Abstract
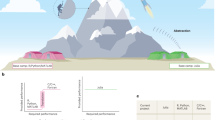
In reviewing the past decade, it is clear that genomics was, and still is, driven by innovative technologies, perhaps more so than any other scientific area in recent memory. From the outset, computing, mathematics and new automated laboratory techniques have been key components in allowing the field to move forward rapidly. We highlight some key innovations that have come together to nurture the explosive growth that makes a new era of genomics a reality. We also document how these new approaches have fueled further innovations and discoveries.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Kuska, B. Beer, Bethesda, and biology: how “genomics” came into being. J. Natl. Cancer Inst. 90, 93 (1998).
Sanger, F., Nicklen, S. & Coulson, A.R. DNA sequencing with chain-terminating inhibitors. Proc. Natl. Acad. Sci. USA 74, 5463–5467 (1977).
Martin-Gallardo, A. et al. Automated DNA sequencing and analysis of 106 kilobases from human chromosome 19q13.3. Nat. Genet. 1, 34–39 (1992).
McCombie, W.R. et al. Expressed genes, Alu repeats and polymorphisms in cosmids sequenced from chromosome 4p16.3. Nat. Genet. 1, 348–353 (1992).
Wada, A. The practicability of and necessity for developing a large-scale DNA-base sequencing system: toward the establishment of international super DNA-sequencing centers. Basic Life Sci. 46, 119–130 (1988).
Wada, A. Fundamental significance of DNA mass-sequencing factory for biological sciences in future. Adv. Biophys. 30, 85–103 (1994).
Understanding Our Genetic Inheritance: The Human Genome Project, The First Five Years, FY 1991–1995. NIH Report 90–1590 (1990).
Goffeau, A. et al. Life with 6000 genes. Science 274, 563–567 (1996).
Blattner, F.R. et al. The complete genome sequence of Escherichia coli K-12. Science 277, 1453–1474 (1997).
Adams, M.D. et al. Complementary DNA sequencing: expressed-sequence tags and human genome project. Science 252, 1651–1656 (1991).
Roberts, L. Gambling on a shortcut to genome sequencing. Science 252, 1618–1619 (1991).
Uberbacher, E.C. & Mural, R.J. Locating protein-coding regions in human DNA sequences by a multiple sensor–neural network approach. Proc. Natl. Acad. Sci. USA 88, 11261–11265 (1991).
Olson, M., Hood, L., Cantor, C. & Botstein, D. A common language for physical mapping of the human genome. Science 245, 1434–1435 (1989).
Putney, S.D., Herlihy, W.C. & Schimmel, P. A new troponin T and cDNA clones for 13 different muscle proteins, found by shotgun sequencing. Nature 302, 718–721 (1983).
Papadopoulos, N. et al. Mutation of a Mutl homolog in hereditary colon cancer. Science 263, 1625–1629 (1994).
Nicolaides, N.C. et al. Mutations of 2 Pms homologs in hereditary nonpolyposis colon cancer. Nature 371, 75–80 (1994).
Somerville, C. & Somerville, S. Plant functional genomics. Science 285, 380–383 (1999).
Venter, J.C. et al. The sequence of the human genome. Science 291, 1304–1351 (2001).
Lander, E.S. et al. Initial sequencing and analysis of the human genome. Nature 409, 860–921 (2001).
Velculescu, V.E., Zhang, L., Vogelstein, B. & Kinzler, K.W. Serial analysis of gene expression. Science 270, 484–487 (1995).
Lockhart, D.J. et al. Expression monitoring by hybridization to high-density oligonucleotide arrays. Nat. Biotechnol. 14, 1675–1680 (1996).
Wodicka, L., Dong, H.L., Mittmann, M., Ho, M.H. & Lockhart, D.J. Genome-wide expression monitoring in Saccharomyces cerevisiae. Nat. Biotechnol. 15, 1359–1367 (1997).
Duggan, D.J., Bittner, M., Chen, Y.D., Meltzer, P. & Trent, J.M. Expression profiling using cDNA microarrays. Nat. Genet. 21, 10–14 (1999).
Waterston, R. et al. A survey of expressed genes in Caenorhabditis elegans. Nat. Genet. 1, 114–123 (1992).
Okubo, K. et al. Large-scale cDNA sequencing for analysis of quantitative and qualitative aspects of gene expression. Nat. Genet. 2, 173–179 (1992).
Adams, M.D., Kerlavage, A.R., Fields, C. & Venter, J.C. 3,400 new expressed-sequence tags identify diversity of transcripts in human brain. Nat. Genet. 4, 256–267 (1993).
Adams, M.D., Soares, M.B., Kerlavage, A.R., Fields, C. & Venter, J.C. Rapid cDNA sequencing (expressed-sequence tags) from a directionally cloned human infant brain cDNA library. Nat. Genet. 4, 373–386 (1993).
Newman, T. et al. Genes galore—a summary of methods for accessing results from large-scale partial sequencing of anonymous Arabidopsis cDNA clones. Plant Physiol. 106, 1241–1255 (1994).
Adams, M.D. et al. Initial assessment of human gene diversity and expression patterns based upon 83 million nucleotides of cDNA sequence. Nature 377, 3–174 (1995).
Hillier, L. et al. Generation and analysis of 280,000 human expressed-sequence tags. Genome Res. 6, 807–828 (1996).
Schuler, G.D. et al. A gene map of the human genome. Science 274, 540–546 (1996).
Altschul, S.F., Boguski, M.S., Gish, W. & Wootton, J.C. Issues in searching molecular sequence databases. Nat. Genet. 6, 119–129 (1994).
Fernandesalnemri, T., Litwack, G. & Alnemri, E.S. Cpp32, a novel human apoptotic protein with homology to Caenorhabditis elegans cell-death protein Ced-3 and mammalian interleukin-1β-converting enzyme. J. Biol. Chem. 269, 30761–30764 (1994).
Simonet, W.S. et al. Osteoprotegerin: a novel secreted protein involved in the regulation of bone density. Cell 89, 309–319 (1997).
Messersmith, E.K. et al. Semaphorin III can function as a selective chemorepellent to pattern sensory projections in the spinal cord. Neuron 14, 949–959 (1995).
Sutton, G.G., White, O., Adams, M.D. & Kerlavage, A.R. TIGR Assembler: a new tool for assembling large shotgun sequencing projects. 1, 9–19 (1995).
Fleischmann, R.D. et al. Whole-genome random sequencing and assembly of Haemophilus influenzae Rd. Science 269, 496–512 (1995).
Himmelreich, R. et al. Complete sequence analysis of the genome of the bacterium Mycoplasma pneumoniae. Nucleic Acids Res. 24, 4420–4449 (1996).
Nelson, K.E. et al. Evidence for lateral gene transfer between Archaea and Bacteria from genome sequence of Thermotoga maritima. Nature 399, 323–329 (1999).
Philipp, W.J. et al. An integrated map of the genome of the tubercle bacillus, Mycobacterium tuberculosis H37Rv, and comparison with Mycobacterium leprae. Proc. Natl. Acad. Sci. USA 93, 3132–3137 (1996).
Brown, J.R. & Doolittle, W.F. Archaea and the prokaryote-to-eukaryote transition. Microbiol. Mol. Biol. Rev. 61, 456–502 (1997).
Rubin, G.M. et al. Comparative genomics of the eukaryotes. Science 287, 2204–2215 (2000).
Mural, R.J. et al. A comparison of whole-genome shotgun-derived mouse chromosome 16 and the human genome. Science 296, 1661–1671 (2002).
Waterston, R.H. et al. Initial sequencing and comparative analysis of the mouse genome. Nature 420, 520–562 (2002).
Tatusov, R.L., Koonin, E.V. & Lipman, D.J. A genomic perspective on protein families. Science 278, 631–637 (1997).
Dujon, B. The yeast genome project: what did we learn? Trends Genet. 12, 263–270 (1996).
Lashkari, D.A. et al. Yeast microarrays for genome-wide parallel genetic and gene-expression analysis. Proc. Natl. Acad. Sci. USA 94, 13057–13062 (1997).
Blackstock, W.P. & Weir, M.P. Proteomics: quantitative and physical mapping of cellular proteins. Trends Biotechnol. 17, 121–127 (1999).
Yates, J.R. Mass spectrometry and the age of the proteome. J. Mass Spectrom. 33, 1–19 (1998).
Govan, J.R.W. & Deretic, V. Microbial pathogenesis in cystic fibrosis: mucoid Pseudomonas aeruginosa and Burkholderia cepacia. Microbiol. Rev. 60, 539–574 (1996).
Freiberg, C. et al. Molecular basis of symbiosis between Rhizobium and legumes. Nature 387, 394–401 (1997).
Zumft, W.G. Cell biology and molecular basis of denitrification. Microbiol. Mol. Biol. Rev. 61, 533–616 (1997).
Merrick, M.J. & Edwards, R.A. Nitrogen control in bacteria. Microbiol. Rev. 59, 604–622 (1995).
Pennisi, E. DNA sequencers' trial by fire. Science 280, 814–817 (1998).
Weber, J.L. & Myers, E.W. Human whole-genome shotgun sequencing. Genome Res. 7, 401–409 (1997).
Green, P. Against a whole-genome shotgun. Genome Res. 7, 410–417 (1997).
Meldrum, D. Automation for genomics, part one: preparation for sequencing. Genome Res. 10, 1081–1092 (2000).
Meldrum, D. Automation for genomics, part two: sequencers, microarrays, and future trends. Genome Res. 10, 1288–1303 (2000).
Stein, L.D. et al. The generic genome browser: a building block for a model organism system database. Genome Res. 12, 1599–1610 (2002).
Doggett, N.A. et al. An integrated physical map of human chromosome 16. Nature 377, 335–365 (1995).
Venter, J.C., Smith, H.O. & Hood, L. A new strategy for genome sequencing. Nature 381, 364–366 (1996).
Ewing, B., Hillier, L., Wendl, M.C. & Green, P. Base-calling of automated sequencer traces using phred. I. Accuracy assessment. Genome Res. 8, 175–185 (1998).
Ewing, B. & Green, P. Base-calling of automated sequencer traces using phred. II. Error probabilities. Genome Res. 8, 186–194 (1998).
Chou, H.H. & Holmes, M.H. DNA sequence quality trimming and vector removal. Bioinformatics 17, 1093–1104 (2001).
Myers, E.W. Toward simplifying and accurately formulating fragment assembly. J. Comput. Biol. 2, 275–290 (1995).
Myers, E.W. et al. A whole-genome assembly of Drosophila. Science 287, 2196–2204 (2000).
Adams, M.D. et al. The genome sequence of Drosophila melanogaster. Science 287, 2185–2195 (2000).
Holt, R.A. et al. The genome sequence of the malaria mosquito Anopheles gambiae. Science 298, 129–149 (2002).
Batzoglou, S. et al. ARACHNE: a whole-genome shotgun assembler. Genome Res. 12, 177–189 (2002).
Aparicio, S. et al. Whole-genome shotgun assembly and analysis of the genome of Fugu rubripes. Science 297, 1301–1310 (2002).
Mullikin, J.C. & Ning, Z. The phusion assembler. Genome Res. 13, 81–90 (2003).
Kent, W.J. & Haussler, D. Assembly of the working draft of the human genome with GigAssembler. Genome Res. 11, 1541–1548 (2001).
Huang, X. & Madan, A. CAP3: a DNA sequence assembly program. Genome Res. 9, 868–877 (1999).
Pevzner, P.A., Tang, H. & Waterman, M.S. An Eulerian path approach to DNA fragment assembly. Proc. Natl. Acad. Sci. USA 98, 9748–9753 (2001).
The C. elegans Sequencing Consortium. Genome sequence of the nematode C. elegans: a platform for investigating biology. Science 282, 2012–2018 (1998).
The Arabidopsis Genome Initiative. Analysis of the genome sequence of the flowering plant Arabidopsis thaliana. Nature 408, 796–815 (2000).
Bankier, A.T. et al. The DNA sequence of the human cytomegalovirus genome. DNA Seq. 2, 1–12 (1991).
Baer, R. et al. DNA sequence and expression of the B95-8 Epstein–Barr virus genome. Nature 310, 207–211 (1984).
Seigneur, M., Bidnenko, V., Ehrlich, S.D. & Michel, B. RuvAB acts at arrested replication forks. Cell 95, 419–430 (1998).
Takahashi, S., Kuzuyama, T., Watanabe, H. & Seto, H. A 1-deoxy-d-xylulose 5-phosphate reductoisomerase catalyzing the formation of 2-c-methyl-d-elythritol 4-phosphate in an alternative non-mevalonate pathway for terpenoid biosynthesis. Proc. Natl. Acad. Sci. USA 95, 9879–9884 (1998).
Razin, S., Yogev, D. & Naot, Y. Molecular biology and pathogenicity of mycoplasmas. Microbiol. Mol. Biol. Rev. 62, 1094–1156 (1998).
Perna, N.T. et al. Molecular evolution of a pathogenicity island from enterohemorrhagic Escherichia coli O157:H7. Infect. Immun. 66, 3810–3817 (1998).
Martin, W. & Muller, M. The hydrogen hypothesis for the first eukaryote. Nature 392, 37–41 (1998).
Hutchison, C.A. et al. Global transposon mutagenesis and a minimal mycoplasma genome. Science 286, 2165–2169 (1999).
Fraser, C.M. et al. The minimal gene complement of Mycoplasma genitalium. Science 270, 397–403 (1995).
Mushegian, A.R. & Koonin, E.V. A minimal gene set for cellular life derived by comparison of complete bacterial genomes. Proc. Natl. Acad. Sci. USA 93, 10268–10273 (1996).
Drake, J.W., Charlesworth, B., Charlesworth, D. & Crow, J.F. Rates of spontaneous mutation. Genetics 148, 1667–1686 (1998).
Lawrence, J.G. & Ochman, H. Molecular archaeology of the Escherichia coli genome. Proc. Natl. Acad. Sci. USA 95, 9413–9417 (1998).
Collins, F.S. Shattuck lecture—medical and societal consequences of the human genome project. N. Engl. J. Med. 341, 28–37 (1999).
Shizuya, H. et al. Cloning and stable maintenance of 300-kilobase-pair fragments of human DNA in Escherichia coli using an F-factor-based vector. Proc. Natl. Acad. Sci. USA 89, 8794–8797 (1992).
Burke, D.T., Carle, G.F. & Olson, M.V. Cloning of large segments of exogenous DNA into yeast by means of artificial chromosome vectors. Science 236, 806–812 (1987).
Boguski, M.S. & Schuler, G.D. Establishing a human transcript map. Nat. Genet. 10, 369–371 (1995).
Schena, M. et al. Parallel human genome analysis: microarray-based expression monitoring of 1,000 genes. Proc. Natl. Acad. Sci. USA 93, 10614–10619 (1996).
Bult, C.J. et al. Complete genome sequence of the methanogenic archaeon, Methanococcus jannaschii. Science 273, 1058–1073 (1996).
Redenbach, M. et al. A set of ordered cosmids and a detailed genetic and physical map for the 8 Mb Streptomyces coelicolor A3(2) chromosome. Mol. Microbiol. 21, 77–96 (1996).
Tomb, J.F. et al. The complete genome sequence of the gastric pathogen Helicobacter pylori. Nature 388, 539–547 (1997).
Kunst, F. et al. The complete genome sequence of the Gram-positive bacterium Bacillus subtilis. Nature 390, 249–256 (1997).
Klenk, H.P. et al. The complete genome sequence of the hyperthermophilic, sulphate-reducing archaeon Archaeoglobus fulgidus. Nature 390, 364–370 (1997).
Fraser, C.M. et al. Genomic sequence of a Lyme disease spirochaete, Borrelia burgdorferi. Nature 390, 580–586 (1997).
Andersson, S.G.E. et al. The genome sequence of Rickettsia prowazekii and the origin of mitochondria. Nature 396, 133–140 (1998).
Deckert, G. et al. The complete genome of the hyperthermophilic bacterium Aquifex aeolicus. Nature 392, 353–358 (1998).
Fraser, C.M. et al. Complete genome sequence of Treponema pallidum, the syphilis spirochete. Science 281, 375–388 (1998).
Gardner, M.J. et al. Chromosome 2 sequence of the human malaria parasite Plasmodium falciparum. Science 282, 1126–1132 (1998).
Tettelin, H. et al. Complete genome sequence of Neisseria meningitidis serogroup B strain MC58. Science 287, 1809–1815 (2000).
Coux, O., Tanaka, K. & Goldberg, A.L. Structure and functions of the 20S and 26S proteasomes. Annu. Rev. Biochem. 65, 801–847 (1996).
Wang, J.C. DNA topoisomerases. Annu. Rev. Biochem. 65, 635–692 (1996).
Paulsen, I.T., Brown, M.H. & Skurray, R.A. Proton-dependent multidrug efflux systems. Microbiol. Rev. 60, 575–608 (1996).
Nikaido, H. Multidrug efflux pumps of gram-negative bacteria. J. Bact. 178, 5853–5859 (1996).
Krokan, H.E., Standal, R. & Slupphaug, G. DNA glycosylases in the base excision repair of DNA. Biochem. J. 325, 1–16 (1997).
Miklos, G.L.G. & Rubin, G.M. The role of the genome project in determining gene function: insights from model organisms. Cell 86, 521–529 (1996).
Silver, S. & Phung, L.T. Bacterial heavy metal resistance: new surprises. Annu. Rev. Microbiol. 50, 753–789 (1996).
Lowe, T.M. & Eddy, S.R. tRNAscan-SE: a program for improved detection of transfer RNA genes in genomic sequence. Nucleic Acids Res. 25, 955–964 (1997).
Gurtler, V. & Stanisich, V.A. New approaches to typing and identification of bacteria using the 16S-23S rDNA spacer region. Microbiology 142, 3–16 (1996).
Cole, S.T. et al. Deciphering the biology of Mycobacterium tuberculosis from the complete genome sequence. Nature 393, 537–544 (1998).
Dunham, I. et al. The DNA sequence of human chromosome 22. Nature 402, 489–495 (1999).
Stover, C.K. et al. Complete genome sequence of Pseudomonas aeruginosa PAO1, an opportunistic pathogen. Nature 406, 959–964 (2000).
Perna, N.T. et al. Genome sequence of enterohaemorrhagic Escherichia coli O157: H7. Nature 409, 529–533 (2001).
Kuroda, M. et al. Whole-genome sequencing of meticillin-resistant Staphylococcus aureus. Lancet 357, 1225–1240 (2001).
Simpson, A.J.G. et al. The genome sequence of the plant pathogen Xylella fastidiosa. Nature 406, 151–157 (2000).
Elliott, S.J. et al. The complete sequence of the locus of enterocyte effacement (LEE) from enteropathogenic Escherichia coli E2348/69. Mol. Microbiol. 28, 1–4 (1998).
Jain, R., Rivera, M.C. & Lake, J.A. Horizontal gene transfer among genomes: the complexity hypothesis. Proc. Natl. Acad. Sci. USA 96, 3801–3806 (1999).
Wallin, E. & von Heijne, G. Genome-wide analysis of integral membrane proteins from eubacterial, archaean, and eukaryotic organisms. Protein Sci. 7, 1029–1038 (1998).
Dandekar, T., Snel, B., Huynen, M. & Bork, P. Conservation of gene order: a fingerprint of proteins that physically interact. Trends Biochem. Sci. 23, 324–328 (1998).
Nunes-Duby, S.E., Kwon, H.J., Tirumalai, R.S., Ellenberger, T. & Landy, A. Similarities and differences among 105 members of the Int family of site-specific recombinases. Nucleic Acids Res. 26, 391–406 (1998).
Datsenko, K.A. & Wanner, B.L. One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc. Natl. Acad. Sci. USA 97, 6640–6645 (2000).
Surette, M.G., Miller, M.B. & Bassler, B.L. Quorum sensing in Escherichia coli, Salmonella typhimurium, and Vibrio harveyi: a new family of genes responsible for autoinducer production. Proc. Natl. Acad. Sci. USA 96, 1639–1644 (1999).
Bochtler, M., Ditzel, L., Groll, M., Hartmann, C. & Huber, R. The proteasome. Annu. Rev. Biophys. Biomol. Struct. 28, 295–317 (1999).
Sargent, F. et al. Overlapping functions of components of a bacterial Sec- independent protein export pathway. EMBO J. 17, 3640–3650 (1998).
Weiner, J.H. et al. A novel and ubiquitous system for membrane targeting and secretion of cofactor-containing proteins. Cell 93, 93–101 (1998).
Hung, L.W. et al. Crystal structure of the ATP-binding subunit of an ABC transporter. Nature 396, 703–707 (1998).
Locher, K.P. et al. Transmembrane signaling across the ligand-gated FhuA receptor: crystal structures of free and ferrichrome-bound states reveal allosteric changes. Cell 95, 771–778 (1998).
Walhout, A.J.M. et al. Protein interaction mapping in C. elegans using proteins involved in vulval developement. Science 287, 116–122 (2000).
Ito, T. et al. A comprehensive two-hybrid analysis to explore the yeast protein interactome. Proc. Natl. Acad. Sci. USA 98, 4569–4574 (2001).
Marcotte, E.M. et al. Detecting protein function and protein–protein interactions from genome sequences. Science 285, 751–753 (1999).
Schwartz, S. et al. PipMaker—a Web server for aligning two genomic DNA sequences. Genome Res. 10, 577–586 (2000).
Lukashin, A.V. & Borodovsky, M. GeneMark.hmm: new solutions for gene finding. Nucleic Acids Res. 26, 1107–1115 (1998).
Kozak, M. Initiation of translation in prokaryotes and eukaryotes. Gene 234, 187–208 (1999).
Richmond, C.S., Glasner, J.D., Mau, R., Jin, H.F. & Blattner, F.R. Genome-wide expression profiling in Escherichia coli K-12. Nucleic Acids Res. 27, 3821–3835 (1999).
Tao, H., Bausch, C., Richmond, C., Blattner, F.R. & Conway, T. Functional genomics: expression analysis of Escherichia coli growing on minimal and rich media. J. Bact. 181, 6425–6440 (1999).
Acknowledgements
The authors thank D. Borton, H. Kowalski and M. Peterson for their help with this manuscript.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Venter, J., Levy, S., Stockwell, T. et al. Massive parallelism, randomness and genomic advances. Nat Genet 33 (Suppl 3), 219–227 (2003). https://doi.org/10.1038/ng1114
Issue Date:
DOI: https://doi.org/10.1038/ng1114
This article is cited by
-
Comparative 16S rDNA metagenomics study of two samples of cassava peel heap from Nigeria and India
3 Biotech (2019)
-
A comparison of sequencing platforms and bioinformatics pipelines for compositional analysis of the gut microbiome
BMC Microbiology (2017)
-
Impact of germline and somatic missense variations on drug binding sites
The Pharmacogenomics Journal (2017)
-
Suppression Subtractive Hybridization Versus Next-Generation Sequencing in Plant Genetic Engineering: Challenges and Perspectives
Molecular Biotechnology (2015)
-
Comparing whole genomes using DNA microarrays
Nature Reviews Genetics (2008)