Abstract
Mutations in PRKCSH, encoding the β-subunit of glucosidase II, an N-linked glycan-processing enzyme in the endoplasmic reticulum (ER), cause autosomal dominant polycystic liver disease. We found that mutations in SEC63, encoding a component of the protein translocation machinery in the ER, also cause this disease. These findings are suggestive of a role for cotranslational protein-processing pathways in maintaining epithelial luminal structure and implicate noncilial ER proteins in human polycystic disease.
Similar content being viewed by others
Main
Polycystic liver disease often occurs in association with autosomal dominant polycystic kidney disease (ADPKD; OMIM 173900 and OMIM 173910), but it also exists as isolated autosomal dominant polycystic liver disease (ADPLD; OMIM 174050). Mutations in PRKCSH1,2 cause ADPLD and there is evidence of genetic heterogeneity3. Of 66 unrelated individuals affected with ADPLD4,5, mutations in PRKCSH were excluded in 57 probands by direct sequencing. Of these, 10 individuals belonged to families with multiple affected individuals. Four of the five largest kindreds (Supplementary Fig. 1 online) were previously reported: families A-6 (ref. 6), B-7 (ref. 7), F-1 and F-4 (ref. 3). We carried out a genome-wide analysis for linkage at ∼10-cM resolution in all ten families (comprising 43 affected individuals, 18 unaffected individuals and 2 spouses) by genotyping 394 microsatellite markers.
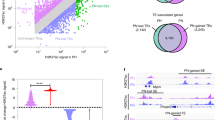
We set the allele frequency for the gene underlying ADPLD to 0.0002, the phenocopy rate to 0.01 and the heterozygous disease penetrance to 95% (ref. 4). We assigned all phenotypic data before genotyping. We calculated lod scores for the entire marker set using MLINK and LODSCORE for two-point analysis and GENEHUNTER (version 2.1) for multipoint analysis (Fig. 1a and Supplementary Table 1 online). We used the suggested genome-wide significance threshold for parametric analyses with many independent families, lod >3.3, to establish linkage.
(a) Multipoint analysis with 16 markers on chromosome 6. The 1-lod support interval (red line) is flanked by D6S1021 and D6S474. (b) Haplotype analysis of recombinant chromosomes for families A-6 (Zmax = 2.3), F-4 (Zmax = 1.8) and B-7 (Zmax = 1.5), where a lod score >1.5 yields a conditional probability of linkage of >99% for α = 0.81. Complete pedigrees are shown in Supplementary Figure 1 online. The disease-associated haplotype (all red) is shown for each family. Critical recombinant chromosomes in four affected family members are shown with the disease-associated haplotype segments in red and the normal segments in black. The boundaries defined by recombination (D6S1021 and D6S474) correspond to the 1-lod support interval. (c–f) Identification of SEC63. (c) Genetic map of interval in chromosome 6q21–q23 showing microsatellite markers used in refining the interval. (d) Physical map of interval between D6S1021 and D6S474, showing known genes and their direction of transcription, based on Build 34 of the public domain genome sequence (MapViewer). (e) Intron-exon structure of SEC63. Exons are shown as vertical bars. (f) Schematic representation of the domain structure of SEC63 protein and the location of mutations detected in individuals with ADPLD. SEC63 is an integral protein of the ER membrane with three transmembrane spans (TM1–TM3), a luminal N terminus, a cytoplasmic C terminus containing a SEC63 domain (yellow) with a coiled-coil region (CC, green) and a DnaJ domain (red) between TM2 and TM3 facing the ER luminal aspect. Within the cotranslational pathway, the DnaJ domain of SEC63 acts as a docking site localizing BiP to the luminal exit site of the Sec translocon11,12. The C terminus is thought to physically interact with other Sec translocon components.
Only one region of the genome, on chromosome 6q, met this criterion, yielding a maximum multipoint lod score, Zmax, of 6.0 (Fig. 1a). The 1-lod support interval among the ten families was ∼7 cM, corresponding to ∼8 Mb on the physical map between D6S1021 and D6S474 (Fig. 1a). Under models of heterogeneity, we obtained a maximum multipoint lod score of 6.4 at α = 0.81 with the same 1-lod support interval (data not shown). The genetic interval between D6S1021 and D6S474 was also supported by haplotype analysis in the three families with the highest individual multipoint lod scores (Fig. 1b).
Examination of the annotated sequence in the human MapViewer identified ∼39 genes and a number of putative open reading frames or hypothetical genes (Fig. 1c–f). We initially focused on genes that are expressed in liver tissue (as shown by RT-PCR) and that may be functionally linked to PRKCSH through a role in protein maturation in the ER. SEC63 met these criteria. We screened the 21-exon coding sequence and flanking splice sequences of SEC63 by direct sequencing of amplified genomic PCR products (primer sequences on request). We detected seven heterozygous sequence variants in 8 of 57 probands, including the five families in whom we found the highest lod scores with chromosome 6 markers (Table 1 and Supplementary Fig. 2 online). Mutations were located throughout the gene from exon 2 to exon 19 (Fig. 1c–f). We found two insertion-deletion mutations resulting in frameshifts with premature chain termination, two nonsense codon mutations and two mutations predicted to disrupt splice donor-acceptor sites. These variants were not found in 192 normal chromosomes. The final variant, 1702delGAA, resulted in the in-frame deletion of one of three successive glutamic acid residues (amino acids 566–568). This variant was not found in 360 normal chromosomes and is probably pathogenic.
Mutation 173G→A, resulting in a nonsense codon W58X, occurred in two probands from the central US who were not known to be related. Three probands from Finland (in families F-1, F-4 and F-226) had different mutations. The unique nature of most mutations is consistent with the idea that mutations in SEC63 that cause ADPLD, like those in PRKCSH1,2 and in the genes associated with polycystic kidney disease8,9,10, probably arose independently. We found an additional 11 sequence variants, 7 of which result in amino acid substitutions, in samples from both affected individuals and controls (Supplementary Table 2 online). Although cysts occur only in the liver, Sec63 was expressed in all tissues tested (Supplementary Fig. 3 online). Expression in the liver was roughly two times higher than in kidney and testis when densitometrically normalized to β-actin loading (Supplementary Fig. 3 online).
In summary, we identified a second gene associated with ADPLD. We found mutations in SEC63 in 8 of 66 probands (∼12%) in our sample that included both familial cases and individual probands not known to have a family history of ADPLD. Mutations in PRKCSH and SEC63 together account for less than one-third of ADPLD cases in this cohort, indicating that there is at least one more locus associated with this disease.
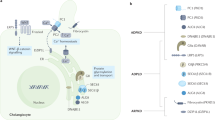
SEC63 encodes an integral membrane protein of the ER that is highly conserved from yeast to man. It is part of the multicomponent translocon that comprises the protein translocation machinery for integral membrane and secreted proteins. There are two targeting pathways to the Sec translocons: the cotranslational or signal recognition particle (SRP)-dependent pathway and the post-translational or SRP-independent pathway (reviewed in ref. 11). SEC63 is required in both post-translational and cotranslational pathways12. The cotranslational pathway, in which the ribosome is directly complexed with the Sec translocon and extrudes the nascent peptide through it, is the main pathway in mammalian cells, including lumen-forming epithelia such as the bile duct. Cotranslational maturation events include signal peptide cleavage, transfer and trimming of N-linked glycans, disulfide bond formation, transmembrane domain integration, chaperone binding and protein folding11,13. The transfer and trimming of N-glycans notably involves the activity of glucosidase II (GII), the β subunit of which, PRKCSH, was the first gene found to be associated with ADPLD. PRKCSH-dependent GII activity promotes proper folding and maturation of glycoproteins in the calnexin-calreticulin cycle14, a process that occurs immediately downstream of the translocon. This is the functional link between the two genes known to be associated with ADPLD11. Other genes involved in these processes are functional candidates for ADPLD.
If ADPLD, like ADPKD, occurs by a cellular recessive, two-hit mechanism, then mutations in either SEC63 or PRKCSH will result in loss of proper folding of integral membrane or secreted glycoproteins in bile duct cells that have undergone somatic second hits. Proteins that do not fold properly are targeted for degradation14. One possible molecular link among polycystic diseases may be that client proteins for SEC63 and GIIβ include cilial components such as polycystin-1, polycystin-2 or polyductin (also called fibrocystin). Somatic loss of the ER polycystic proteins GIIβ or SEC63 results in functional loss of one or more of the cilial polycystic proteins. A corollary to this would be that a substantial proportion of pathogenic amino acid substitution mutations seen with high frequency in polycystin-1 (ref. 9) and polyductin8 may be trafficking mutations rather than loss-of-function mutations. The lack of an observed abnormal kidney phenotype in ADPLD may be due to the existence of alternative pathways for maturation of client proteins in that tissue15 or potential tissue-specific cellular lethality after homozygous loss of the respective gene associated with ADPLD due to somatic second hits. The identification of SEC63 as a gene underlying ADPLD implicates noncilial pathways in polycystic disease in the liver and provides a new cellular and molecular entry point to understanding human polycystic disease processes in general.
URL. MapViewer is available at http://ncbi.nih.gov/mapview/.
Note: Supplementary information is available on the Nature Genetics website.
References
Li, A. et al. Am. J. Hum. Genet. 72, 691–703 (2003).
Drenth, J.P., Te Morsche, R.H., Smink, R., Bonifacino, J.S. & Jansen, J.B. Nat. Genet 33, 345–347 (2003).
Tahvanainen, P. et al. J. Hepatol. 38, 39–43 (2003).
Reynolds, D.M. et al. Am. J. Hum. Genet. 67, 1598–1604 (2000).
Qian, Q. et al. Hepatology 37, 164–171 (2003).
Iglesias, D.M. et al. Dig. Dis. Sci. 44, 385–388 (1999).
Pirson, Y. et al. Hepatology 23, 249–252 (1996).
Furu, L. et al. J. Am. Soc. Nephrol. 14, 2004–2014 (2003).
Rossetti, S. et al. Am. J. Hum. Genet. 68, 46–63 (2001).
Wu, G. & Somlo, S. Mol. Genet. Metab. 69, 1–15 (2000).
Schnell, D.J. & Hebert, D.N. Cell 112, 491–505 (2003).
Young, B.P., Craven, R.A., Reid, P.J., Willer, M. & Stirling, C.J. EMBO J. 20, 262–271 (2001).
Daniels, R., Kurowski, B., Johnson, A.E. & Hebert, D.N. Mol. Cell 11, 79–90 (2003).
Helenius, A. & Aebi, M. Science 291, 2364–2369 (2001).
Moore, S.E. & Spiro, R.G. J. Biol. Chem. 265, 13104–13112 (1990).
Acknowledgements
We thank the affected individuals and family members for their participation; K. Cornwell and P. Urban for help with recruiting study subjects; and R. Torra, X.M. Lens, M. Ott and Y. Pei for referring study subjects. The Keck Biotechnology Resource at Yale provided automated genotyping services and the Mayo Clinic General Clinical Research Center assisted with evaluations of study subjects. P.T, E.T, H.K. and K.H. received financial support from Mary and Georg C. Ehrnrooth Foundation. This work was supported by the US National Institutes of Health (S.S. and V.E.T.). S.S. is a member of the Yale Digestive Diseases Research Core Center; S.D., L.F, X.T., T.O, A.L., Y.C. and S.S. are members of the Yale Center for the Study of Polycystic Kidney Disease.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Davila, S., Furu, L., Gharavi, A. et al. Mutations in SEC63 cause autosomal dominant polycystic liver disease. Nat Genet 36, 575–577 (2004). https://doi.org/10.1038/ng1357
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng1357
This article is cited by
-
LncRNA WFDC21P interacts with SEC63 to promote gastric cancer malignant behaviors by regulating calcium homeostasis signaling pathway
Cancer Cell International (2024)
-
Genetics, pathobiology and therapeutic opportunities of polycystic liver disease
Nature Reviews Gastroenterology & Hepatology (2022)
-
Expanding the variability of the ADPKD-GANAB clinical phenotype in a family of Italian ancestry
Journal of Nephrology (2022)
-
Polycystic liver disease with lethal abdominal wall rupture: a case report
Journal of Medical Case Reports (2021)
-
Pathobiology of inherited biliary diseases: a roadmap to understand acquired liver diseases
Nature Reviews Gastroenterology & Hepatology (2019)