Abstract
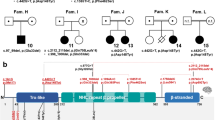
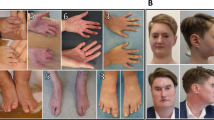
Alagille syndrome is an autosomal dominant disorder characterized by abnormal development of liver, heart, skeleton, eye, face and, less frequently, kidney. Analyses of many patients with cytogenetic deletions or rearrangements have mapped the gene to chromosome 20p12, although deletions are found in a relatively small proportion of patients (< 7%). We have mapped the human Jagged1 gene (JAG1), encoding a ligand for the developmentally important Notch transmembrane receptor, to the Alagille syndrome critical region within 20p12. The Notch intercellular signalling pathway has been shown to mediate cell fate decisions during development in invertebrates and vertebrates. We demonstrate four distinct coding mutations in JAG1 from four Alagille syndrome families, providing evidence that it is the causal gene for Alagille syndrome. All four mutations lie within conserved regions of the gene and cause translational f rameshifts, resulting in gross alterations of the protein product. Patients with cytogenetically detectable deletions including JAG1 have Alagille syndrome, supporting the hypothesis that haploinsufficiency for this gene is one of the mechanisms causing the Alagille syndrome phenotype.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Alagille, D. et al. Syndromic paucity of interlobular bile ducts. J. Pediatr. 110, 195–200 (1987).
Krantz, I.D., Piccoli, D.A. & Spinner, N.B. Alagille syndrome. J. Med. Genet. 34, 152–157(1997).
Danks, D.M., Campbell, P.E., Jack, I., Rogers, J. & Smith, A.L. Studies of the aetiology of neonatal hepatitis and biliary atresia. Arch. Dis. Child. 52, 360–367 (1977).
Krantz, I.D. et al. Deletions of 20p12 in Alagille syndrome: frequency and molecular characterization. Am. J. Med. Genet. 70, 80–86 (1997).
Piccoli, D.A. & Witzleben, C.L. Disorders of the intrahepatic bile ducts.. in Gastrointestinal disease: pathophysiology, Diagnosis, Management, 3rd ed (eds Walker, D . A. et al.) 1124–1140 (B.C. Decker, Philadelphia, 1991).
Watson, G.H. & Miller, V. Arteriohepatic dysplasia: familial pulmonary arterial stenosis with neonatal liver disease. Arch. Dis. Child. 48, 459–466 (1973).
Dhorne-Pollet, S., Deleuze, J.-F., Hadchouel, M. & Bonaiti-Pellie, C. Segregation analysis of Alagille syndrome. J. Med. Genet. 31, 453–457 (1994).
Schnittger, S., Hofers, C., Heidemann, P., Beermann, F. & Hansmann, I. Molecular and cytogenetic analysis of an interstitial 20p deletion associated with syndromic intrahepatic ductular hypoplasia (Alagille syndrome). Hum. Genet. 83, 239–244 (1989).
Spinner, N.B. et al. Cologically balanced t(2;20) in a two generation family with Alagille syndrome: cytogenetic and molecular studies. Am. J. Hum. Genet. 55, 238–243 (1994).
Rand, E.B., Spinner, N.B., Piccoli, D.A., Whitington, P.F. & Taub, R. Molecular analysis of 24 Alagille syndrome families identifies a single submicroscopic deletion and further localizes the Alagille region within 20p12. Am. J. Hum. Genet. 57, 1068–1073 (1995).
Deleuze, J.-F., Hazan, J., Dhorne, S., Weissenbach, J. & Hadchouel, M. Mapping of microsatellite markers in the Alagille region and screening of microdeletions by genotyping 23 patients. Eur. J. Hum. Genet. 2, 185–190 (1994).
Li, L. et al. Human homolog of rat Jagged, JAG 1, inhibits granulocytic differentiation of 32D myeloid progenitors through interaction with Notch 1. Immunity (in press).
Greenwald, I. & Rubin, G.M. Making a difference: the role of cell-cell interactions in establishing separate identities for equivalent cells. Cell. 68, 271–281 (1992).
Fortini, M.E., Rebay, I., Caron, L.A. & Artavan-Tsakonas, S An activated Notch receptor blocks cell-fate commitment in the developing Drosophila eye . Nature 365, 555–557 (1993).
Artavanis-Tsakonas, S., Matsuno, K. & Fortini, M.E. Notch signaling. Science 268, 225–232 (1995).
Franco del Amo, F. et al. Expression pattern of Motch, a mouse homolog of Drosophila Notch, suggests an important role in early postimplantation mouse development. Development 115, 737–744 (1992).
Reaume, A.G., Conlon, R.A., Zirngibl, R., Yamaguchi, T. & Rossant, J. Expression analysis of a Notch homologue in the mouse embryo. Dev. Biol. 154, 377–387 (1992).
Kopan, R. & Weintraub, H., Notch: expression in hair follicles correlates with cell fate determination. Cell Biol. 121, 631–641 (1993).
Lardeli, M., Dahlstrand, J. & Lendahl, U. The novel Notch homolog mouse Notch3 lacks specific epidermal growth factor-repeats and is expressed in proliferating neuroepithelium. Mech. Dev. 46, 123–136 (1994).
Weinmaster, G., Roberts, V.J. & Lemke, G. A homolog of Drosophila Notch expressed during mammalian development. Development 113, 199–205 (1991).
Weinmaster, G., Roberts, V.J. & Lemke, G. Notch2: a second mammalian Notch gene. Development 116, 931–941 (1992).
Uyttendaele, H. et al. Notch4/int-3, a mammary proto-oncogene, is an endothelial cell-specific mammalian Notch gene. Deve/opment 122, 2251–2259 (1996).
Robey, E. et al. An activated form of Notch influences the choice between CD4 and CDS T cell lineages. Cell 87, 483–492 (1996).
Washburn, T. et al. Notch activity influences the αβ versus γδ T cell lineage decision. Cell 88, 833–643 (1997).
Swiatek, P.J., Lindsell, C.E., del Amo, F., Weinmaster, G. amp; Gridley, T. Notch1 is essential for postimplantation development in mice. Genes Dev. 8, 707–719 (1994).
Bao, Z.Z., Cepko, C.L. The expression and function of Notch pathway genes in the developing rat eye. J. Neurosci. 17, 1425–1434.
Ellisen, L.W. et al. TAN-1, the human homolog of the Drosophila Notch gene, is broken by chromosomal translocations in T lymphoblastic neoplasms. Cell. 66, 649–661 (1991).
Sugaya, K. et al. Three genes in the human MHC class III region near the junction with the class II: gene for receptor of advanced glycosylation end products, PBX2 homeobox gene and a Notch homolog, human counterpart of mouse mammary tumor gene int-3 . Genomics. 23, 408–119 (1994).
Muskavitch, M.A. & Hoffman, P.M. Homologs of vertebrate growth factors in Drosophila melanogaster and other vertebrates. Curr. Topics Dev. Biol. 24, 289–328 (1990).
Lindsell, C.E., Shawber, C.J., Boulter, J. & Weinmaster, G. Jagged: a mammalian ligand that activates Notch 1. cell 80, 909–917 (1995).
Henderson, S.T., Gao, D., Lambie, E.J. & Kimble, J. Lag-2 may encode a signaling ligand for the GLP-1 and LIN-12 receptors of C elegans. Development 120, 2913–2924 (1994).
Lieber, T. et al. Single amino acid substitutions in EGF-like elements of Notch and Delta modify Drosophila development and affect cell adhesion in vitro . Neuron. 9, 847–859 (1992).
Shawber, C.J., Boulter, J., Lindsell, C.E. & Weinmaster, G. Jagged2: A Serrate-like gene expressed during rat embryogenesis. Dev. Biol. 180, 370–376 (1996).
Zimrin, A.B. et al. An antisense oligonucleotide to the Notch ligand Jagged enhances fibroblast growth factor-induced angiogenesis in vitro. J. Biol. Chem. 51, 32499–32502(1996).
Spinner, N.B. et al. Mapping the Alagille syndrome critical region within 20p12. Proc. Single Chromosome 20 Workshop, February 1997, Hinxton, Cambridgeshire, UK Cytogenetic. Cell Genet. (in press).
Oda, T. et al. Mutations in the human Jagged! Gene (JAGL1) are responsible for the Alagille syndrome. Nat. Genet. 16, 235–242 (1997).
Delwart, E.L. et al. Genetic relationships determined by a DNA heteroduplex mobility assay: analysis of HIV-1 env genes. Science 262, 1257–1261 (1993).
Kahn, E. et al. Nonsyndromic paucity of interlobular bile ducts: light and electron microscopic evaluation of sequential liver biopsies in early childhood. Hepatology 6, 890–901 (1986).
Novotny, N.M., Zetterman, R.K., Antonson, D.L. & Vanderhoof, J.A. Variation in liver histology in Alagille's syndrome. Am. J. Gastroenterol. 75, 449–506 (1981).
Lindsell, C.E., Boulter, J. diSibio, G., Gossler, A. & Weinmaster, G. Expression patterns of Jagged, Deltal, NotcM, Notch2, and NotchS genes identify ligand-receptor pairs that may function in neural development. Mol. Cell. Neurosd. 8, 14–27 (1996).
Conlon, R.A., Reaume, A.G. & Rossant, J. Notch1 is required for the coordinate segmentation of somites. Development 121, 1533–1545 (1995).
de Angeles, M.H., Mclntyre, J., Gossler, A. Maintenance of somite borders in mice requires the Delta homologue DII1. Nature. 386, 717–721 (1997).
Joutel, A. et al.Notch3 mutations in CADASIL, a hereditary adult-onset condition causing stroke and dementia. Nature 383, 707–710 (1996).
Wilkie, A.O.M. The molecular basis of genetic dominance. Med. Genet. 31, 89–98
Nickerson, E., Greenberg, F., Keating, M.T., McCaskill, C. & Shaffer, L.G. Deletions of the elastin gene at 7q11.23 occur in ∼90% of patients with Williams syndrome. Am. J. Hum. Genet. 56, 1156–1161 (1995).
Petrij, F. et al. Rubinstein-Taybi syndrome caused by mutations in the transcriptional co-activator CBP. Nature 376, 348–351 (1995).
Lindsley, D.L. & Zimm, G.G. The genome of Drosophila melanogaster (Academic Press, New York, 1992).
Heitzler, P. & Simpson, P. The choice of cell fate in the epidermis of Drosophila . Cell 64, 1083–1092(1991).
Pollet, N et al .Construction of a 3.7 Mb physical map within human chromosome 20p12 ordering 18 markers in the Alagille syndrome locus. Genomics 27, 467–474 (1995).
Shepherd, N.S. et al. Preparation and screening of an arrayed human genomic library generated with the P1 cloning system. Proc. Natl. Acad. Sci USA 91, 2629–2633 (1994).
Stokke, T. et al. A physical map of chromosome 20 established using fluorescence in situ hybridization and digital image analysis. Genomics 26, 134–137 (1995).
Shizuya, H. et al. Cloning and stable maintenance of 300-kilobase-pair fragments of human DNA in Escherichia coli using an F-factor based vector. Proc Natl. Acad. Sci USA. 89, 8794–8797 (1992).
Kim, U.-J. et al. Construction and characterization of a human bacterial artificial chromosome library. Genomics 34, 213–218 (1996).
Smith, T.M. et al. complete genomic sequence and analysis of 117 kb of human DNA containing the gene BRCA1. Genome Res. 6, 1029–1049(1996).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Li, L., Krantz, I., Deng, Y. et al. Alagille syndrome is caused by mutations in human Jagged1, which encodes a ligand for Notch1. Nat Genet 16, 243–251 (1997). https://doi.org/10.1038/ng0797-243
Issue Date:
DOI: https://doi.org/10.1038/ng0797-243
This article is cited by
-
Novel JAG1 variants leading to Alagille syndrome in two Chinese cases
Scientific Reports (2024)
-
Fetal liver development and implications for liver disease pathogenesis
Nature Reviews Gastroenterology & Hepatology (2023)
-
A child with chronic kidney disease and hepatic dysfunction: Answers
Pediatric Nephrology (2023)
-
The Notch-mediated circuitry in the evolution and generation of new cell lineages: the tooth model
Cellular and Molecular Life Sciences (2023)
-
Management of adults with Alagille syndrome
Hepatology International (2023)