Abstract
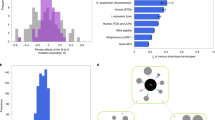
How do the fitness effects of several mutations combine? Despite its simplicity, this question is central to the understanding of multilocus evolution. Epistasis (the interaction between alleles at different loci), especially epistasis for fitness traits such as reproduction and survival, influences evolutionary predictions1,2 “almost whenever multilocus genetics matters”3. Yet very few models4,5 have sought to predict epistasis, and none has been empirically tested. Here we show that the distribution of epistasis can be predicted from the distribution of single mutation effects, based on a simple fitness landscape model6. We show that this prediction closely matches the empirical measures of epistasis that have been obtained for Escherichia coli7 and the RNA virus vesicular stomatitis virus8. Our results suggest that a simple fitness landscape model may be sufficient to quantitatively capture the complex nature of gene interactions. This model may offer a simple and widely applicable alternative to complex metabolic network models, in particular for making evolutionary predictions.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Phillips, P., Otto, S.P. & Whitlock, M.C. in Epistasis and the Evolutionary Process (eds. Wolf, J.B., Brodie, E.D. & Wade, M.J.) 20–38 (Oxford University Press, Oxford, 2000).
Otto, S.P. & Lenormand, T. Resolving the paradox of sex and recombination. Nat. Rev. Genet. 3, 252–261 (2002).
Michalakis, Y. & Roze, D. Epistasis in RNA viruses. Science 306, 1492–1493 (2004).
Szàthmary, E. Do deleterious mutations act synergistically - metabolic control-theory provides a partial answer. Genetics 133, 127–132 (1993).
Segré, D., DeLuna, A., Church, G.M. & Kishony, R. Modular epistasis in yeast metabolism. Nat. Genet. 37, 77–83 (2005).
Martin, G. & Lenormand, T. A multivariate extension of Fisher's geometrical model and the distribution of mutation fitness effects across species. Evolution 60, 893–907 (2006).
Elena, S.F. & Lenski, R.E. Test of synergistic interactions among deleterious mutations in bacteria. Nature 390, 395–398 (1997).
Sanjuàn, R., Moya, A. & Elena, S.F. The contribution of epistasis to the architecture of fitness in an RNA virus. Proc. Natl. Acad. Sci. USA 101, 15376–15379 (2004).
Malmberg, R.L. & Mauricio, R. QTL-based evidence for the role of epistasis in evolution. Genet. Res. 86, 89–95 (2005).
Burch, C.L., Turner, P.E. & Hanley, K.A. Patterns of epistasis in RNA viruses: a review of the evidence from vaccine design. J. Evol. Biol. 16, 1223–1235 (2003).
Bonhoeffer, S., Chappey, C., Parkin, N.T., Whitcomb, J.M. & Petropoulos, C.J. Evidence for positive epistasis in HIV-1. Science 306, 1547–1550 (2004).
Wloch, D.M., Borts, R.H. & Korona, R. Epistatic interactions of spontaneous mutations in haploid strains of the yeast Saccharomyces cerevisiae. J. Evol. Biol. 14, 310–316 (2001).
Fong, S.S. & Palsson, B.O. Metabolic gene-deletion strains of Escherichia coli evolve to computationally predicted growth phenotypes. Nat. Genet. 36, 1056–1058 (2004).
Fisher, R.A. The Genetical Theory of Natural Selection, (Oxford University Press, Oxford, 1930).
Orr, H.A. The genetic theory of adaptation: a brief history. Nat. Rev. Genet. 6, 119–127 (2005).
Barton, N.H. & Keightley, P.D. Understanding quantitative genetic variation. Nat. Rev. Genet. 3, 11–21 (2002).
Orr, H.A. The “sizes” of mutations fixed in phenotypic evolution: a response to Clarke and Arthur. Evol. Dev. 3, 121–123 (2001).
Waxman, D. & Welch, J.J. Fisher's microscope and Haldane's ellipse. Am. Nat. 166, 447–457 (2005).
Martin, G. & Lenormand, T. The fitness effect of mutations in stressful environments: a survey in the light of fitness landscape models. Evolution 60, 2413–2427 (2006).
Akaike, H. A new look at the statistical model identification. IEEE Trans. Automatic Control 19, 716–723 (1974).
Otto, S.P. & Feldman, M.W. Deleterious mutations, variable epistatic interactions and the evolution of recombination. Theor. Popul. Biol. 51, 134–147 (1997).
Kouyos, R.D., Otto, S.P. & Bonhoeffer, S. Effect of varying epistasis on the evolution of recombination. Genetics 173, 589–597 (2006).
Lande, R. The genetic covariance between characters maintained by pleiotropic mutations. Genetics 94, 203–215 (1980).
Burger, R. in The Mathematical Theory of Selection, Recombination, Mutation Ch. V, 158–160 (John Wiley & Sons, Chichester, UK, 2000).
Mathai, A.M. & Provost, S.B. Quadratic Forms in Random Variables (Marcel Dekker, New York, 1992).
Whitlock, M.C. & Bourguet, D. Factors affecting the genetic load in Drosophila: synergistic epistasis and correlations among fitness components. Evolution 54, 1654–1660 (2000).
Lenski, R.E. & Travisano, M. Dynamics of adaptation and diversification - a 10,000-generation experiment with bacterial-populations. Proc. Natl. Acad. Sci. USA 91, 6808–6814 (1994).
Elena, S.F., Ekunwe, L., Hajela, N., Oden, S.A. & Lenski, R.E. Distribution of fitness effects caused by random insertion mutations in Escherichia coli. Genetica 103, 349–358 (1998).
Sanjuàn, R., Moya, A. & Elena, S.F. The distribution of fitness effects caused by single-nucleotide substitutions in an RNA virus. Proc. Natl. Acad. Sci. USA 101, 8396–8401 (2004).
Kidwell, M.G. & Lisch, D. Transposable elements as sources of variation in animals and plants. Proc. Natl. Acad. Sci. USA 94, 7704–7711 (1997).
Acknowledgements
We thank D. Waxman and P. Jarne for comments on this work. G.M. thanks J. Goudet for hosting him during part of this work. This work was supported by an Action Concertée Incitative from the French Ministry of Research (T.L.), a PhD fellowship from the French Ministry of Research (G.M.), the Swiss National Science Foundation (grant 31-108194/1 to G.M.) and the Spanish Ministerio de Educación y Ciencia (MEC)-FEDER grant BMC2003-00066 to S.F.E.
Author information
Authors and Affiliations
Contributions
G.M. and T.L. designed the model, did the analysis and wrote the paper. S.F.E. compiled the data and wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Fig. 1
Power curves for the tests shown in Table 1. (PDF 134 kb)
Supplementary Fig. 2
Robustness of the model to non-additivity in the phenotypic effects of mutations. (PDF 56 kb)
Supplementary Fig. 3
Agreement of the predicted approximate distributions with simulations. (PDF 384 kb)
Supplementary Table 1
Parameter estimation for the MCMC fit. (PDF 29 kb)
Rights and permissions
About this article
Cite this article
Martin, G., Elena, S. & Lenormand, T. Distributions of epistasis in microbes fit predictions from a fitness landscape model. Nat Genet 39, 555–560 (2007). https://doi.org/10.1038/ng1998
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng1998
This article is cited by
-
Sign epistasis caused by hierarchy within signalling cascades
Nature Communications (2018)
-
Modeling genome-wide enzyme evolution predicts strong epistasis underlying catalytic turnover rates
Nature Communications (2018)
-
The utility of fitness landscapes and big data for predicting evolution
Heredity (2018)
-
Beneficial mutation-selection dynamics in finite asexual populations: a free boundary approach
Scientific Reports (2017)
-
Causes of molecular convergence and parallelism in protein evolution
Nature Reviews Genetics (2016)