Abstract
Tissue-resident memory T cells (TRM cells) provide superior protection against infection in extralymphoid tissues. Here we found that CD103+CD8+ TRM cells developed in the skin from epithelium-infiltrating precursor cells that lacked expression of the effector-cell marker KLRG1. A combination of entry into the epithelium plus local signaling by interleukin 15 (IL-15) and transforming growth factor-β (TGF-β) was required for the formation of these long-lived memory cells. Notably, differentiation into TRM cells resulted in the progressive acquisition of a unique transcriptional profile that differed from that of circulating memory cells and other types of T cells that permanently reside in skin epithelium. We provide a comprehensive molecular framework for the local differentiation of a distinct peripheral population of memory cells that forms a first-line immunological defense system in barrier tissues.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
References
von Andrian, U.H. & Mackay, C.R. T-cell function and migration. Two sides of the same coin. N. Engl. J. Med. 343, 1020–1034 (2000).
Kündig, T.M. et al. On the role of antigen in maintaining cytotoxic T-cell memory. Proc. Natl. Acad. Sci. USA 93, 9716–9723 (1996).
Yang, L., Yu, Y., Kalwani, M., Tseng, T.W. & Baltimore, D. Homeostatic cytokines orchestrate the segregation of CD4 and CD8 memory T-cell reservoirs in mice. Blood 118, 3039–3050 (2011).
Masopust, D. et al. Dynamic T cell migration program provides resident memory within intestinal epithelium. J. Exp. Med. 207, 553–564 (2010).
Gebhardt, T. et al. Different patterns of peripheral migration by memory CD4+ and CD8+ T cells. Nature 477, 216–219 (2011).
Woodland, D.L. & Kohlmeier, J.E. Migration, maintenance and recall of memory T cells in peripheral tissues. Nat. Rev. Immunol. 9, 153–161 (2009).
Bevan, M.J. Memory T cells as an occupying force. Eur. J. Immunol. 41, 1192–1195 (2011).
Gebhardt, T., Mueller, S.N., Heath, W.R. & Carbone, F.R. Peripheral tissue surveillance and residency by memory T cells. Trends Immunol. 34, 27–32 (2013).
Gebhardt, T. et al. Memory T cells in nonlymphoid tissue that provide enhanced local immunity during infection with herpes simplex virus. Nat. Immunol. 10, 524–530 (2009).
Kim, S.K., Reed, D.S., Heath, W.R., Carbone, F. & Lefrancois, L. Activation and migration of CD8 T cells in the intestinal mucosa. J. Immunol. 159, 4295–4306 (1997).
Wakim, L.M., Woodward-Davis, A. & Bevan, M.J. Memory T cells persisting within the brain after local infection show functional adaptations to their tissue of residence. Proc. Natl. Acad. Sci. USA 107, 17872–17879 (2010).
Hofmann, M. & Pircher, H. E-cadherin promotes accumulation of a unique memory CD8 T-cell population in murine salivary glands. Proc. Natl. Acad. Sci. USA 108, 16741–16746 (2011).
Jiang, X. et al. Skin infection generates non-migratory memory CD8+ TRM cells providing global skin immunity. Nature 483, 227–231 (2012).
Mackay, L.K. et al. Long-lived epithelial immunity by tissue-resident memory T (TRM) cells in the absence of persisting local antigen presentation. Proc. Natl. Acad. Sci. USA 109, 7037–7042 (2012).
Shin, H. & Iwasaki, A. A vaccine strategy that protects against genital herpes by establishing local memory T cells. Nature 491, 463–467 (2012).
Sallusto, F., Lenig, D., Forster, R., Lipp, M. & Lanzavecchia, A. Two subsets of memory T lymphocytes with distinct homing potentials and effector functions. Nature 401, 708–712 (1999).
Klonowski, K.D. et al. Dynamics of blood-borne CD8 memory T cell migration in vivo. Immunity 20, 551–562 (2004).
Sheridan, B.S. & Lefrancois, L. Regional and mucosal memory T cells. Nat. Immunol. 12, 485–491 (2011).
Kaech, S.M. & Wherry, E.J. Heterogeneity and cell-fate decisions in effector and memory CD8+ T cell differentiation during viral infection. Immunity 27, 393–405 (2007).
Obar, J.J. & Lefrancois, L. Early events governing memory CD8+ T-cell differentiation. Int. Immunol. 22, 619–625 (2010).
Masopust, D., Vezys, V., Marzo, A.L. & Lefrancois, L. Preferential localization of effector memory cells in nonlymphoid tissue. Science 291, 2413–2417 (2001).
Masopust, D., Vezys, V., Wherry, E.J., Barber, D.L. & Ahmed, R. Cutting edge: gut microenvironment promotes differentiation of a unique memory CD8+ T cell population. J. Immunol. 176, 2079–2083 (2006).
Tang, V.A. & Rosenthal, K.L. Intravaginal infection with herpes simplex virus type-2 (HSV-2) generates a functional effector memory T cell population that persists in the murine genital tract. J. Reprod. Immunol. 87, 39–44 (2010).
Kane, C.J., Knapp, A.M., Mansbridge, J.N. & Hanawalt, P.C. Transforming growth factor-beta 1 localization in normal and psoriatic epidermal keratinocytes in situ. J. Cell Physiol. 144, 144–150 (1990).
Koyama, S.Y. & Podolsky, D.K. Differential expression of transforming growth factors alpha and beta in rat intestinal epithelial cells. J. Clin. Invest. 83, 1768–1773 (1989).
Wang, D. et al. Regulation of CD103 expression by CD8+ T cells responding to renal allografts. J. Immunol. 172, 214–221 (2004).
Lee, Y.T. et al. Environmental and antigen receptor-derived signals support sustained surveillance of the lungs by pathogen-specific cytotoxic T lymphocytes. J. Virol. 85, 4085–4094 (2011).
Casey, K.A. et al. Antigen-independent differentiation and maintenance of effector-like resident memory T cells in tissues. J. Immunol. 188, 4866–4875 (2012).
Spangrude, G.J., Sacchi, F., Hill, H.R., Van Epps, D.E. & Daynes, R.A. Inhibition of lymphocyte and neutrophil chemotaxis by pertussis toxin. J. Immunol. 135, 4135–4143 (1985).
Bromley, S.K., Thomas, S.Y. & Luster, A.D. Chemokine receptor CCR7 guides T cell exit from peripheral tissues and entry into afferent lymphatics. Nat. Immunol. 6, 895–901 (2005).
Debes, G.F. et al. CC chemokine receptor 7 expression by effector/memory CD4+ T cells depends on antigen specificity and tissue localization during influenza A virus infection. J. Virol. 78, 7528–7535 (2004).
Kaech, S.M., Hemby, S., Kersh, E. & Ahmed, R. Molecular and functional profiling of memory CD8 T cell differentiation. Cell 111, 837–851 (2002).
El-Asady, R. et al. TGF-{beta}-dependent CD103 expression by CD8+ T cells promotes selective destruction of the host intestinal epithelium during graft-versus-host disease. J. Exp. Med. 201, 1647–1657 (2005).
Tripp, R.A., Hou, S. & Doherty, P.C. Temporal loss of the activated L-selectin-low phenotype for virus-specific CD8+ memory T cells. J. Immunol. 154, 5870–5875 (1995).
Wherry, E.J. et al. Lineage relationship and protective immunity of memory CD8 T cell subsets. Nat. Immunol. 4, 225–234 (2003).
Kaech, S.M. et al. Selective expression of the interleukin 7 receptor identifies effector CD8 T cells that give rise to long-lived memory cells. Nat. Immunol. 4, 1191–1198 (2003).
Sarkar, S. et al. Functional and genomic profiling of effector CD8 T cell subsets with distinct memory fates. J. Exp. Med. 205, 625–640 (2008).
Wakim, L.M., Waithman, J., van Rooijen, N., Heath, W.R. & Carbone, F.R. Dendritic cell-induced memory T cell activation in nonlymphoid tissues. Science 319, 198–202 (2008).
Çuburu, N. et al. Intravaginal immunization with HPV vectors induces tissue-resident CD8+ T cell responses. J. Clin. Invest. 122, 4606–4620 (2012).
Debes, G.F. et al. Chemokine receptor CCR7 required for T lymphocyte exit from peripheral tissues. Nat. Immunol. 6, 889–894 (2005).
Wakim, L.M. et al. The molecular signature of tissue resident memory CD8+ T cells isolated from the brain. J. Immunol. 189, 3462–3471 (2012).
Matloubian, M. et al. Lymphocyte egress from thymus and peripheral lymphoid organs is dependent on S1P receptor 1. Nature 427, 355–360 (2004).
Skon, C.N. et al. Transcriptional downregulation of S1PR1 is required for establishment of resident memory CD8+ T cells. Nat. Immunol. 10.1038/ni.2745 (2013).
Shiow, L.R. et al. CD69 acts downstream of interferon-α/β to inhibit S1P1 and lymphocyte egress from lymphoid organs. Nature 440, 540–544 (2006).
Jenne, C.N. et al. T-bet-dependent S1P5 expression in NK cells promotes egress from lymph nodes and bone marrow. J. Exp. Med. 206, 2469–2481 (2009).
Druey, K.M., Blumer, K.J., Kang, V.H. & Kehrl, J.H. Inhibition of G-protein-mediated MAP kinase activation by a new mammalian gene family. Nature 379, 742–746 (1996).
Gebhardt, T. & Mackay, L.K. Local immunity by tissue-resident CD8+ memory T cells. Front. Immunol. 3, 340 (2012).
Jameson, J. et al. A role for skin γδ T cells in wound repair. Science 296, 747–749 (2002).
Sharp, L.L., Jameson, J.M., Cauvi, G. & Havran, W.L. Dendritic epidermal T cells regulate skin homeostasis through local production of insulin-like growth factor 1. Nat. Immunol. 6, 73–79 (2005).
Waithman, J., Gebhardt, T., Davey, G.M., Heath, W.R. & Carbone, F.R. Cutting edge: Enhanced IL-2 signaling can convert self-specific T cell response from tolerance to autoimmunity. J. Immunol. 180, 5789–5793 (2008).
Livak, K.J. & Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(−ΔΔC(T)) method. Methods 25, 402–408 (2001).
Irizarry, R.A et al. Summaries of Affymetrix GeneChip probe level data. Nucleic Acids Res. 31, e15 (2003).
Smyth, G.K. Linear models and empirical bayes methods for assessing differential expression in microarray experiments. Stat. Appl. Genet. Mol. Biol. 3, Article 3 (2004).
Acknowledgements
We thank W. Weninger (University of Sydney) for Ccr7−/− mice; N. Hunt (University of Sydney) for Cxcr3−/− mice; M. Bevan (University of Washington) for Tgfbr2f/fdLck-Cre OT-I mice; I. Frazer (University of Queensland) for Tcrd−/− mice; members of the Carbone and Heath laboratories for discussions; and N. McBain and J. Smith for technical assistance. Supported by National Health and Medical Research Council of Australia and Australian Research Council.
Author information
Authors and Affiliations
Contributions
L.K.M., F.R.C. and T.G. designed the experiments; L.K.M., A.R., J.Z.M., N.C., A.T.S., M.-L.H. and T.G. did the experiments; L.K.M., A.R., J.Z.M., A.T.S., W.R.H., M.I., F.R.C. and T.G. analyzed the data; J.V.-R., P.L., S.N.M., T.S. and D.C.T. contributed reagents; F.R.C. led the research program and, with the help of L.K.M. and T.G. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Phenotype and anatomical localization of TRM cells.
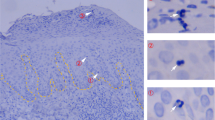
(a,b) Analysis of CD103, Bcl-2 and CD69 expression by endogenous Vα2+ T cells in skin at the indicated times p.i. Plots are gated on Vα2+ CD45.2+ cells; numbers indicate the percentage of events in the respective gates. Data representative of 2–3 experiments. (c) Mice received naïve gBT-I.GFP cells prior to HSV infection. Arrows indicate examples of gBT-I.GFP cells in the epidermis and hair follicle epithelium 14 d p.i. Photo representative of >5 experiments. (d) Wild-type (WT) and Cd69−/− gBT-I cells were transferred into WT mice prior to HSV infection. Enumeration of gBT-I cells in dorsal root ganglia (DRG) 30 d p.i. Data from one experiment (n = 5 mice/group). (e) WT (GFP-expressing, green) and Itgae−/− (DsRed-expressing, red) gBT-I cells were co-transferred into WT mice prior to HSV infection. Microscopy of skin 8 d p.i. Staining with anti-keratin antibody to delineate the epidermis and hair follicle epithelium. Arrows in insert indicate gBT-I cells in the epidermis. Photo representative of n = 3 mice analyzed.
Supplementary Figure 2 Development of virus-specific CD8+ memory T cells after infection of skin with HSV-1.
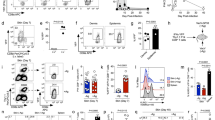
(a) Mice received naïve gBT-I cells prior to infection. Analysis of CD62L, CD127 and KLRG1 expression by gBT-I cells in the spleen at different time points after infection. Plots are gated on Vα2+CD45.2+CD45.1+ cells and are representative of n = 4–8 mice/time point. Numbers indicate the percentage of events in the respective gates (as in b). (b) Analysis of CD103 and KLRG1 expression by endogenous Vα2+ T cells in skin at the indicated times p.i. Plots are gated on Vα2+CD45.2+ cells and are representative of 3 experiments (n = 12 mice/time point). (c) Effector gBT-I cells were sorted into KLRG1+ and KLRG1− subsets from spleens of infected mice (6 d p.i.) and transferred into infected recipients (4 d p.i.). Enumeration of CD103+ gBT-I cells in dorsal root ganglia (DRG) 3 weeks p.i. Data from 3 experiments (n = 12 mice/group); *, P< 0.05 by two-tailed Mann-Whitney test.
Supplementary Figure 3 Epidermal localization of CD8+ T cells after intradermal transfer.
(a) CD103 and CD69 expression by gBT-I cells in skin at the indicated times after in vitro activation by gB peptide-coated splenocytes and intradermal transfer. Plots are gated on Vα2+CD45.2+CD45.1+ cells and are representative of 2–3 experiments (n = 6–12 mice/time point). Numbers indicate the percentage of events in the respective gates. (b) In vitro activated gBT-I.GFP cells were transferred into the skin by intradermal injection. Microscopy of skin 30 d after transfer. Staining with anti-keratin antibody to denote epithelium in the epidermis and hair follicles. Arrows indicate examples of gBT-I.GFP cells (green). Photos (2 examples shown) are representative of 2 experiments. (c) In vitro activated gBT-I cells were left untreated (Ctrl, DsRed-expressing, red) or treated with PTx (+PTx, GFP-expressing, green) and co-transferred into mice by intradermal injection. Microscopy of skin 4 weeks after transfer; anti-keratin staining denotes the epidermal layer and hair follicle epithelium. Red arrows indicate control cells in the epithelium, green arrows indicate PTx-treated cells in the dermis. Photos representative of 2 experiments. (d) In vitro activated gBT-I cells were untreated (Ctrl) or treated with PTx (+PTx). Analysis of CD103 expression following subsequent in vitro incubation of the cells in the presence (solid or dashed black lines, as indicated) or absence (orange area) of TGF-b (5 ng/ml; 24 hours). Plots are gated on Vα2+CD45.2+CD45.1+ cells and are representative of 2 experiments.
Supplementary Figure 4 Chemokine expression in skin and ex vivo migration of KLRG1+ and KLRG1− effector cells.
(a) mRNA expression for genes encoding CXCL9 and CXCL10 in CD45.2−EpCAM+ keratinocytes sorted from naïve (Ctrl) or HSV-infected skin (d 6 p.i.). Data pooled from 3 independent experiments (represented by individual symbols). *, P< 0.05 by two-tailed paired t-test. (b) Ex vivo migration towards CXCL9 gradients by KLRG1+ (closed symbols) and KLRG1− (open symbols) gBT-I cells enriched from spleens 7 d after HSV skin infection (cells pooled from n = 5 donor mice).
Supplementary Figure 5 Requirement for the TGF-β receptor and IL-15 signaling in the development of CD103+ TRM cells.
(a) Wild-type (WT) and TGF-R-deficient (Tgfbr2f/f.dLck-Cre) OT-I cells were activated by culture with ovalbumin peptide-coated splenocytes and transferred into the skin by intradermal injection. Depicted are the ratios of WT relative to Tgfbr2f/f.dLck-Cre cells in the skin and spleen at the indicated times after transfer. Data representative of 2 experiments (n = 3–4 mice/group). (b,c) Effector gBT-I cells were enriched from spleens of WT mice (6 d p.i.) and transferred into infected (4 d p.i.) WT or Il15−/− recipient mice. (b) Enumeration of CD103+ gBT-I cells in dorsal root ganglia (DRG) 4 wks p.i. Data from one experiment with n = 5 mice/group. (c) CD103 and Bcl-2 expression by gBT-I cells in WT and Il15−/− mice (11 d p.i.). Plots are gated on Vα2+CD45.2+CD45.1+ cells; numbers depict percentages of events in the respective gates. Data are representative of 2 experiments (n = 8 mice/group).
Supplementary Figure 6 The developmental pathway for the formation of TRM cells.
KLRG1− TRM precursors enter the dermis where they may (i) die in situ, (ii) exit the skin and return to the blood in a CCR7-dependent manner or (iii) migrate to the epidermis, partly under the influence of CXCR3 ligands. Both tissue exit and epidermal entry are sensitive to treatment with pertussis toxin (PTx) indicating the involvement of G protein-coupled molecules such as chemokine receptors. Following epidermal entry, TRM precursors undergo maturation into long-lived CD103+ TRM cells in a TGF-β- and IL-15-dependent manner. Epithelial TRM cells and their counterparts in the circulation are depicted with different symbols to highlight their distinct transcriptional profiles.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6 (PDF 9504 kb)
Rights and permissions
About this article
Cite this article
Mackay, L., Rahimpour, A., Ma, J. et al. The developmental pathway for CD103+CD8+ tissue-resident memory T cells of skin. Nat Immunol 14, 1294–1301 (2013). https://doi.org/10.1038/ni.2744
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ni.2744
This article is cited by
-
Getting under the skin: resident memory CD8+ T cells have a second residence in the draining lymph node
Genes & Immunity (2024)
-
Aging modifies endometrial dendritic cell function and unconventional double negative T cells in the human genital mucosa
Immunity & Ageing (2023)
-
IKK2/NFkB signaling controls lung resident CD8+ T cell memory during influenza infection
Nature Communications (2023)
-
A virtual memory CD8+ T cell-originated subset causes alopecia areata through innate-like cytotoxicity
Nature Immunology (2023)
-
T cell egress via lymphatic vessels is tuned by antigen encounter and limits tumor control
Nature Immunology (2023)