Abstract
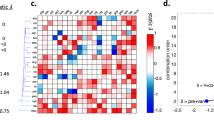
Multidrug treatments are increasingly important in medicine and for probing biological systems1,2,3,4,5,6. Although many studies have focused on interactions between specific drugs, little is known about the system properties of a full drug interaction network6. Like their genetic counterparts, two drugs may have no interaction, or they may interact synergistically or antagonistically to increase or suppress their individual effects. Here we use a sensitive bioluminescence technique7,8 to provide quantitative measurements of pairwise interactions among 21 antibiotics that affect growth rate in Escherichia coli. We find that the drug interaction network possesses a special property: it can be separated into classes of drugs such that any two classes interact either purely synergistically or purely antagonistically. These classes correspond directly to the cellular functions affected by the drugs. This network approach provides a new conceptual framework for understanding the functional mechanisms of drugs and their cellular targets and can be applied in systems intractable to mutant screening, biochemistry or microscopy.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Walsh, C. Molecular mechanisms that confer antibacterial drug resistance. Nature 406, 775–781 (2000).
Hurley, L.H. DNA and its associated processes as targets for cancer therapy. Nat. Rev. Cancer 2, 188–200 (2002).
Leeb, M. Antibiotics: a shot in the arm. Nature 431, 892–893 (2004).
Levy, S.B. & Marshall, B. Antibacterial resistance worldwide: causes, challenges and responses. Nat. Med. 10, S122–S129 (2004).
Nathan, C. Antibiotics at the crossroads. Nature 431, 899–902 (2004).
Keith, C.T., Borisy, A.A. & Stockwell, B.R. Multicomponent therapeutics for networked systems. Nat. Rev. Drug Discov. 4, 71–78 (2005).
Bjarnason, J., Southward, C.M. & Surette, M.G. Genomic profiling of iron-responsive genes in Salmonella enterica serovar typhimurium by high-throughput screening of a random promoter library. J. Bacteriol. 185, 4973–4982 (2003).
Kishony, R. & Leibler, S. Environmental stresses can alleviate the average deleterious effect of mutations. J. Biol. 2, 14 (2003).
Schreiber, S.L. The small-molecule approach to biology. Chem. Eng. News 81, 51–61 (2003).
Hartman, J.L., Garvik, B. & Hartwell, L. Cell biology - Principles for the buffering of genetic variation. Science 291, 1001–1004 (2001).
Tong, A.H. et al. Global mapping of the yeast genetic interaction network. Science 303, 808–813 (2004).
Davierwala, A.P. et al. The synthetic genetic interaction spectrum of essential genes. Nat. Genet. 37, 1147–1152 (2005).
Loewe, S. The problem of synergism and antagonism of combined drugs. Arzneimittel-Forschung-Drug Research 3, 285–290 (1953).
Bliss, C.I. The toxicity of poisons applied jointly. Ann. Appl. Biol. 26, 585–615 (1939).
Parsons, A.B. et al. Integration of chemical-genetic and genetic interaction data links bioactive compounds to cellular target pathways. Nat. Biotechnol. 22, 62–69 (2004).
Segre, D., DeLuna, A., Church, G.M. & Kishony, R. Modular epistasis in yeast metabolism. Nat. Genet. 37, 77–83 (2005).
Borisy, A.A. et al. Systematic discovery of multicomponent therapeutics. Proc. Natl. Acad. Sci. USA 100, 7977–7982 (2003).
Scott, G.M. & Kyi, M.S. Handbook of Essential Antibiotics (Harwood Academic, Amsterdam, 2001).
Walsh, C. Antibiotics: Actions, Origins, Resistance (American Society for Microbiology, Washington, D.C., 2003).
Hoffman, L.R. et al. Aminoglycoside antibiotics induce bacterial biofilm formation. Nature 436, 1171–1175 (2005).
Perlman, Z. et al. Multidimensional drug profiling by automated microscopy. Science 306, 1194–1198 (2004).
Hecht, S.M. Bleomycin: New perspectives on the mechanism of action. J. Nat. Prod. 63, 158–168 (2000).
Gardner, T.S., di Bernardo, D., Lorenz, D. & Collins, J.J. Inferring genetic networks and identifying compound mode of action via expression profiling. Science 301, 102–105 (2003).
Swinney, D.C. Biochemical mechanisms of drug action: what does it take for success? Nat. Rev. Drug Discov. 3, 801–808 (2004).
Milo, R. et al. Network motifs: Simple building blocks of complex networks. Science 298, 824–827 (2002).
Shen-Orr, S.S., Milo, R., Mangan, S. & Alon, U. Network motifs in the transcriptional regulation network of Escherichia coli. Nat. Genet. 31, 64–68 (2002).
Barabasi, A.L. & Oltvai, Z.N. Network biology: Understanding the cell's functional organization. Nat. Rev. Genet. 5, 101–113 (2004).
Remold, S.K. & Lenski, R.E. Pervasive joint influence of epistasis and plasticity on mutational effects in Escherichia coli. Nat. Genet. 36, 423–426 (2004).
Balaban, N.Q., Merrin, J., Chait, R., Kowalik, L. & Leibler, S. Bacterial persistence as a phenotypic switch. Science 305, 1622–1625 (2004).
Komarova, N.L. & Wodarz, D. Drug resistance in cancer: Principles of emergence and prevention. Proc. Natl. Acad. Sci. USA 102, 9714–9719 (2005).
Acknowledgements
We thank N. Barkai, J. Clardy, A. De Luna, M. Elowitz, L. Garwin, M. Hegreness, H. Hofmann, D. Kahne, G. Lahav, M. Laub, T. Mitchison, A. Murray, E. O'Shea, S. Renn, V. Savage, D. Segrè, N. Shoresh and C. Walsh for helpful suggestions and for comments on the manuscript. We acknowledge support from the Bauer Center for Genomics Research.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Fig. 1
Reproducibility of growth rate measurements using the bioluminescence technique. (PDF 40 kb)
Supplementary Fig. 2
Single-drug dose-reponse measurements. (PDF 52 kb)
Supplementary Fig. 3
Examining dose dependence drug interactions. (PDF 57 kb)
Supplementary Fig. 4
Monochromaticity exhibited by the drug network is a special property not exhibited in random networks. (PDF 21 kb)
Rights and permissions
About this article
Cite this article
Yeh, P., Tschumi, A. & Kishony, R. Functional classification of drugs by properties of their pairwise interactions. Nat Genet 38, 489–494 (2006). https://doi.org/10.1038/ng1755
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng1755
This article is cited by
-
Knowledge integration and decision support for accelerated discovery of antibiotic resistance genes
Nature Communications (2022)
-
Antibiotic combinations reduce Staphylococcus aureus clearance
Nature (2022)
-
The physiology and genetics of bacterial responses to antibiotic combinations
Nature Reviews Microbiology (2022)
-
Investigation of the effects of mTOR inhibitors rapamycin and everolimus in combination with carboplatin on canine malignant melanoma cells
BMC Veterinary Research (2021)
-
A multi-scale pipeline linking drug transcriptomics with pharmacokinetics predicts in vivo interactions of tuberculosis drugs
Scientific Reports (2021)