Abstract
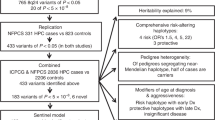
After the recent discovery that common genetic variation in 8q24 influences inherited risk of prostate cancer, we genotyped 2,973 SNPs in up to 7,518 men with and without prostate cancer from five populations. We identified seven risk variants, five of them previously undescribed, spanning 430 kb and each independently predicting risk for prostate cancer (P = 7.9 × 10−19 for the strongest association, and P < 1.5 × 10−4 for five of the variants, after controlling for each of the others). The variants define common genotypes that span a more than fivefold range of susceptibility to cancer in some populations. None of the prostate cancer risk variants aligns to a known gene or alters the coding sequence of an encoded protein.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Freedman, M.L. et al. Admixture mapping identifies 8q24 as a prostate cancer risk locus in African-American men. Proc. Natl. Acad. Sci. USA 103, 14068–14073 (2006).
Amundadottir, L.T. et al. A common variant associated with prostate cancer in European and African populations. Nat. Genet. 38, 652–658 (2006).
de Bakker, P.I. et al. Efficiency and power in genetic association studies. Nat. Genet. 37, 1217–1223 (2005).
van Duin, M. et al. High-resolution array comparative genomic hybridization of chromosome arm 8q: evaluation of genetic progression markers for prostate cancer. Genes Chromosom. Cancer 44, 438–449 (2005).
Visakorpi, T. et al. Genetic changes in primary and recurrent prostate cancer by comparative genomic hybridization. Cancer Res. 55, 342–347 (1995).
Kolonel, L.N. et al. A multiethnic cohort in Hawaii and Los Angeles: baseline characteristics. Am. J. Epidemiol. 151, 346–357 (2000).
John, E.M., Schwartz, G.G., Koo, J., Van Den Berg, D. & Ingles, S. Sun exposure, vitamin D receptor gene polymorphisms, and risk of advanced prostate cancer. Cancer Res. 65, 5470–5479 (2005).
Oakley-Girvan, I. et al. Risk of early-onset prostate cancer in relation to germline polymorphisms of the vitamin D receptor. Cancer Epidemiol. Biomarkers Prev. 13, 1325–1330 (2004).
Cooney, K.A. et al. Age-specific distribution of serum prostate-specific antigen in a community-based study of African-American men. Urology 57, 91–96 (2001).
Cooney, K.A. et al. Prostate cancer susceptibility locus on chromosome 1q: a confirmatory study. J. Natl. Cancer Inst. 89, 955–959 (1997).
The International HapMap Consortium. A haplotype map of the human genome. Nature 437, 1299–1320 (2005).
Hosono, S. et al. Unbiased whole-genome amplification directly from clinical samples. Genome Res. 13, 954–964 (2003).
Fan, J.B. et al. Highly parallel SNP genotyping. Cold Spring Harb. Symp. Quant. Biol. 68, 69–78 (2003).
Tang, K. et al. Chip-based genotyping by mass spectrometry. Proc. Natl. Acad. Sci. USA 96, 10016–10020 (1999).
Lee, L.G., Connell, C.R. & Bloch, W. Allelic discrimination by nick-translation PCR with fluorogenic probes. Nucleic Acids Res. 21, 3761–3766 (1993).
Patterson, N. et al. Methods for high-density admixture mapping of disease genes. Am. J. Hum. Genet. 74, 979–1000 (2004).
Breslow, N.E. & Day, N.E. Statistical Methods in Cancer Research, Vol 1: the Analysis of Case-Control Studies (International Agency on Cancer Research, Lyon, 1980).
Acknowledgements
We thank the men with and without prostate cancer who volunteered to participate in this study. We thank E. Lander, A. Price, J. Seidman and E. Ziv for critical comments and researchers from the NCI Cancer Genetic Markers of Susceptibility Study (CGEMS) Project, whose online data we cited to provide confirmation of the association at rs6983267. Genotyping and personnel were supported by NIH grants CA63464 to B.E.H., C.A.H., D.A., M.L.F. and D.R. and by funds from the Harvard Medical School Department of Genetics (D.R.). M.L.F. was supported by a Department of Defense Health Disparity Training-Prostate Scholar Award (17-02-1-0246) and Dana-Farber/Harvard Partners Cancer Care Prostate SPORE. N.P. was supported by NIH career transition award HG02758. S.C.G. is a Hospital for Sick Children and March of Dimes Fellow of the Pediatric Scientist Development Program, and is also supported by NICHD grant HD00850. C.A.H., D.O.S., M.C.P., L.L.M., L.N.K. and B.E.H. were supported by CA54281, E.M.J. and S.A.I. were supported by California Cancer Research Program Grants 99-00527V-10182 and 99-00524V-10258; S.A.I. was also supported by NIH grant CA84979. The Flint Men's Health Study was supported by the University of Michigan SPORE in Prostate Cancer (CA69568), the University of Michigan Department of Urology and the University of Michigan Comprehensive Cancer Center. K.A.C. was supported by grants CA69568 and CA79596, and I.O.G. and A.S.W. were supported by CA67044. D.A. is a Charles E. Culpeper Scholar of the Rockefeller Brothers Fund and a Burroughs Wellcome Fund Clinical Scholar in Translational Research. D.R. is supported by a Burroughs-Wellcome Career Development Award in the Biomedical Sciences.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Fig. 1
Proportion of genetic variation in HapMap samples captured by genotyping at 8q24. (PDF 56 kb)
Supplementary Fig. 2
Linkage disequilibrium scans across 8q24 for each of five populations. (PDF 477 kb)
Supplementary Fig. 3
Outlier removal and ancestry estimates. (PDF 114 kb)
Supplementary Table 1
Whole-genome admixture scan. (XLS 618 kb)
Supplementary Table 2
Linkage disequilibrium scan in five populations across the admixture peak. (XLS 2069 kb)
Supplementary Table 3
Phenotype and genotype data for each sample. (XLS 5846 kb)
Supplementary Table 4
Genotype data for 547 newly characterized polymorphisms in HapMap samples. (XLS 872 kb)
Supplementary Table 5
Linkage disequilibrium patterns among key polymorphisms in regions 1–3. (PDF 60 kb)
Supplementary Table 6
Recalculation of Table 2 for prospectively collected multiethnic cohort samples. (PDF 22 kb)
Supplementary Table 7
No evidence for epistatic interactions or nonmultiplicative effects among polymorphisms. (PDF 22 kb)
Supplementary Table 8
Association of variants to specific phenotypes of prostate cancer. (PDF 69 kb)
Supplementary Table 9
Polymorphisms used for outlier removal and ancestry estimates. (XLS 44 kb)
Rights and permissions
About this article
Cite this article
Haiman, C., Patterson, N., Freedman, M. et al. Multiple regions within 8q24 independently affect risk for prostate cancer. Nat Genet 39, 638–644 (2007). https://doi.org/10.1038/ng2015
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng2015
This article is cited by
-
Genetic and biological drivers of prostate cancer disparities in Black men
Nature Reviews Urology (2024)
-
Long-range gene regulation in hormone-dependent cancer
Nature Reviews Cancer (2023)
-
PRState: Incorporating genetic ancestry in prostate cancer risk scores for men of African ancestry
BMC Cancer (2022)
-
Prostate cancer risk in men of differing genetic ancestry and approaches to disease screening and management in these groups
British Journal of Cancer (2022)
-
Association of genetic variants with prostate cancer in Africa: a concise review
Egyptian Journal of Medical Human Genetics (2021)