Abstract
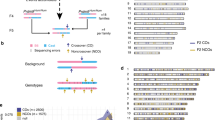
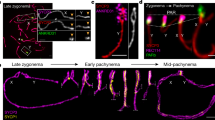
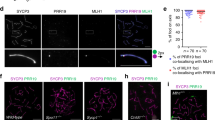
During meiosis, the reductional segregation of homologous chromosomes at the first meiotic division requires reciprocal exchange (crossing over) between homologs1. The number of crossovers is tightly regulated (one to two per homolog in mice2), and their distribution in the genome is not random—recombination 'hot' and 'cold' regions can be identified3,4. We developed a molecular assay to study these events directly in mouse germ cells. This analysis was developed with reference to the proteosome subunit β type 9 (Psmb9, previously called Lmp2) hot-spot region identified through genetic analysis5,6. Here we show that this hot spot is an initiation site of meiotic recombination on the basis of two observations: (i) crossover density is maximal in an interval of 210 bp and decreases on both sides of this region; (ii) a high frequency of gene conversion is found in the region of highest crossover density. We then used this strategy to carry out the first temporal analysis of meiotic recombination in mouse spermatogenesis and demonstrate that crossover events occur during the pachytene stage of meiotic prophase.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Change history
01 October 2002
This was incorrect in AOP and the October issue. Replaced incorrect Table A with corrected Web Table A.
References
Roeder, G.S. Meiotic chromosomes: it takes two to tango. Genes Dev. 11, 2600–2621 (1997).
Lawrie, N.M., Tease, C. & Hulten, M.A. Chiasma frequency, distribution and interference maps of mouse autosomes. Chromosoma 104, 308–314 (1995).
Lichten, M. & Goldman, A.S. Meiotic recombination hotspots. Annu. Rev. Genet. 29, 423–444 (1995).
Petes, T.D. Meiotic recombination hot spots and cold spots. Nat. Rev. Genet. 2, 360–369 (2001).
Shiroishi, T., Koide, T., Yoshino, M., Sagai, T. & Moriwaki, K. Hotspots of homologous recombination in mouse meiosis. Adv. Biophys. 31, 119–132 (1995).
Shiroishi, T. et al. Recombinational hotspot specific to female meiosis in the mouse major histocompatibility complex. Immunogenetics 31, 79–88 (1990).
Shiroishi, T., Sagai, T., Hanzawa, N., Gotoh, H. & Moriwaki, K. Genetic control of sex-dependent meiotic recombination in the major histocompatibility complex of the mouse. EMBO J. 10, 681–686 (1991).
Szostak, J.W., Orr-Weaver, T.L., Rothstein, R.J. & Stahl, F.W. The double-strand-break repair model for recombination. Cell 33, 25–35 (1983).
Goetz, P., Chandley, A.C. & Speed, R.M. Morphological and temporal sequence of meiotic prophase development at puberty in the male mouse. J. Cell Sci. 65, 249–263 (1984).
Mizuno, K., Koide, T., Sagai, T., Moriwaki, K. & Shiroishi, T. Molecular analysis of a recombinational hotspot adjacent to Lmp2 gene in the mouse MHC: fine location and chromatin structure. Mamm. Genome 7, 490–496 (1996).
Detloff, P., White, M.A. & Petes, T.D. Analysis of a gene conversion gradient at the HIS4 locus in Saccharomyces cerevisiae. Genetics 132, 113–123 (1992).
Schultes, N.P. & Szostak, J.W. Decreasing gradients of gene conversion on both sides of the initiation site for meiotic recombination at the ARG4 locus in yeast. Genetics 126, 813–822 (1990).
Borts, R.H. & Haber, J.E. Length and distribution of meiotic gene conversion tracts and crossovers in Saccharomyces cerevisiae. Genetics 123, 69–80 (1989).
Symington, L.S., Brown, A., Oliver, S.G., Greenwell, P. & Petes, T.D. Genetic analysis of a meiotic recombination hotspot on chromosome III of Saccharomyces cerevisiae. Genetics 128, 717–727 (1991).
Jeffreys, A.J., Kauppi, L. & Neumann, R. Intensely punctate meiotic recombination in the class II region of the major histocompatibility complex. Nature Genet. 29, 217–222 (2001).
Baker, S.M. et al. Involvement of mouse Mlh1 in DNA mismatch repair and meiotic crossing over. Nature Genet. 13, 336–342 (1996).
Anderson, L.K., Reeves, A., Webb, L.M. & Ashley, T. Distribution of crossing over on mouse synaptonemal complexes using immunofluorescent localization of Mlh1 protein. Genetics 151, 1569–1579 (1999).
Cobb, J., Cargile, B. & Handel, M.A. Acquisition of competence to condense metaphase I chromosomes during spermatogenesis. Dev. Biol. 205, 49–64 (1999).
Jeffreys, A.J., Neumann, R. & Wilson, V. Repeat unit sequence variation in minisatellites: a novel source of DNA polymorphism for studying variation and mutation by single molecule analysis. Cell 60, 473–485 (1990).
Peters, A.H., Plug, A.W., van Vugt, M.J. & de Boer, P. A drying-down technique for the spreading of mammalian meiocytes from the male and female germline. Chromosome Res. 5, 66–68 (1997).
Jeffreys, A.J., Ritchie, A. & Neumann, R. High resolution analysis of haplotype diversity and meiotic crossover in the human TAP2 recombination hotspot. Hum. Mol. Genet. 9, 725–733 (2000).
Acknowledgements
We thank C. Ferraz for assistance in DNA sequencing; C. Heyting and P. de Boer for advice on testis cell preparation and spreading; C. Heyting and M.A. Handel for providing Sycp3 and H1t antibodies; J. Buard for advice on ASO–PCR; T. Shiroishi for providing the mouse strains and stimulating discussions about the project; and C. Mézard, H. te Riele and C. White for comments on the manuscript. This work was supported by research grants from the Centre National de la Recherche Scientifique (ATIPE program), the Association pour la Recherche contre le Cancer, the Fondation pour la Recherche Médicale and the Commissariat à l'Energie Atomique.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
About this article
Cite this article
Guillon, H., de Massy, B. An initiation site for meiotic crossing-over and gene conversion in the mouse. Nat Genet 32, 296–299 (2002). https://doi.org/10.1038/ng990
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng990