Abstract
The activation of immune-defense mechanisms in response to a microbial attack must be robust and appropriately tailored to fight particular types of pathogens. Infection with intracellular microorganisms elicits a type 1 inflammatory response characterized by mobilization of T helper type 1 (TH1) cells to the site of infection, where they are responsible for the recruitment and activation of macrophages. At the center of the type 1 inflammatory response is the transcription factor T-bet, a critical regulator of the TH1 differentiation program. T-bet induces the production of interferon-γ (IFN-γ) and orchestrates the TH1 cell–migratory program by regulating the expression of chemokines and chemokine receptors. However, tight regulation of the type 1 inflammatory response is essential for the prevention of immunopathology and the development of organ-specific autoimmunity. In this review, we discuss how T-bet expression drives autoaggressive and inflammatory processes and how its function in vivo must be delicately balanced to avoid disease.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

Katie Vicari

Katie Vicari

Katie Vicari

Katie Vicari
Similar content being viewed by others
References
Mosmann, T.R., Cherwinski, H., Bond, M.W., Giedlin, M.A. & Coffman, R.L. Two types of murine helper T cell clone. I. Definition according to profiles of lymphokine activities and secreted proteins. J. Immunol. 136, 2348–2357 (1986).
Szabo, S.J. et al. A novel transcription factor, T-bet, directs Th1 lineage commitment. Cell 100, 655–669 (2000).
Zheng, W. & Flavell, R.A. The transcription factor GATA-3 is necessary and sufficient for Th2 cytokine gene expression in CD4 T cells. Cell 89, 587–596 (1997).
Harrington, L.E. et al. Interleukin 17-producing CD4+ effector T cells develop via a lineage distinct from the T helper type 1 and 2 lineages. Nat. Immunol. 6, 1123–1132 (2005).
Park, H. et al. A distinct lineage of CD4 T cells regulates tissue inflammation by producing interleukin 17. Nat. Immunol. 6, 1133–1141 (2005).
Hwang, E.S., Szabo, S.J., Schwartzberg, P.L. & Glimcher, L.H. T helper cell fate specified by kinase-mediated interaction of T-bet with GATA-3. Science 307, 430–433 (2005).
Lazarevic, V. et al. T-bet represses TH17 differentiation by preventing Runx1-mediated activation of the gene encoding RORγt. Nat. Immunol. 12, 96–104 (2011).
Villarino, A.V., Gallo, E. & Abbas, A.K. STAT1-activating cytokines limit Th17 responses through both T-bet-dependent and -independent mechanisms. J. Immunol. 185, 6461–6471 (2010).
Esensten, J.H., Lee, M.R., Glimcher, L.H. & Bluestone, J.A. T-bet-deficient NOD mice are protected from diabetes due to defects in both T cell and innate immune system function. J. Immunol. 183, 75–82 (2009).
Wang, J. et al. Transcription factor T-bet regulates inflammatory arthritis through its function in dendritic cells. J. Clin. Invest. 116, 414–421 (2006).
Sullivan, B.M., Juedes, A., Szabo, S.J., von Herrath, M. & Glimcher, L.H. Antigen-driven effector CD8 T cell function regulated by T-bet. Proc. Natl. Acad. Sci. USA 100, 15818–15823 (2003).
Peng, S.L., Szabo, S.J. & Glimcher, L.H. T-bet regulates IgG class switching and pathogenic autoantibody production. Proc. Natl. Acad. Sci. USA 99, 5545–5550 (2002).
Townsend, M.J. et al. T-bet regulates the terminal maturation and homeostasis of NK and Va14i NKT cells. Immunity 20, 477–494 (2004).
Afkarian, M. et al. T-bet is a STAT1-induced regulator of IL-12R expression in naive CD4+ T cells. Nat. Immunol. 3, 549–557 (2002).
Lighvani, A.A. et al. T-bet is rapidly induced by interferon-γ in lymphoid and myeloid cells. Proc. Natl. Acad. Sci. USA 98, 15137–15142 (2001).
Mullen, A.C. et al. Role of T-bet in commitment of TH1 cells before IL-12-dependent selection. Science 292, 1907–1910 (2001).
Schulz, E.G., Mariani, L., Radbruch, A. & Hofer, T. Sequential polarization and imprinting of type 1 T helper lymphocytes by interferon-γ and interleukin-12. Immunity 30, 673–683 (2009).
Hu, X. & Ivashkiv, L.B. Cross-regulation of signaling pathways by interferon-γ: implications for immune responses and autoimmune diseases. Immunity 31, 539–550 (2009).
Jenner, R.G. et al. The transcription factors T-bet and GATA-3 control alternative pathways of T-cell differentiation through a shared set of target genes. Proc. Natl. Acad. Sci. USA 106, 17876–17881 (2009).
Lord, G.M. et al. T-bet is required for optimal proinflammatory CD4+ T-cell trafficking. Blood 106, 3432–3439 (2005).
Sullivan, B.M. et al. Increased susceptibility of mice lacking T-bet to infection with Mycobacterium tuberculosis correlates with increased IL-10 and decreased IFN-γ production. J. Immunol. 175, 4593–4602 (2005).
Szabo, S.J. et al. Distinct effects of T-bet in TH1 lineage commitment and IFN-γ production in CD4 and CD8 T cells. Science 295, 338–342 (2002).
Hultgren, O.H., Verdrengh, M. & Tarkowski, A. T-box transcription-factor-deficient mice display increased joint pathology and failure of infection control during staphylococcal arthritis. Microbes Infect. 6, 529–535 (2004).
Ravindran, R., Foley, J., Stoklasek, T., Glimcher, L.H. & McSorley, S.J. Expression of T-bet by CD4 T cells is essential for resistance to Salmonella infection. J. Immunol. 175, 4603–4610 (2005).
Neurath, M.F. et al. The transcription factor T-bet regulates mucosal T cell activation in experimental colitis and Crohn's disease. J. Exp. Med. 195, 1129–1143 (2002).
Bettelli, E. et al. Loss of T-bet, but not STAT1, prevents the development of experimental autoimmune encephalomyelitis. J. Exp. Med. 200, 79–87 (2004).
Lovett-Racke, A.E. et al. Silencing T-bet defines a critical role in the differentiation of autoreactive T lymphocytes. Immunity 21, 719–731 (2004).
Juedes, A.E., Rodrigo, E., Togher, L., Glimcher, L.H. & von Herrath, M.G. T-bet controls autoaggressive CD8 lymphocyte responses in type 1 diabetes. J. Exp. Med. 199, 1153–1162 (2004).
Flynn, J.L. et al. An essential role for interferon γ in resistance to Mycobacterium tuberculosis infection. J. Exp. Med. 178, 2249–2254 (1993).
Huang, S. et al. Immune response in mice that lack the interferon-γ receptor. Science 259, 1742–1745 (1993).
Harty, J.T. & Bevan, M.J. Specific immunity to Listeria monocytogenes in the absence of IFN γ. Immunity 3, 109–117 (1995).
Way, S.S. & Wilson, C.B. Cutting edge: immunity and IFN-γ production during Listeria monocytogenes infection in the absence of T-bet. J. Immunol. 173, 5918–5922 (2004).
Fuss, I.J. et al. Disparate CD4+ lamina propria (LP) lymphokine secretion profiles in inflammatory bowel disease. Crohn's disease LP cells manifest increased secretion of IFN-γ, whereas ulcerative colitis LP cells manifest increased secretion of IL-5. J. Immunol. 157, 1261–1270 (1996).
Plevy, S.E. et al. A role for TNF-α and mucosal T helper-1 cytokines in the pathogenesis of Crohn's disease. J. Immunol. 159, 6276–6282 (1997).
Targan, S.R., Deem, R.L., Liu, M., Wang, S. & Nel, A. Definition of a lamina propria T cell responsive state. Enhanced cytokine responsiveness of T cells stimulated through the CD2 pathway. J. Immunol. 154, 664–675 (1995).
Matsuoka, K. et al. T-bet upregulation and subsequent interleukin 12 stimulation are essential for induction of Th1 mediated immunopathology in Crohn's disease. Gut 53, 1303–1308 (2004).
Yang, Y., Xu, J., Niu, Y., Bromberg, J.S. & Ding, Y. T-bet and eomesodermin play critical roles in directing T cell differentiation to Th1 versus Th17. J. Immunol. 181, 8700–8710 (2008).
Sasaki, Y. et al. Identification of a novel type 1 diabetes susceptibility gene, T-bet. Hum. Genet. 115, 177–184 (2004).
Hultgren, B., Huang, X., Dybdal, N. & Stewart, T.A. Genetic absence of γ-interferon delays but does not prevent diabetes in NOD mice. Diabetes 45, 812–817 (1996).
Serreze, D.V. et al. Interferon-gamma receptor signaling is dispensable in the development of autoimmune type 1 diabetes in NOD mice. Diabetes 49, 2007–2011 (2000).
Kaplan, M.H., Schindler, U., Smiley, S.T. & Grusby, M.J. Stat6 is required for mediating responses to IL-4 and for development of Th2 cells. Immunity 4, 313–319 (1996).
Kurata, H., Lee, H.J., O'Garra, A. & Arai, N. Ectopic expression of activated Stat6 induces the expression of Th2-specific cytokines and transcription factors in developing Th1 cells. Immunity 11, 677–688 (1999).
Zhu, J., Guo, L., Watson, C.J., Hu-Li, J. & Paul, W.E. Stat6 is necessary and sufficient for IL-4's role in Th2 differentiation and cell expansion. J. Immunol. 166, 7276–7281 (2001).
Djuretic, I.M. et al. Transcription factors T-bet and Runx3 cooperate to activate Ifng and silence Il4 in T helper type 1 cells. Nat. Immunol. 8, 145–153 (2007).
Avni, O. et al. TH cell differentiation is accompanied by dynamic changes in histone acetylation of cytokine genes. Nat. Immunol. 3, 643–651 (2002).
Finotto, S. et al. Development of spontaneous airway changes consistent with human asthma in mice lacking T-bet. Science 295, 336–338 (2002).
Finotto, S. et al. Asthmatic changes in mice lacking T-bet are mediated by IL-13. Int. Immunol. 17, 993–1007 (2005).
Munthe-Kaas, M.C. et al. T cell-specific T-box transcription factor haplotype is associated with allergic asthma in children. J. Allergy Clin. Immunol. 121, 51–56 (2008).
Raby, B.A. et al. T-bet polymorphisms are associated with asthma and airway hyperresponsiveness. Am. J. Respir. Crit. Care Med. 173, 64–70 (2006).
Kiwamoto, T. et al. Transcription factors T-bet and GATA-3 regulate development of airway remodeling. Am. J. Respir. Crit. Care Med. 174, 142–151 (2006).
Bettelli, E. et al. Reciprocal developmental pathways for the generation of pathogenic effector TH17 and regulatory T cells. Nature 441, 235–238 (2006).
Ivanov, I.I. et al. The orphan nuclear receptor RORγt directs the differentiation program of proinflammatory IL-17+ T helper cells. Cell 126, 1121–1133 (2006).
Mangan, P.R. et al. Transforming growth factor-beta induces development of the TH17 lineage. Nature 441, 231–234 (2006).
Veldhoen, M., Hocking, R.J., Atkins, C.J., Locksley, R.M. & Stockinger, B. TGFβ in the context of an inflammatory cytokine milieu supports de novo differentiation of IL-17-producing T cells. Immunity 24, 179–189 (2006).
Korn, T. et al. IL-21 initiates an alternative pathway to induce proinflammatory TH17 cells. Nature 448, 484–487 (2007).
Zhou, L. et al. IL-6 programs TH-17 cell differentiation by promoting sequential engagement of the IL-21 and IL-23 pathways. Nat. Immunol. 8, 967–974 (2007).
Nurieva, R. et al. Essential autocrine regulation by IL-21 in the generation of inflammatory T cells. Nature 448, 480–483 (2007).
Brustle, A. et al. The development of inflammatory TH-17 cells requires interferon-regulatory factor 4. Nat. Immunol. 8, 958–966 (2007).
Schraml, B.U. et al. The AP-1 transcription factor Batf controls TH17 differentiation. Nature 460, 405–409 (2009).
Zhang, F., Meng, G. & Strober, W. Interactions among the transcription factors Runx1, RORγt and Foxp3 regulate the differentiation of interleukin 17-producing T cells. Nat. Immunol. 9, 1297–1306 (2008).
Mukasa, R. et al. Epigenetic instability of cytokine and transcription factor gene loci underlies plasticity of the T helper 17 cell lineage. Immunity 32, 616–627 (2010).
Bending, D. et al. Epigenetic changes at il12rb2 and tbx21 in relation to plasticity behavior of th17 cells. J. Immunol. 186, 3373–3382 (2011).
Doodes, P.D. et al. IFN-γ regulates the requirement for IL-17 in proteoglycan-induced arthritis. J. Immunol. 184, 1552–1559 (2010).
Rutitzky, L.I., Smith, P.M. & Stadecker, M.J. T-bet protects against exacerbation of schistosome egg-induced immunopathology by regulating Th17-mediated inflammation. Eur. J. Immunol. 39, 2470–2481 (2009).
Yuan, X. et al. A novel role of CD4 Th17 cells in mediating cardiac allograft rejection and vasculopathy. J. Exp. Med. 205, 3133–3144 (2008).
Guo, S., Cobb, D. & Smeltz, R.B. T-bet inhibits the in vivo differentiation of parasite-specific CD4+ Th17 cells in a T cell-intrinsic manner. J. Immunol. 182, 6179–6186 (2009).
Durrant, D.M., Gaffen, S.L., Riesenfeld, E.P., Irvin, C.G. & Metzger, D.W. Development of allergen-induced airway inflammation in the absence of T-bet regulation is dependent on IL-17. J. Immunol. 183, 5293–5300 (2009).
Furuta, S. et al. Overlapping and distinct roles of STAT4 and T-bet in the regulation of T cell differentiation and allergic airway inflammation. J. Immunol. 180, 6656–6662 (2008).
Manoury-Schwartz, B. et al. High susceptibility to collagen-induced arthritis in mice lacking IFN-γ receptors. J. Immunol. 158, 5501–5506 (1997).
Vermeire, K. et al. Accelerated collagen-induced arthritis in IFN-γ receptor-deficient mice. J. Immunol. 158, 5507–5513 (1997).
Nistala, K. et al. Th17 plasticity in human autoimmune arthritis is driven by the inflammatory environment. Proc. Natl. Acad. Sci. USA 107, 14751–14756 (2010).
Cannon, G.W. et al. Double-blind trial of recombinant γ-interferon versus placebo in the treatment of rheumatoid arthritis. Arthritis Rheum. 32, 964–973 (1989).
Panitch, H.S., Hirsch, R.L., Schindler, J. & Johnson, K.P. Treatment of multiple sclerosis with γ interferon: exacerbations associated with activation of the immune system. Neurology 37, 1097–1102 (1987).
Skurkovich, S. et al. Randomized study of antibodies to IFN-γ and TNF-α in secondary progressive multiple sclerosis. Mult. Scler. 7, 277–284 (2001).
Ferber, I.A. et al. Mice with a disrupted IFN-γ gene are susceptible to the induction of experimental autoimmune encephalomyelitis (EAE). J. Immunol. 156, 5–7 (1996).
Willenborg, D.O., Fordham, S., Bernard, C.C., Cowden, W.B. & Ramshaw, I.A. IFN-γ plays a critical down-regulatory role in the induction and effector phase of myelin oligodendrocyte glycoprotein-induced autoimmune encephalomyelitis. J. Immunol. 157, 3223–3227 (1996).
Frisullo, G. et al. pSTAT1, pSTAT3, and T-bet expression in peripheral blood mononuclear cells from relapsing-remitting multiple sclerosis patients correlates with disease activity. J. Neurosci. Res. 84, 1027–1036 (2006).
Nos, C. et al. Clinical impact of intravenous methylprednisolone in attacks of multiple sclerosis. Mult. Scler. 10, 413–416 (2004).
Liberman, A.C. et al. The activated glucocorticoid receptor inhibits the transcription factor T-bet by direct protein-protein interaction. FASEB J. 21, 1177–1188 (2007).
Frisullo, G. et al. Glucocorticoid treatment reduces T-bet and pSTAT1 expression in mononuclear cells from relapsing remitting multiple sclerosis patients. Clin. Immunol. 124, 284–293 (2007).
Drulovic, J. et al. Expression of Th1 and Th17 cytokines and transcription factors in multiple sclerosis patients: does baseline T-bet mRNA predict the response to interferon-beta treatment? J. Neuroimmunol. 215, 90–95 (2009).
Chen, Y. et al. Anti-IL-23 therapy inhibits multiple inflammatory pathways and ameliorates autoimmune encephalomyelitis. J. Clin. Invest. 116, 1317–1326 (2006).
Yang, X.O. et al. Regulation of inflammatory responses by IL-17F. J. Exp. Med. 205, 1063–1075 (2008).
Hu, Y. et al. IL-17RC is required for IL-17A- and IL-17F-dependent signaling and the pathogenesis of experimental autoimmune encephalomyelitis. J. Immunol. 184, 4307–4316 (2010).
Yang, Y. et al. T-bet is essential for encephalitogenicity of both Th1 and Th17 cells. J. Exp. Med. 206, 1549–1564 (2009).
Reboldi, A. et al. C–C chemokine receptor 6-regulated entry of TH-17 cells into the CNS through the choroid plexus is required for the initiation of EAE. Nat. Immunol. 10, 514–523 (2009).
Kebir, H. et al. Preferential recruitment of interferon-γ-expressing TH17 cells in multiple sclerosis. Ann. Neurol. 66, 390–402 (2009).
Abromson-Leeman, S., Bronson, R.T. & Dorf, M.E. Encephalitogenic T cells that stably express both T-bet and RORγt consistently produce IFNγ but have a spectrum of IL-17 profiles. J. Neuroimmunol. 215, 10–24 (2009).
Ghoreschi, K. et al. Generation of pathogenic TH17 cells in the absence of TGF-β signalling. Nature 467, 967–971 (2010).
Chitnis, T. et al. Effect of targeted disruption of STAT4 and STAT6 on the induction of experimental autoimmune encephalomyelitis. J. Clin. Invest. 108, 739–747 (2001).
Cua, D.J. et al. Interleukin-23 rather than interleukin-12 is the critical cytokine for autoimmune inflammation of the brain. Nature 421, 744–748 (2003).
Hirota, K. et al. Fate mapping of IL-17-producing T cells in inflammatory responses. Nat. Immunol. 12, 255–263 (2011).
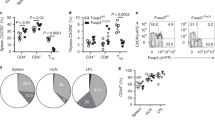
Koch, M.A. et al. The transcription factor T-bet controls regulatory T cell homeostasis and function during type 1 inflammation. Nat. Immunol. 10, 595–602 (2009).
Fontenot, J.D., Gavin, M.A. & Rudensky, A.Y. Foxp3 programs the development and function of CD4+CD25+ regulatory T cells. Nat. Immunol. 4, 330–336 (2003).
Garrett, W.S. et al. Communicable ulcerative colitis induced by T-bet deficiency in the innate immune system. Cell 131, 33–45 (2007).
Dardalhon, V. et al. IL-4 inhibits TGF-beta-induced Foxp3+ T cells and, together with TGF-β, generates IL-9+IL-10+Foxp3− effector T cells. Nat. Immunol. 9, 1347–1355 (2008).
Veldhoen, M. et al. Transforming growth factor-β 'reprograms' the differentiation of T helper 2 cells and promotes an interleukin 9–producing subset. Nat. Immunol. 9, 1341–1346 (2008).
Jager, A., Dardalhon, V., Sobel, R.A., Bettelli, E. & Kuchroo, V.K. Th1, Th17, and Th9 effector cells induce experimental autoimmune encephalomyelitis with different pathological phenotypes. J. Immunol. 183, 7169–7177 (2009).
Nurieva, R.I. et al. Generation of T follicular helper cells is mediated by interleukin-21 but independent of T helper 1, 2, or 17 cell lineages. Immunity 29, 138–149 (2008).
Nurieva, R.I. et al. Bcl6 mediates the development of T follicular helper cells. Science 325, 1001–1005 (2009).
Cruz-Guilloty, F. et al. Runx3 and T-box proteins cooperate to establish the transcriptional program of effector CTLs. J. Exp. Med. 206, 51–59 (2009).
Pearce, E.L. et al. Control of effector CD8+ T cell function by the transcription factor Eomesodermin. Science 302, 1041–1043 (2003).
Intlekofer, A.M. et al. Effector and memory CD8+ T cell fate coupled by T-bet and eomesodermin. Nat. Immunol. 6, 1236–1244 (2005).
Intlekofer, A.M. et al. Anomalous type 17 response to viral infection by CD8+ T cells lacking T-bet and eomesodermin. Science 321, 408–411 (2008).
Joshi, N.S. et al. Inflammation directs memory precursor and short-lived effector CD8+ T cell fates via the graded expression of T-bet transcription factor. Immunity 27, 281–295 (2007).
Rao, R.R., Li, Q., Odunsi, K. & Shrikant, P.A. The mTOR kinase determines effector versus memory CD8+ T cell fate by regulating the expression of transcription factors T-bet and Eomesodermin. Immunity 32, 67–78 (2010).
Oldstone, M.B., Nerenberg, M., Southern, P., Price, J. & Lewicki, H. Virus infection triggers insulin-dependent diabetes mellitus in a transgenic model: role of anti-self (virus) immune response. Cell 65, 319–331 (1991).
Ohashi, P.S. et al. Ablation of “tolerance” and induction of diabetes by virus infection in viral antigen transgenic mice. Cell 65, 305–317 (1991).
Burrell, B.E., Csencsits, K., Lu, G., Grabauskiene, S. & Bishop, D.K. CD8+ Th17 mediate costimulation blockade-resistant allograft rejection in T-bet-deficient mice. J. Immunol. 181, 3906–3914 (2008).
Rangachari, M. et al. T-bet negatively regulates autoimmune myocarditis by suppressing local production of interleukin 17. J. Exp. Med. 203, 2009–2019 (2006).
Gerth, A.J., Lin, L. & Peng, S.L. T-bet regulates T-independent IgG2a class switching. Int. Immunol. 15, 937–944 (2003).
Liu, N., Ohnishi, N., Ni, L., Akira, S. & Bacon, K.B. CpG directly induces T-bet expression and inhibits IgG1 and IgE switching in B cells. Nat. Immunol. 4, 687–693 (2003).
Werneck, M.B., Lugo-Villarino, G., Hwang, E.S., Cantor, H. & Glimcher, L.H. T-bet plays a key role in NK-mediated control of melanoma metastatic disease. J. Immunol. 180, 8004–8010 (2008).
Lugo-Villarino, G., Ito, S., Klinman, D.M. & Glimcher, L.H. The adjuvant activity of CpG DNA requires T-bet expression in dendritic cells. Proc. Natl. Acad. Sci. USA 102, 13248–13253 (2005).
Lugo-Villarino, G., Maldonado-Lopez, R., Possemato, R., Penaranda, C. & Glimcher, L.H. T-bet is required for optimal production of IFN-γ and antigen-specific T cell activation by dendritic cells. Proc. Natl. Acad. Sci. USA 100, 7749–7754 (2003).
Lipscomb, M.W. et al. Ectopic T-bet expression licenses dendritic cells for IL-12-independent priming of type 1 T cells in vitro. J. Immunol. 183, 7250–7258 (2009).
Veiga, P. et al. Bifidobacterium animalis subsp. lactis fermented milk product reduces inflammation by altering a niche for colitogenic microbes. Proc. Natl. Acad. Sci. USA 107, 18132–18137 (2010).
Garrett, W.S. et al. Colitis-associated colorectal cancer driven by T-bet deficiency in dendritic cells. Cancer Cell 16, 208–219 (2009).
Hegazy, A.N. et al. Interferons direct Th2 cell reprogramming to generate a stable GATA-3+T-bet+ cell subset with combined Th2 and Th1 cell functions. Immunity 32, 116–128 (2010).
Lee, Y.K. et al. Late developmental plasticity in the T helper 17 lineage. Immunity 30, 92–107 (2009).
Acknowledgements
Supported by the US National Institutes of Health (P01 NS038037 and CA112663 to L.H.G.), the Danone Group and the Cancer Research Institute (V.L.).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
L.H.G. is a member of the board of directors of and holds equity in Bristol Myers Squibb.
Rights and permissions
About this article
Cite this article
Lazarevic, V., Glimcher, L. T-bet in disease. Nat Immunol 12, 597–606 (2011). https://doi.org/10.1038/ni.2059
Published:
Issue Date:
DOI: https://doi.org/10.1038/ni.2059
This article is cited by
-
The effect of the anti-leukemia inhibitory factor on the immune system in the Balb/c mice bearing breast cancer induced with 4T1 cells
European Journal of Medical Research (2023)
-
Aiolos represses CD4+ T cell cytotoxic programming via reciprocal regulation of TFH transcription factors and IL-2 sensitivity
Nature Communications (2023)
-
Direct AKT activation in tumor-infiltrating lymphocytes markedly increases interferon-γ (IFN-γ) for the regression of tumors resistant to PD-1 checkpoint blockade
Scientific Reports (2022)
-
Fungal sensing by dectin-1 directs the non-pathogenic polarization of TH17 cells through balanced type I IFN responses in human DCs
Nature Immunology (2022)
-
Combined overexpression of four transcription factors promotes effector T cell dedifferentiation toward early phenotypes
Immunogenetics (2022)