Abstract
Remodeling of the cytoskeleton is central to the modulation of cell shape and migration. Filamin A, encoded by the gene FLNA, is a widely expressed protein that regulates re-organization of the actin cytoskeleton by interacting with integrins, transmembrane receptor complexes and second messengers1,2. We identified localized mutations in FLNA that conserve the reading frame and lead to a broad range of congenital malformations, affecting craniofacial structures, skeleton, brain, viscera and urogenital tract, in four X-linked human disorders: otopalatodigital syndrome types 1 (OPD1; OMIM 311300) and 2 (OPD2; OMIM 304120), frontometaphyseal dysplasia (FMD; OMIM 305620) and Melnick–Needles syndrome (MNS; OMIM 309350). Several mutations are recurrent, and all are clustered into four regions of the gene: the actin-binding domain and rod domain repeats 3, 10 and 14/15. Our findings contrast with previous observations that loss of function of FLNA is embryonic lethal in males but manifests in females as a localized neuronal migration disorder, called periventricular nodular heterotopia (PVNH; refs. 3–6). The patterns of mutation, X-chromosome inactivation and phenotypic manifestations in the newly described mutations indicate that they have gain-of-function effects, implicating filamin A in signaling pathways that mediate organogenesis in multiple systems during embryonic development.
Similar content being viewed by others
Main
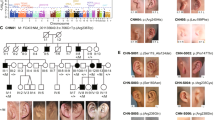
We previously described linkage of a kindred segregating OPD2 to a 2.1-Mb interval in distal Xq28 (refs. 7,8) that coincided with a broader 6-Mb localization for the related condition OPD1 (refs. 9,10). Clinically, OPD2 manifests with abnormally modeled, bowed bones, small or absent first digits and, more variably, cleft palate, posterior fossa brain anomalies, omphalocele and cardiac defects (refs. 11,12; Fig. 1a–c). Males with OPD1 have cleft palate, malformations of the ossicles causing deafness and milder bone and limb defects than those associated with OPD2 (refs. 13,14; Fig. 1d,e). Obligate female carriers of mutations causing both OPD1 and OPD2 have variable (often milder) expression of a similar phenotypic spectrum. Two other disorders, FMD and MNS, have not been conclusively localized genetically but have features that overlap clinically with the OPD syndromes12. Males with FMD have supraorbital hyperostosis, deafness and digital anomalies (ref. 15; Fig. 1f,g); females with MNS have an osteodysplasia (ref. 16; Fig. 1h–j); and males with MNS have a lethal phenotype indistinguishable from severe OPD2 (ref. 17). We developed two additional polymorphic markers to reduce the candidate interval in the family that showed linkage to OPD2 (refs. 7,8) to 1.95 Mb, bounded by IRAK1 proximally and the Xq telomere distally (data not shown).
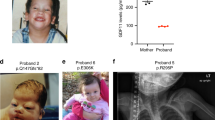
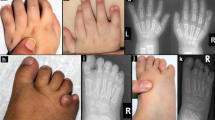
a–c, OPD2 in the male presents with camptodactyly, bowed bones (campomelia), thoracic hypoplasia, omphalocoele and hypoplasia of the thumbs and great toes (individuals are from refs. 7,8). d,e, A male with OPD1 (individual 2042) who has conductive deafness and a repaired cleft palate, showing the prominent brow (supraorbital hyperostosis; d), spatulate fingers with short first digits and hypoplastic terminal phalanges (e). f,g, A carrier mother and her son (individual 2436) with FMD, showing prominence of the brow and hypertelorism. Skull radiograph of the son (g) shows thickening of the skull vault, supraorbital hyperostosis and underdeveloped frontal sinuses. Images in f and g reproduced with permission from Wiley-Liss. h–j, Females with MNS have prominent eyes, full cheeks, small chin (individual 2239) and bones with an irregular contour resulting in deformities such as thoracic hypoplasia and bowed long bones (individual 2819).
We screened candidate genes in this interval using single-strand conformation polymorphism analysis and Southern blotting in a multi-ethnic panel of 41 unrelated individuals with diverse phenotypes (6 with OPD1, 13 with OPD2, 7 with FMD, 10 with MNS and 5 atypical). After excluding 24 other genes, we examined FLNA. Mutations in FLNA (predominantly nonsense and frameshift mutations but including two missense mutations) lead to the clinically unrelated neuronal migration disorder PVNH3,5,6. FLNA comprises 48 exons and encodes a protein of 280 kDa that possesses an N-terminal actin-binding domain (ABD) containing two calponin homology domains (CHD1 and CHD2) and an extended region made up of 24 repeated rod subdomains that bind to multiple proteins (refs. 1,2; Fig. 2a). Filamin A associates as a homodimer and may heterodimerize with filamin B, the product of a paralogous gene, FLNB18.
a, Cartoon of a filamin A dimer (left) showing the primary structural motifs. Regions containing pathogenic substitutions are circled and are presented in more detail on the right. b, Sequence trace from an affected male (marked with an asterisk in the pedigree below) and segregation of the recurrent 760G→A mutation in the family with OPD2 used for the linkage analysis7,8. The mutation destroys a Bsp1286I site. Fragment sizes are indicated in base pairs. c, Sequence trace from an affected male (marked with an asterisk in the pedigree below) and segregation of the recurrent 620C→T mutation in a family segregating OPD1 (ref. 14). A new EcoN1 restriction site is created. Fragment sizes are indicated in base pairs. Dots within symbols indicate obligate or clinically manifesting female carriers. d, Sequence trace from a female (marked with an asterisk in the pedigree below) affected with MNS heterozygous with respect to the 3562G→A mutation compared with a normal control; the mutation creates an additional Bsp1286I restriction site. Below, segregation of new Bsp1286I fragment (160 bp) in familial MNS (ref. 28; lanes 1–9) and two sporadic cases (lanes 12,13), one of which is shown to have arisen de novo (lanes 10–12). Fragment sizes are indicated in base pairs. N, normal control. e, Sequence trace from a female (marked with an asterisk in the pedigree below) with MNS heterozygous with respect to the 3596C→T mutation compared with a normal control. Below, allele-specific oligonucleotide (ASO) hybridization showing that the mutation has occurred de novo in one individual (lane 3) and in three other sporadic cases (lanes 4–6) but not in an unaffected sibling (lane 7) or a control individual (lane 8). N, normal control.
Black and gray backgrounds indicate high and intermediate levels of conservation, respectively. The distal portion of CHD1 and helices A, C, E and G in the tertiary structure of the CHD2 are indicated by blue bars. Sites where missense mutations lead to the OPD-spectrum disorders (red), PVNH (green), Duchenne and Becker muscular dystrophy (yellow), hereditary spherocytosis (blue) and focal segmental glomerulosclerosis (magenta) are indicated with color. The specific substitutions identified in the OPD-spectrum disorders are indicated above their corresponding position in filamin A. The pair of asterisks indicate residues participating in a conserved salt bridge.
We examined all coding exons of FLNA by denaturing high-performance liquid chromatography (dHPLC). We identified 15 different mutations in four discrete regions of FLNA (Table 1 and Fig. 2a) in 26 members of the primary screening panel (3 with OPD1, 10 with OPD2, 3 with FMD and 10 with MNS). We later found four additional mutations in a focused screen of the four mutation hotspots. Overall, we identified 17 distinct mutations in 30 unrelated individuals.
All mutations leading to OPD1 and OPD2 in males were located in the CHD2 portion of the ABD. A recurrent mutation, 760G→A (resulting in the amino-acid substitution E254K), accounted for all families segregating OPD2 with omphalocele (n = 4), including the family in which we initially identified linkage of OPD2 to distal Xq28 (refs. 7,8; Fig. 2b). We identified another recurrent mutation, 620C→T (resulting in the amino-acid substitution P207L), in two large families14,19 segregating OPD1 (Fig. 2c). Clustered substitutions in rod-domain repeat 10 accounted for all cases of MNS in the panel. Two substitutions, 3562G→A (resulting in the amino-acid substitution A1188T) and 3596C→T (resulting in the amino-acid substitution S1199L) were recurrent, and both were shown to have arisen de novo (Fig. 2d,e). We identified additional substitutions in rod-domain repeats 3, 10 and 14/15 in males with FMD and females with OPD2 (Table 1). Five distinct mutations were shown to have arisen de novo (Table 1); the other 12 mutations predict changes in highly conserved residues across a wide range of vertebrate and invertebrate filamins (Fig. 3 and Supplementary Fig. 1 online), segregate with the disease phenotype in a total of 59 meioses (data not shown) and were not observed in at least 100 control chromosomes. We conclude that OPD1, OPD2, FMD and MNS are allelic conditions12, which we collectively term 'OPD-spectrum disorders'. All (17 of 17) mutations reported here, in contrast with a small minority (2 of 14) associated with PVNH3,5,6, conserve the reading frame and are predicted to produce full-length filamin A.
We compared the structure of filamin A and the distribution of mutations in the ABD with selected proteins containing a homologous domain (Fig. 3). Human disorders have been identified in association with missense mutations in the ABDs of three other proteins: dystrophin (Duchenne and Becker muscular dystrophy), β-spectrin (hereditary spherocytosis; ref. 20) and α-actinin-4 (focal segmental glomerulosclerosis; ref. 21). The missense substitutions in these proteins cluster in the same regions as do the substitutions in filamin A (Fig. 3). We modeled the tertiary structure of the CHD2 domain of filamin A based on sequence homology to the previously obtained structures of the CHD2 in dystrophin22 and β-spectrin23. Most notably, the recurrent OPD2 mutation E254K is predicted to disrupt a highly conserved salt bridge with residue Lys169 (Figs. 3,4). All substitutions (except L172F) lie on one face of the domain (Fig. 4), and those within helix A map to a region previously implicated in actin binding22. Similarly distributed mutations in the dystrophin CHD2 are associated with residual function (Becker phenotype), and those in α-actinin-4 were proposed to enhance actin binding21. Collectively, these observations suggest that mutations in FLNA affecting the CHD2 domain do not result in simple loss of function.
Helices A, C, E and G are specifically identified and the predicted sites of the substituted residues in filamin A are identified by gray spheres (α-carbon atom). The side chains of residues 169 and 254 predicted to form a salt bridge in wild-type filamin A, which would be destroyed by the E254K substitution, are indicated in purple. Drawn with BOBSCRIPT29.
The substitutions in the rod repeats of filamin A are also highly non-random. Interspecies sequence comparisons around mutation hotspots show high sequence conservation among vertebrates and moderate conservation in more distantly related phyla, especially at the sites recurrently mutated in rod-domain repeat 10 (see Supplementary Fig. 1 online). Sequence alignment and comparison with the structure of Dictyostelium rod-domain repeat 5 (ref. 24) suggests that most of these mutations should not grossly disrupt the structure. Repeats 15–23 recruit multiple proteins to the cytoskeleton, but no binding partners have yet been identified for repeat 10; we speculate that these mutations alter binding affinity for unidentified protein partners.
To explore further the differences in pathophysiology between OPD-spectrum disorders and PVNH, we examined X-inactivation patterns in leukocytes using the androgen receptor (AR) assay25. X inactivation was reported to be random in females with PVNH caused by truncating mutations in FLNA3, indicating that filamin A is dispensable for cell-autonomous survival in leukocytes. In females heterozygous with respect to a mutation associated with OPD-spectrum disorders, by contrast, we found that X inactivation was markedly skewed, especially for mutations associated with more severe phenotypes (Fig. 5). In all (7 of 7) familial cases in which the AR alleles could be unambiguously assigned to the X chromosome harboring the mutated or wild-type allele, inactivation was always biased towards the mutated allele (Fig. 5). These results suggest that leukocytes from individuals with OPD-spectrum disorders that express mutant filamin A are at a selective disadvantage.
Diagnostic categories are presented on the horizontal axis. Controls are individuals who are related to known mutation carriers but who were shown to have two wild-type alleles. Individuals for whom preferential inactivation of the mutated allele could be shown are indicated with triangular symbols. Individuals indicated with an asterisk were previously studied28. The values shown are mean ± s.e.m. calculated for 3–7 independent measurements.
Our finding of clustered in-frame mutations in FLNA associated with OPD1, OPD2, FMD and MNS raises two key questions. What are the mechanisms underlying the genotype–phenotype correlation in the OPD-spectrum disorders? What is the biological basis of these phenotypes, which are entirely distinct from PVNH? We emphasize the following points. First, individuals with mutations in FLNA associated with OPD-spectrum disorders do not have a higher prevalence of seizures or the characteristic phenotypic features of PVNH26,27. Second, 12 of 14 mutations causing PVNH predict protein truncation and/or mRNA instability and are assumed to lead to loss of function3,5,6, whereas 17 of 17 mutations causing OPD-spectrum disorders conserve the reading frame, compatible with specific altered functions. Third, the distinct genotype–phenotype correlations that we observe (Table 1) imply that different mutations confer different altered functions. Finally, the X-inactivation data suggest that although loss-of-function mutations do not alter leukocyte survival, the altered function associated with mutations causing OPD-spectrum disorders is detrimental, presumably by conferring a selective growth disadvantage. Consequently we propose that PVNH and the OPD-spectrum disorders encompass discrete mutational categories with little or no overlap despite being allelic entities. Identification of the roles of filamin A in modulating signaling during organogenesis in multiple tissues is an unexpected and new challenge that requires further investigation.
Methods
Subjects and consent.
Informed consent was obtained from all participating individuals and the study was approved by the East Anglia Multicentre Research Ethics Committee.
Refinement of the disease interval.
We used two new polymorphisms to refine the candidate interval in the family showing linkage to OPD2: a (TAAA)n repeat in intron 9 of RENBP and a single-nucleotide polymorphism (IVS5+22G/A) in IRAK1. PCR primers for both polymorphisms are available on request.
Analysis of FLNA.
We carried out mutation screening by WAVE dHPLC according to the manufacturer's specifications (Transgenomic). We designed amplimers to incorporate all known coding sequences and splice sites (GenBank L44140). Primer sequences, PCR conditions and dHPLC analysis temperatures are available on request. We mixed genomic DNA derived from male subjects with DNA from an unaffected male before doing PCR. We subjected all anomalous traces to automated sequencing on a 3100 capillary sequencer (Applied Biosystems). In cases of apparent de novo mutation, we examined the segregation of eight unlinked microsatellite markers to confirm the correct assignment of paternity and maternity. Mutations not shown to be de novo were shown to be absent in a panel of 100 chromosomes from north European individuals, either by allele-specific oligonucleotide hybridization (see Supplementary Table 1 online) or by restriction enzyme digestion. Additional variants identified that were considered to be non-pathogenic are listed in Supplementary Table 2 and include one (5290G→A, encoding A1764T) previously reported to cause PVNH5.
X-inactivation assay.
We measured skewing of X inactivation using the AR triplet repeat assay25, except that we predigested genomic DNA (1 μg) with RsaI either in the presence (+) or absence (−) of the methylation-sensitive enzyme HpaII (20 IU). We analyzed the products on an ABI 377 sequencer using GeneScan software. Differences in peak areas for the two alleles in the HpaII(+) assay were corrected for differences in amplification efficiency measured in the HpaII(−) assay, and the final results were expressed as a ratio. Forty-two of 59 subjects analyzed were informative in the assay.
URL.
Information on Duchenne and Becker muscular dystrophy is available at http://www.DMD.nl/.
Note: Supplementary information is available on the Nature Genetics website.
Accession codes
References
Stossel, T.P. et al. Filamins as integrators of cell mechanics and signalling. Nat. Rev. Mol. Cell Biol. 2, 138–145 (2001).
van der Flier, A. & Sonnenberg, A. Structural and functional aspects of filamins. Biochim. Biophys. Acta 1538, 99–117 (2001).
Fox, J.W. et al. Mutations in filamin 1 prevent migration of cerebral cortical neurons in human periventricular heterotopia. Neuron 21, 1315–1325 (1998).
Eksioglu, Y.Z. et al. Periventricular heterotopia: an X-linked dominant epilepsy locus causing aberrant cerebral cortical development. Neuron 16, 77–87 (1996).
Sheen, V.L. et al. Mutations in the X-linked filamin 1 gene cause periventricular nodular heterotopia in males as well as in females. Hum. Mol. Genet. 10, 1775–1783 (2001).
Moro, F. et al. Familial periventricular heterotopia. Missense and distal truncating mutations of the FLN1 gene. Neurology 58, 916–921 (2002).
Robertson, S., Gunn, T., Allen, B., Chapman, C. & Becroft, D. Are Melnick–Needles syndrome and otopalatodigital syndrome type II allelic? Observations in a four generation kindred. Am. J. Med. Genet. 71, 341–347 (1997).
Robertson, S.P. et al. Linkage of otopalatodigital syndrome type 2 (OPD2) to distal Xq28: evidence for allelism with OPD1. Am. J. Hum. Genet. 69, 223–227 (2001).
Biancalana, V., Le Marec, B., Odent, S., van den Hurk, J.A. & Hanauer, A. Oto-palato-digital syndrome type1: further evidence for assignment of the locus to Xq28. Hum. Genet. 88, 228–230 (1991).
Kosho, T. et al. Refined mapping of the gene for otopalatodigital syndrome type I. J. Med. Genet. 39, E7 (2002).
Fitch, N., Jequier, S. & Gorlin, R. The otopalatodigital syndrome, proposed type II. Am. J. Med. Genet. 15, 655–664 (1983).
Verloes, A. et al. Fronto-otopalatodigital osteodysplasia: clinical evidence for a single entity encompassing Melnick–Needles syndrome, otopalatodigital syndrome types 1 and 2, and frontometaphyseal dysplasia. Am. J. Med. Genet. 90, 407–422 (2000).
Taybi, H. Generalized skeletal dysplasia with multiple anomalies. Am. J. Roentgen. 88, 450–457 (1962).
Dudding, B.A., Gorlin, R.J. & Langer, L.O. A new symptom complex consisting of deafness, dwarfism, cleft palate, characteristic facies and a generalised bone dysplasia. Am. J. Dis. Child. 113, 214–221 (1967).
Gorlin, R.J. & Cohen, M.M. Frontometaphyseal dysplasia. A new syndrome. Am. J. Dis. Child. 118, 487–494 (1969).
Melnick, J.C. & Needles, C.F. An undiagnosed bone dysplasia. A 2 family study of 4 generations and 3 generations. Am. J. Roentgen. 97, 39–48 (1966).
Donnenfeld, A.E., Conard, K.A., Roberts, N.S., Borns, P.F. & Zackai, E.H. Melnick–Needles syndrome in males: a lethal multiple congenital anomalies syndrome. Am. J. Med. Genet. 27, 159–173 (1987).
Sheen, V.L. et al. Filamin A and filamin B are co-expressed within neurons during periods of neuronal migration and can physically interact. Hum. Mol. Genet. 11, 2845–2854 (2002).
Le Marec, B. et al. Syndrome oto-palato-digital de type I atteignant cinq générations relations avec la forme de type II. Ann. Génét. 31, 155–161 (1988).
Hassoun, H. et al. Characterization of the underlying molecular defect in hereditary spherocytosis associated with spectrin deficiency. Blood 90, 398–406 (1997).
Kaplan, J.M. et al. Mutations in ACTN4, encoding α-actinin-4, cause familial focal segmental glomerulosclerosis. Nat. Genet. 24, 251–256 (2000).
Norwood, F.L., Sutherland-Smith, A.J., Keep, N.H. & Kendrick-Jones, J. The structure of the N-terminal actin-binding domain of human dystrophin and how mutations in this domain may cause Duchenne or Becker muscular dystrophy. Structure 8, 481–491 (2000).
Carugo, K.D., Bañuelos, S. & Saraste, M. Crystal structure of a calponin homology domain. Nat. Struct. Biol. 4, 175–179 (1997).
McCoy, A.J., Fucini, P., Noegel, A.A. & Stewart, M. Structural basis for dimerization of the Dictyostelium gelation factor (ABP120) rod. Nat. Struct. Biol. 6, 836–841 (1999).
Allen, R.C., Zoghbi, H.Y., Moseley, A.B., Rosenblatt, H.M. & Belmont, J.W. Methylation of HpaII and HhaI sites near the polymorphic CAG repeat in the human androgen-receptor gene correlates with X-chromosome inactivation. Am. J. Hum. Genet. 51, 1229–1239 (1992).
André, M., Vigneron, J. & Didier, F. Abnormal facies, cleft palate and generalised dysostosis: a lethal X-linked syndrome. J. Pediatr. 98, 747–752 (1981).
Stratton, R.F. & Bluestone, D.F. Oto-palatal-digital syndrome type II with X-linked cerebellar hypoplasia/hydrocephalus. Am. J. Med. Genet. 41, 169–172 (1991).
Kristiansen, M., Knudsen, G.P., Søyland, A., Westvik, J. & Ørstavik, K.H. Phenotypic variation in Melnick–Needles syndrome is not reflected in X inactivation patterns from blood or buccal smear. Am. J. Med. Genet. 108, 120–127 (2002).
Esnouf, R.M. An extensively modified version of MolScript that includes greatly enhanced coloring capabilities. J. Mol. Graph. Model. 15, 132–134 (1997).
Acknowledgements
We are grateful to the individuals, families and members of the MNS Support Group who participated in this research, A. McCoy for discussions on filamin A structure, M. Cossee and B. Hane for sharing unpublished results and N. Elanko, I. Taylor, S. Butler and K. Clark for technical assistance. This work was supported by a Nuffield Medical Fellowship (S.P.R.) and a Wellcome Trust Senior Research Fellowship in Clinical Science (A.O.M.W.).
Author information
Authors and Affiliations
Consortia
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Robertson, S., Twigg, S., Sutherland-Smith, A. et al. Localized mutations in the gene encoding the cytoskeletal protein filamin A cause diverse malformations in humans. Nat Genet 33, 487–491 (2003). https://doi.org/10.1038/ng1119
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng1119
This article is cited by
-
Prenatal detection and molecular cytogenetic characterization of Xp deletion and Xq duplication: a case report and literature review
BMC Medical Genomics (2024)
-
Clinical genetics evaluation and testing of connective tissue disorders: a cross-sectional study
BMC Medical Genomics (2022)
-
A novel FLNA variant in a fetus with skeletal dysplasia
Human Genome Variation (2022)
-
Bipolar disorder with Melnick–Needles syndrome and periventricular nodular heterotopia: two case reports and a review of the literature
Journal of Medical Case Reports (2021)
-
The variants at FLNA and FLNB contribute to the susceptibility of hypertension and stroke with differentially expressed mRNA
The Pharmacogenomics Journal (2021)