Abstract
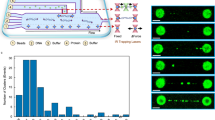
Cytoplasmic DNA triggers activation of the innate immune system. Although 'downstream' signaling components have been characterized, the DNA-sensing components remain elusive. Here we present a systematic proteomics screen for proteins that associate with DNA, 'crossed' to a screen for transcripts induced by interferon-β, which identified AIM2 as a candidate cytoplasmic DNA sensor. AIM2 showed specificity for double-stranded DNA. It also recruited the inflammasome adaptor ASC and localized to ASC 'speckles'. A decrease in AIM2 expression produced by RNA-mediated interference impaired DNA-induced maturation of interleukin 1β in THP-1 human monocytic cells, which indicated that endogenous AIM2 is required for DNA recognition. Reconstitution of unresponsive HEK293 cells with AIM2, ASC, caspase-1 and interleukin 1β showed that AIM2 was sufficient for inflammasome activation. Our data suggest that AIM2 is a cytoplasmic DNA sensor for the inflammasome.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
References
Pichlmair, A. & Reis, E.S.C. Innate recognition of viruses. Immunity 27, 370–383 (2007).
Akira, S. TLR signaling. Curr. Top. Microbiol. Immunol. 311, 1–16 (2006).
Takaoka, A. et al. DAI (DLM-1/ZBP1) is a cytosolic DNA sensor and an activator of innate immune response. Nature 448, 501–508 (2007).
Wang, Z. et al. Regulation of innate immune responses by DAI (DLM-1/ZBP1) and other DNA-sensing molecules. Proc. Natl. Acad. Sci. USA 105, 5477–5482 (2008).
Ishii, K.J. et al. TANK-binding kinase-1 delineates innate and adaptive immune responses to DNA vaccines. Nature 451, 725–729 (2008).
Ishii, K.J. et al. A Toll-like receptor-independent antiviral response induced by double-stranded B-form DNA. Nat. Immunol. 7, 40–48 (2006).
Stetson, D.B. & Medzhitov, R. Recognition of cytosolic DNA activates an IRF3-dependent innate immune response. Immunity 24, 93–103 (2006).
Petrilli, V., Dostert, C., Muruve, D.A. & Tschopp, J. The inflammasome: a danger sensing complex triggering innate immunity. Curr. Opin. Immunol. 19, 615–622 (2007).
Muruve, D.A. et al. The inflammasome recognizes cytosolic microbial and host DNA and triggers an innate immune response. Nature 452, 103–107 (2008).
Shinozuka, K., Morita, T. & Sawai, H. Synthesis and nuclease susceptibility of alpha-oligodeoxyribonucleotide phosphorothioate. Nucleic Acids Symp. Ser. 25, 101–102 (1991).
Mazur, D.J. & Perrino, F.W. Excision of 3′ termini by the Trex1 and TREX2 3′→5′ exonucleases. Characterization of the recombinant proteins. J. Biol. Chem. 276, 17022–17029 (2001).
Morita, M. et al. Gene-targeted mice lacking the Trex1 (DNase III) 3′→5′ DNA exonuclease develop inflammatory myocarditis. Mol. Cell. Biol. 24, 6719–6727 (2004).
Stetson, D.B., Ko, J.S., Heidmann, T. & Medzhitov, R. Trex1 prevents cell-intrinsic initiation of autoimmunity. Cell 134, 587–598 (2008).
Choubey, D. & Panchanathan, R. Interferon-inducible Ifi200-family genes in systemic lupus erythematosus. Immunol. Lett. 119, 32–41 (2008).
Caposio, P. et al. A novel role of the interferon-inducible protein IFI16 as inducer of proinflammatory molecules in endothelial cells. J. Biol. Chem. 282, 33515–33529 (2007).
Moriya, M. et al. Role of charged and hydrophobic residues in the oligomerization of the PYRIN domain of ASC. Biochemistry 44, 575–583 (2005).
Albrecht, M., Choubey, D. & Lengauer, T. The HIN domain of IFI-200 proteins consists of two OB folds. Biochem. Biophys. Res. Commun. 327, 679–687 (2005).
Masumoto, J., Taniguchi, S. & Sagara, J. Pyrin N-terminal homology domain- and caspase recruitment domain-dependent oligomerization of ASC. Biochem. Biophys. Res. Commun. 280, 652–655 (2001).
Natarajan, A., Ghose, R. & Hill, J.M. Structure and dynamics of ASC2, a pyrin domain-only protein that regulates inflammatory signaling. J. Biol. Chem. 281, 31863–31875 (2006).
Yu, H.B. & Finlay, B.B. The caspase-1 inflammasome: a pilot of innate immune responses. Cell Host Microbe 4, 198–208 (2008).
Hornung, V. et al. Silica crystals and aluminum salts activate the NALP3 inflammasome through phagosomal destabilization. Nat. Immunol. 9, 847–856 (2008).
Kanneganti, T.D., Lamkanfi, M. & Nunez, G. Intracellular NOD-like receptors in host defense and disease. Immunity 27, 549–559 (2007).
Johnston, J.B. et al. A poxvirus-encoded pyrin domain protein interacts with ASC-1 to inhibit host inflammatory and apoptotic responses to infection. Immunity 23, 587–598 (2005).
Dorfleutner, A. et al. A Shope Fibroma virus PYRIN-only protein modulates the host immune response. Virus Genes 35, 685–694 (2007).
Baechler, E.C. et al. Interferon-inducible gene expression signature in peripheral blood cells of patients with severe lupus. Proc. Natl. Acad. Sci. USA 100, 2610–2615 (2003).
Woerner, S.M. et al. The putative tumor suppressor AIM2 is frequently affected by different genetic alterations in microsatellite unstable colon cancers. Genes Chromosom. Cancer 46, 1080–1089 (2007).
Burckstummer, T. et al. An efficient tandem affinity purification procedure for interaction proteomics in mammalian cells. Nat. Methods 3, 1013–1019 (2006).
Irizarry, R.A. et al. Summaries of Affymetrix GeneChip probe level data. Nucleic Acids Res. 31, e15 (2003).
Acknowledgements
We acknowledge O. Hantschel and A. Pichlmair for critical reading of the manuscript, and M. Brehme for help with illustrations. Supported by the Austrian Academy of Sciences (for the Research Center for Molecular Medicine), the GenAU Program of the Austrian Federal Ministry of Science and Research (Austrian Proteomics Platform II (GZ200.145/I-VI/I/2006) and DRAGON (GZ200.142/I-VI/I/2006)), the European Commission (PIEF-GA-2008-220596 to C.B.) and the Austrian Science Fund (FWF W1205 to E.D.).
Author information
Authors and Affiliations
Contributions
T.B. did most of the experiments, developed the experimental design, supervised the project and wrote the manuscript; C.B. contributed to the cloning and did the immunofluorescence excperiments; S.B. and H.J. contributed to experiments; E.D. and G.D. did the initial screen; J.C. supervised the bioinformatics analysis; M.B. did the microarray experiments; M.P. and K.L.B. did the mass spectrometry analysis; and G.S.-F. conceived the overall strategy, cosupervised the project and contributed to writing the manuscript.
Corresponding author
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8 (PDF 1063 kb)
Rights and permissions
About this article
Cite this article
Bürckstümmer, T., Baumann, C., Blüml, S. et al. An orthogonal proteomic-genomic screen identifies AIM2 as a cytoplasmic DNA sensor for the inflammasome. Nat Immunol 10, 266–272 (2009). https://doi.org/10.1038/ni.1702
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ni.1702
This article is cited by
-
Candida albicans extracellular vesicles trigger type I IFN signalling via cGAS and STING
Nature Microbiology (2024)
-
Saharan dust induces NLRP3-dependent inflammatory cytokines in an alveolar air-liquid interface co-culture model
Particle and Fibre Toxicology (2023)
-
Assembly mechanism of the inflammasome sensor AIM2 revealed by single molecule analysis
Nature Communications (2023)
-
Mitochondrial E3 ligase MARCH5 is a safeguard against DNA-PKcs-mediated immune signaling in mitochondria-damaged cells
Cell Death & Disease (2023)
-
Effects of neutrophil fate on inflammation
Inflammation Research (2023)