Abstract
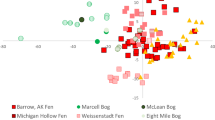
Peat bogs store up to a third of all terrestrial carbon on Earth1, and are one of the largest natural sources of atmospheric methane2. Anaerobic degradation of submerged Sphagnum species—mosses that are prevalent in peat bogs across the globe—produces significant quantities of methane in these systems. However, a study on peat mosses in the Netherlands revealed that a large fraction of this methane is consumed by aerobic methane-oxidizing bacteria, known as methanotrophs3; in return, the methanotrophs provide Sphagnum mosses with carbon3. Here, we show that Sphagnum-associated methane oxidation occurs ubiquitously across the globe. We collected Sphagnum mosses from pools, lawns and hummocks in nine Sphagnum-dominated peatlands across the world, and measured their capacity to oxidize methane in a series of laboratory incubations. All mosses were capable of oxidizing methane. The rate of methane oxidation increased with temperature, and was most pronounced in submerged mosses, collected from peatland pools. According to DNA microarray analyses, the methanotrophic community responsible for methane oxidation was highly diverse. 13C labelling revealed that methane-derived carbon was incorporated into plant lipids when mosses were submerged, indicative of a mutually beneficial symbiosis between mosses and methanotrophs. Our findings suggest that the interaction between methanotrophs and Sphagnum mosses may play a role in carbon recycling in waterlogged Sphagnum vegetation, potentially reducing methane emissions.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Smith, L. C. et al. Siberian peatlands a net carbon sink and global methane source since the early Holocene. Science 303, 353–356 (2004).
Gorham, E. Northern peatlands—role in the carbon-cycle and probable responses to climatic warming. Ecol. Appl. 1, 182–195 (1991).
Raghoebarsing, A. A. et al. Methanotrophic symbionts provide carbon for photosynthesis in peat bogs. Nature 436, 1153–1156 (2005).
Conrad, R. The global methane cycle: Recent advances in understanding the microbial processes involved. Environ. Microbiol. Rep. 1, 285–292 (2009).
Forster, P. et al. IPCC Climate Change 2007: The Physical Science Basis. Contribution of Working Group I to the Fourth Assessment Report of the Intergovernmental Panel on Climate Change (Cambridge Univ. Press, 2007).
Hanson, R. S. & Hanson, T. E. Methanotrophic bacteria. Microbiol. Rev. 60, 439–471 (1996).
Op den Camp, H. J. M. et al. Environmental, genomic and taxonomic perspectives on methanotrophic Verrucomicrobia. Environ. Microbiol. Rep. 1, 293–306 (2009).
Bodrossy, L. et al. Development and validation of a diagnostic microbial microarray for methanotrophs. Environ. Microbiol. 5, 566–582 (2003).
Bodelier, P. L. E. et al. A reanalysis of phospholipid fatty acids as ecological biomarkers for methanotrophic bacteria. ISME J. 3, 606–617 (2009).
Rohmer, M., Bouviernave, P. & Ourisson, G. Distribution of hopanoid triterpenes in prokaryotes. J. Gen. Microbiol. 130, 1137–1150 (1984).
Talbot, H. M., Watson, D. F., Murrell, J. C., Carter, J. F. & Farrimond, P. Analysis of intact bacteriohopanepolyols from methanotrophic bacteria by reversed-phase high-performance liquid chromatography-atmospheric pressure chemical ionisation mass spectrometry. J. Chromatogr. 921, 175–185 (2001).
Pancost, R. D., Baas, M., van Geel, B. & Sinninghe Damsté, J. S. Biomarkers as proxies for plant inputs to peats: An example from a sub-boreal ombrotrophic bog. Org. Geochem. 33, 675–690 (2002).
Pancost, R. D. & Sinninghe Damsté, J. S. Carbon isotopic compositions of prokaryotic lipids as tracers of carbon cycling in diverse settings. Chem. Geol. 195, 29–58 (2003).
Basiliko, N., Knowles, R. & Moore, T. R. Roles of moss species and habitat in methane oxidation in a northern peatland. Wetlands 24, 178–185 (2004).
Williams, R. T. & Crawford, R. L. Methane production in Minnesota Peatlands. Appl. Environ. Microbiol. 47, 1266–1271 (1984).
Frenzel, P. & Karofeld, E. CH4 emission from a hollow-ridge complex in a raised bog: The role of CH4 production and oxidation. Biogeochemistry 51, 91–112 (2000).
Cvejic, J. H., Bodrossy, L., Kovacs, K. L. & Rohmer, M. Bacterial triterpenoids of the hopene series from the methanotrophic bacteria Methylocaldum spp.: Phylogenetic implications and first evidence for an unsaturated aminobacteriopanepolyol. FEMS Microbiol. Lett. 182, 361–365 (2000).
Smolders, A. J. P., Tomassen, H. B. M., Pijnappel, H. W., Lamers, L. P. M. & Roelofs, J. G. M. Substrate-derived CO2 is important in the development of Sphagnum spp. New Phytol. 152, 325–332 (2001).
White, J. W. C. et al. A high-resolution record of atmospheric CO2 content from carbon isotopes in peat. Nature 367, 153–156 (1994).
Dedysh, S. Exploring methanotroph diversity in acidic northern wetlands: Molecular and cultivation-based studies. Microbiology 78, 655–669 (2009).
Chen, Y. et al. Revealing the uncultivated majority: Combining DNA stable-isotope probing, multiple displacement amplification and metagenomic analyses of uncultivated Methylocystis in acidic peatlands. Environ. Microbiol. 10, 2609–2622 (2008).
Chen, Y. et al. Diversity of the active methanotrophic community in acidic peatlands as assessed by mRNA and SIP-PLFA analyses. Environ. Microbiol. 10, 446–459 (2008).
Cebron, A. et al. Identity of active methanotrophs in landfill cover soil as revealed by DNA stable isotope probing. FEMS Microbiol. Ecol. 62, 12–23 (2007).
Vishwakarma, P. et al. Ecological and molecular analyses of the rhizospheric methanotroph community in tropical rice soil: Effect of crop phenology and land-use history. Curr. Sci. 96, 1082–1089 (2009).
Dedysh, S. N. et al. Differential detection of type II methanotrophic bacteria in acidic peatlands using newly developed 16S rRNA-targeted fluorescent oligonucleotide probes. FEMS Microbiol. Ecol. 43, 299–308 (2003).
Dedysh, S. N. Methanotrophic bacteria of acidic Sphagnum peat bogs. Microbiology 71, 638–650 (2002).
Dedysh, S. N. et al. Methylocella palustris gen. nov., sp nov., a new methane-oxidizing acidophilic bacterium from peat bogs, representing a novel subtype of serine-pathway methanotrophs. Int. J. Syst. Evol. Microbiol. 50, 955–969 (2000).
Miguez, C. B., Bourque, D., Sealy, J. A., Greer, C. W. & Groleau, D. Detection and isolation of methanotrophic bacteria possessing soluble methane monooxygenase (sMMO) genes using the polymerase chain reaction (PCR). Microb. Ecol. 33, 21–31 (1997).
McDonald, I. R., Kenna, E. M. & Murrell, J. C. Detection of methanotrophic bacteria in environmental samples with the PCR. Appl. Environ. Microbiol. 61, 116–121 (1995).
Auman, A. J., Stolyar, S., Costello, A. M. & Lidstrom, M. E. Molecular characterization of methanotrophic isolates from freshwater lake sediment. Appl. Environ. Microbiol. 66, 5259–5266 (2000).
Acknowledgements
We thank P. Wijnhoven and S. Schouten for technical assistance. C. Fritz, L. Lamers, J. van Huissteden, F. Keuper, T. Moore, N. Filippov and W. Bleuten provided Sphagnum species from different locations. N.K. and J.F.v.W. were partially supported by a grant (142.16.1060) provided by the Darwin Centre for Biogeosciences.
Author information
Authors and Affiliations
Contributions
N.K. carried out methane oxidation measurements. J.F.v.W. analysed 13C-labelled biomarkers. Y.P., L.B. and N.K. carried out the microarray experiment and analysis. The research was conceived by M.S.M.J., J.S.S.D., H.J.M.O.d.C. and G-J.R. N.K., J.F.v.W., A.J.P.S., M.S.M.J., J.S.S.D., H.J.M.O.d.C, L.B. and G-J.R. contributed to interpreting the data and writing the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Information (PDF 494 kb)
Rights and permissions
About this article
Cite this article
Kip, N., van Winden, J., Pan, Y. et al. Global prevalence of methane oxidation by symbiotic bacteria in peat-moss ecosystems. Nature Geosci 3, 617–621 (2010). https://doi.org/10.1038/ngeo939
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ngeo939
This article is cited by
-
Photosynthetic microorganisms effectively contribute to bryophyte CO2 fixation in boreal and tropical regions
ISME Communications (2022)
-
Integrating Decomposers, Methane-Cycling Microbes and Ecosystem Carbon Fluxes Along a Peatland Successional Gradient in a Land Uplift Region
Ecosystems (2022)
-
The bacterial communities of Alaskan mosses and their contributions to N2-fixation
Microbiome (2021)
-
Methane emissions from fens in Alberta’s boreal region: reference data for functional evaluation of restoration outcomes
Wetlands Ecology and Management (2020)
-
Microbial consortia including methanotrophs: some benefits of living together
Journal of Microbiology (2019)