Abstract
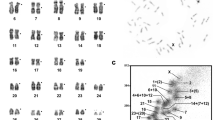
Expressed-sequence tag (EST) maps are an adjunct to sequence-based analytical methods of gene detection and localization for those species for which such data are available, and provide anchors for high-density homology and orthology mapping in species for which large-scale sequencing has yet to be done1. Species for which radiation hybrid–based transcript maps have been established include human2, rat3,4,5, mouse6, dog7, cat8 and zebrafish9,10. We have established a comprehensive first-generation–placement radiation hybrid map of the mouse consisting of 5,904 mapped markers (3,993 ESTs and 1,911 sequence-tagged sites (STSs)). The mapped ESTs, which often originate from small-EST clusters, are enriched for genes expressed during early mouse embryogenesis and are probably different from those localized in humans. We have confirmed by in situ hybridization that even singleton ESTs, which are usually not retained for mapping studies, may represent bona fide transcribed sequences. Our studies on mouse chromosomes 12 and 14 orthologous to human chromosome 14 show the power of our radiation hybrid map as a predictive tool for orthology mapping in humans.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Yang, Y.P. & Womack, J.E. Parallel radiation hybrid mapping: a powerful tool for high-resolution genomic comparison. Genome Res. 8, 731–736 (1998).
Deloukas, P. et al. A physical map of 30,000 human genes. Science 282, 744–746 (1998).
Watanabe, T.K. et al. A radiation hybrid map of the rat genome containing 5,255 markers. Nature Genet. 22, 27–36 (1999).
Steen, R.G. et al. A high-density integrated genetic linkage and radiation hybrid map of the laboratory rat. Genome Res. 9, AP1–8 (1999).
Scheetz T.E. et al. Generation of a high-density rat EST map. Genome Res. 11, 497–502 (2001).
Van Etten, W.J. et al. Radiation hybrid map of the mouse genome. Nature Genet. 22, 384–387 (1999).
Mellersh, C.S. et al. An integrated linkage–radiation hybrid map of the canine genome. Mamm. Genome 11, 120–130 (2000).
Murphy, W.J. et al. A radiation hybrid map of the cat genome: implications for comparative mapping. Genome Res. 10, 691–702 (2000).
Geisler, R. et al. A radiation hybrid map of the zebrafish genome. Nature Genet. 23, 86–89 (1999).
Hukriede, N.A. et al. Radiation hybrid mapping of the zebrafish genome. Proc. Natl Acad. Sci. USA 96, 9745–9750 (1999).
Dunwoodie, S.L., Henrique, D., Harrison, S.M. & Beddington, R.S.P. Mouse Dll3: a novel divergent Delta gene which may complement the function of other Delta homologues during early pattern formation in the mouse embryo. Development 124, 3065–3076 (1997).
Harrison, S.M., Dunwoodie, S.L., Arkell, R.M., Lehrach, H. & Beddington, R.S.P. Isolation of novel tissue-specific genes from cDNA libraries representing the individual tissue constituents of the gastrulating mouse embryo. Development 121, 2479–2489 (1995).
McCarthy, L.C. et al. A first generation whole genome-radiation hybrid map spanning the mouse genome. Genome Res. 7, 1153–1161 (1997).
Breen, M. et al. Towards high resolution maps of the mouse and human genomes—a facility for ordering markers to 0.1 cM resolution. Hum. Mol. Genet. 3, 621–627 (1994).
Dietrich, W. F. et al. A comprehensive genetic map of the mouse genome. Nature 380, 149–152 (1996).
Elliott, R.W., Manly, K.F. & Hohman, C. A radiation hybrid map of mouse chromosome 13. Genomics 57, 365–370 (1999).
Sabile, A., Poras, I., Cherif, D., Goodfellow, P. & Avner, P. Isolation of monochromosomal hybrids for mouse chromosomes 3, 6, 10, 12, 14 and 18. Mamm. Genome 8, 81–85 (1997).
Gyapay, G. et al. A radiation hybrid map of the human genome. Hum. Mol. Genet. 5, 339–46 (1996).
Hudson, T.J. et al. A radiation hybrid map of mouse genes. Nature Genet. 27, 201–205.
DeBry, R.W. & Seldin, M.F. Human/mouse homology relationships. Genomics 33, 337–351 (1996).
Nolan, P.M. et al. A systematic, genome-wide, phenotype-driven mutagenesis programme for gene function studies in the mouse. Nature Genet. 25, 440–443 (2000).
Hrabe de Angelis, M.H. et al. Genome-wide, large-scale production of mutant mice by ENU mutagenesis. Nature Genet. 25, 444–447 (2000).
Kusumi, K. et al. The mouse pudgy mutation disrupts Delta homologue Dll3 and initiation of early somite boundaries. Nature Genet. 19, 274–278 (1998).
Parsons, J.D. & Rodriguez-Tome, P. JESAM: CORBA software components to create and publish EST alignments and clusters. Bioinformatics 16, 313–325 (2000).
Rodriguez-Tome, P. & Lijnzaad, P. RHdb: the Radiation Hybrid Database. Nucleic Acids Res. 28, 146–147 (2000).
Agarwala, R., Applegate, D.L., Maglott, D., Schuler, G.D. & Schaffer, A.A. A fast and scalable radiation hybrid map construction and integration strategy. Genome Res. 10, 350–364 (2000).
Applegate, D., Bixby, R., Chvatal, V. & Cook, W. On the solution of traveling salesman problems. Documenta Mathematica Journal der Deutschen Mathematiker-Vereinigung International Congress of Mathematics III, 645–656 (1998).
Gusfield, D. Algorithms on strings, trees, and sequences: computer science & computational biology (Cambridge University Press, Cambridge, UK, 1997).
Boehnke, M., Lange, K. & Cox, D.R. Statistical methods for multipoint radiation hybrid mapping. Am. J. Hum. Genet. 49, 1174–1188 (1991).
Wilkinson, D.G. Whole-mount in situ hybridisation of vertebrate embryos. in In Situ Hybridisation (ed. Wilkinson, D.G.) 75–83 (Oxford University Press, Oxford, UK, 1992).
Bruls, T. et al. A physical map of human chromosome 14. Nature 409, 947–948 (2001).
Acknowledgements
We dedicate this article to our friend and colleague Rosa Beddington (March 23, 1956–May 18, 2001), a scientist of great biological insight. This work was supported by EEC Contract PL 962414. We thank B. Gorick and the Human Genome Mapping Project at Hinxton, UK, for help with the replication of the 7.5-dpc mouse endoderm library; A.M. Mallon and S. Greenaway of the informatics group at the Medical Research Council, Harwell; and V. Taghavi and E. Sartory for technical assistance in marker typing at MRC Harwell.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Avner, P., Bruls, T., Poras, I. et al. A radiation hybrid transcript map of the mouse genome. Nat Genet 29, 194–200 (2001). https://doi.org/10.1038/ng1001-194
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/ng1001-194
This article is cited by
-
Fine mapping of regulatory loci for mammalian gene expression using radiation hybrids
Nature Genetics (2008)