Abstract
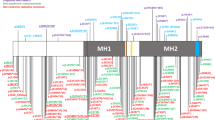
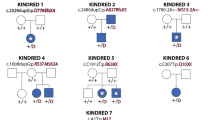
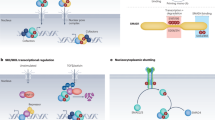
We report heterozygous mutations in the genes encoding either type I or type II transforming growth factor β receptor in ten families with a newly described human phenotype that includes widespread perturbations in cardiovascular, craniofacial, neurocognitive and skeletal development. Despite evidence that receptors derived from selected mutated alleles cannot support TGFβ signal propagation, cells derived from individuals heterozygous with respect to these mutations did not show altered kinetics of the acute phase response to administered ligand. Furthermore, tissues derived from affected individuals showed increased expression of both collagen and connective tissue growth factor, as well as nuclear enrichment of phosphorylated Smad2, indicative of increased TGFβ signaling. These data definitively implicate perturbation of TGFβ signaling in many common human phenotypes, including craniosynostosis, cleft palate, arterial aneurysms, congenital heart disease and mental retardation, and suggest that comprehensive mechanistic insight will require consideration of both primary and compensatory events.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Change history
03 February 2005
The full text and PDF were updated to contain the correct wording of sentence 5 in paragraph 8.
Notes
*Note: In the version of this article initially published online, an error appeared in paragraph 8. The paragraph should read, "Some individuals with mutations in TGFBR1 or TGFBR2 have phenotypes that overlap somewhat with MFS, but none met the diagnostic criteria for MFS15. All affected individuals had manifestations in multiple organ systems that are not associated with MFS (Table 2). In these individuals, aneurysms tend to be particularly aggressive and to rupture at an early age (e.g., individuals 4.II-1 and 5.II-1) or be of a size not associated with high risk in MFS16 (e.g., individual 1.II-1). Therefore, from a management perspective, the distinction from MFS is neither ambiguous nor unimportant. Comprehensive mutation analysis of fibrillin-1 in 93 consecutive individuals referred with classic MFS identified 86 mutations17. We sequenced TGFBR1 and TGFBR2 in the other seven individuals and found no mutations (data not shown)." This mistake has been corrected for the HTML and print versions of the article.
References
Annes, J.P., Munger, J.S. & Rifkin, D.B. Making sense of latent TGFbeta activation. J. Cell Sci. 116, 217–224 (2003).
ten Dijke, P. & Hill, C.S. New insights into TGF-beta-Smad signalling. Trends Biochem. Sci. 29, 265–273 (2004).
Cohen, M.M. Jr. TGF beta/Smad signaling system and its pathologic correlates. Am. J. Med. Genet. 116A, 1–10 (2003).
Mizuguchi, T. et al. Heterozygous TGFBR2 mutations in Marfan syndrome. Nat. Genet. 36, 855–860 (2004).
Dietz, H.C. et al. Marfan syndrome caused by a recurrent de novo missense mutation in the fibrillin gene. Nature 352, 337–339 (1991).
Neptune, E.R. et al. Dysregulation of TGF-beta activation contributes to pathogenesis in Marfan syndrome. Nat. Genet. 33, 407–411 (2003).
Ng, C.M. et al. TGFbeta-dependent pathogenesis of mitral valve prolapse in a mouse model of Marfan syndrome. J. Clin. Invest. 114, 1586–1592 (2004).
Azhar, M. et al. Transforming growth factor beta in cardiovascular development and function. Cytokine Growth Factor Rev. 14, 391–407 (2003).
Sanford, L.P. et al. TGFbeta2 knockout mice have multiple developmental defects that are non-overlapping with other TGFbeta knockout phenotypes. Development 124, 2659–2670 (1997).
Ito, Y. et al. Conditional inactivation of Tgfbr2 in cranial neural crest causes cleft palate and calvaria defects. Development 130, 5269–5280 (2003).
Grady, W.M. et al. Mutational inactivation of transforming growth factor beta receptor type II in microsatellite stable colon cancers. Cancer Res. 59, 320–324 (1999).
Wieser, R., Wrana, J.L. & Massague, J. GS domain mutations that constitutively activate T beta R-I, the downstream signaling component in the TGF-beta receptor complex. EMBO J. 14, 2199–2208 (1995).
Verrecchia, F., Chu, M.L. & Mauviel, A. Identification of novel TGF-beta/Smad gene targets in dermal fibroblasts using a combined cDNA microarray/promoter transactivation approach. J. Biol. Chem. 276, 17058–17062 (2001).
Leask, A. et al. The control of ccn2 (ctgf) gene expression in normal and scleroderma fibroblasts. Mol. Pathol. 54, 180–183 (2001).
De Paepe, A., Devereux, R.B., Dietz, H.C., Hennekam, R.C. & Pyeritz, R.E. Revised diagnostic criteria for the Marfan syndrome. Am. J. Med. Genet. 62, 417–426 (1996).
Kouchoukos, N.T. & Dougenis, D. Surgery of the thoracic aorta. N. Engl. J. Med. 336, 1876–188 (1997).
Loeys, B. et al. Comprehensive molecular screening of the FBN1 gene favors locus homogeneity of classical Marfan syndrome. Hum. Mutat. 24, 140–146 (2004).
Greally, M.T. et al. Shprintzen-Goldberg syndrome: a clinical analysis. Am. J. Med. Genet. 76, 202–212 (1998).
Charbonneau, N.L., Ono, R.N., Corson, G.M., Keene, D.R. & Sakai, L.Y. Fine tuning of growth factor signals depends on fibrillin microfibril networks. Birth Defects Res. Part C Embryo Today 72, 37–50 (2004).
Isogai, Z. et al. Latent transforming growth factor beta-binding protein 1 interacts with fibrillin and is a microfibril-associated protein. J. Biol. Chem. 278, 2750–2757 (2003).
Arteaga-Solis, E. et al. Regulation of limb patterning by extracellular microfibrils. J. Cell. Biol. 154, 275–281 (2001).
Kaartinen, V. & Warburton, D. Fibrillin controls TGF-beta activation. Nat. Genet. 33, 331–332 (2003).
de Caestecker, M. The transforming growth factor-beta superfamily of receptors. Cytokine Growth Factor Rev. 15, 1–11 (2004).
Bottinger, E.P. et al. Expression of a dominant-negative mutant TGF-beta type II receptor in transgenic mice reveals essential roles for TGF-beta in regulation of growth and differentiation in the exocrine pancreas. EMBO J. 16, 2621–2633 (1997).
Denton, C.P. et al. Fibroblast-specific expression of a kinase-deficient type II transforming growth factor beta (TGFbeta) receptor leads to paradoxical activation of TGFbeta signaling pathways with fibrosis in transgenic mice. J. Biol. Chem. 278, 25109–25119 (2003).
Koli, K. et al. Disruption of LTBP-4 function reduces TGF-{beta} activation and enhances BMP-4 signaling in the lung. J. Cell. Biol. 167, 123–133 (2004).
Acknowledgements
We thank the families for their interest and cooperation; S. Kahler, C. Scott and H. Collmann for referring affected individuals; M. Awad for assistance in preparing the figures; and D. Devos and J. Bauer for radiological support. This work was supported by the National Marfan Foundation, the William Smilow Center for Marfan Syndrome Research, the Howard Hughes Medical Institute, the Robert Wood Johnson Foundation, the Dana and Albert Broccoli Center for Aortic Diseases, the Bijzonder Onderzoeksfonds of Ghent University, the Institute for the Promotion of Innovation by Science and Technology in Flanders, the National Institutes of Health and the Fund for Scientific Research-Flanders.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Fig. 1
Sequencing of 9 subcloned RT-PCR amplicons from individual 5.II-1 that span the exon 1-2 junction in TGFBR2 mRNA revealed altered splicing in 4. (PDF 43 kb)
Rights and permissions
About this article
Cite this article
Loeys, B., Chen, J., Neptune, E. et al. A syndrome of altered cardiovascular, craniofacial, neurocognitive and skeletal development caused by mutations in TGFBR1 or TGFBR2. Nat Genet 37, 275–281 (2005). https://doi.org/10.1038/ng1511
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng1511
This article is cited by
-
The roles and regulatory mechanisms of TGF-β and BMP signaling in bone and cartilage development, homeostasis and disease
Cell Research (2024)
-
Diagnosis and treatment of cardiovascular disease in patients with heritable connective tissue disorders or heritable thoracic aortic diseases
Cardiovascular Intervention and Therapeutics (2024)
-
Severity of Autism Spectrum Disorder Symptoms Associated with de novo Variants and Pregnancy-Induced Hypertension
Journal of Autism and Developmental Disorders (2024)
-
Truncating variants in the penultimate exon of TGFBR1 escaping nonsense-mediated mRNA decay cause Loeys-Dietz syndrome
European Journal of Human Genetics (2023)
-
Genetics and mechanisms of thoracic aortic disease
Nature Reviews Cardiology (2023)