thpokpd effector T cells derived from naïve CD4+CD8– cells as in Fig. 5b. (b) Heat map indicates relative expression (log2 values) on genes with 3-fold or greater change in expression in Thpokpd CD4+CD8+ vs. wild-type CD4+CD8– effector T cells.
Supplementary Figure 6 Post-thymic ThPOK restrains cytotoxic gene expression during in vivo TH1 responses.
(a) Contour plots assess uninfected (left) or day 10-infected (right) Thpok+/+ or Thpokpd spleen CD4+ T cells for expression of intra-cellular IFN-γ versus IL-17A. (b) Parasite load was quantified as percentage of RFP+ splenocytes in mice infected with RFP-expressing T. gondii. Each symbol represents one mouse; data are from two independent experiments. No significant difference by t-test. (c) Contour plots of Eomes versus T-bet expression in spleen CD4+ T cells from uninfected (left) or day 10-infected (right) Thpok+/+ or Thpokpd mice. Data (a,c) are representative of three experiments for each panel.
Supplementary Figure 7 Overlapping functions of LRF and ThPOK in CD4 expression.
(a) Histogram overlays show CD4 expression on sorted CD4lo (1), CD4+ CD8– (2) CD44lo spleen T cells from Thpokpd mice or their Thpok+/+ CD4+ CD8- counterparts (3) ex vivo (left panel), or after 5 d culture under TH1 conditions (right). Representative of three experiments. (b,c) Contours plots assess 6–8-week-old littermates of the indicated genotype for expression of CD4 and CD8 on total (left) and TCRβhiCD24lo (right) thymocytes (b), or (c) on total (left) or gated TCR+ YFP+ CD44lo (right) spleen T cells. Figures are representative of three experiments, summarized in d. (d) Plot summarizes numbers (× 106) of mature (TCRhi CD24lo) CD4 and CD8SP thymocytes (top) and of CD4+ (including CD4lo and pDP) and CD8+ T cells from 6–8 week-old mice of the indicated genotype. Significance was determined by Student’s t-test (*P < 0.05, ** P < 0.01; not significant otherwise). (e) Immunoblot analysis of Runx protein expression in effector T cells obtained after 5-d culture under TH1 conditions from naïve CD8+ T cells from Thpok+/+ Zbtb7a+/+ mice or CD4+ T cells from Thpokpd Zbtb7a+/+ or Thpokpd Zbtb7apd mice. The same membrane was subsequently stripped and reprobed for β-actin expression to evaluate lane loading. Representative of two immunoblots.
Supplementary information
Supplementary Text and Figures
Supplementary figures 1–7 (PDF 1170 kb)
Rights and permissions
About this article
Cite this article
Vacchio, M., Wang, L., Bouladoux, N. et al. A ThPOK-LRF transcriptional node maintains the integrity and effector potential of post-thymic CD4+ T cells. Nat Immunol 15, 947–956 (2014). https://doi.org/10.1038/ni.2960
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ni.2960
This article is cited by
-
ZBTB7B is a permissive regulator of hepatocellular carcinoma initiation by repressing c-Jun expression and function
Cell Death & Disease (2024)
-
Transcription factor EGR2 controls homing and pathogenicity of TH17 cells in the central nervous system
Nature Immunology (2023)
-
CD4 expression in effector T cells depends on DNA demethylation over a developmentally established stimulus-responsive element
Nature Communications (2022)
-
The transcription factor LRF promotes integrin β7 expression by and gut homing of CD8αα+ intraepithelial lymphocyte precursors
Nature Immunology (2022)
-
Leukemia/lymphoma-related factor (LRF) or osteoclast zinc finger protein (OCZF) overexpression promotes osteoclast survival by increasing Bcl-xl mRNA: A novel regulatory mechanism mediated by the RNA binding protein SAM68
Laboratory Investigation (2022)