Abstract
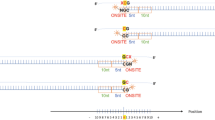
Narcolepsy is a rare sleep disorder with the strongest human leukocyte antigen (HLA) association ever reported. Since the associated HLA-DRB1*1501-DQB1*0602 haplotype is common in the general population (15–25%), it has been suggested that it is almost necessary but not sufficient for developing narcolepsy. To further define the genetic basis of narcolepsy risk, we performed a genome-wide association study (GWAS) in 562 European individuals with narcolepsy (cases) and 702 ethnically matched controls, with independent replication in 370 cases and 495 controls, all heterozygous for DRB1*1501-DQB1*0602. We found association with a protective variant near HLA-DQA2 (rs2858884; P < 3 × 10−8). Further analysis revealed that rs2858884 is strongly linked to DRB1*03-DQB1*02 (P < 4 × 10−43) and DRB1*1301-DQB1*0603 (P < 3 × 10−7). Cases almost never carried a trans DRB1*1301-DQB1*0603 haplotype (odds ratio = 0.02; P < 6 × 10−14). This unexpected protective HLA haplotype suggests a virtually causal involvement of the HLA region in narcolepsy susceptibility.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

Similar content being viewed by others
Change history
27 October 2010
In the version of this article initially published, the name of author Peter Vollenweider was incorrectly written as Peter Vollenwider. Also, Claudio Bassetti’s affiliation was incorrectly listed as Neurocentro (Ente ospedaliero cantonale) della Svizzera Italiana, Ospedale Civico, Lugano, Switzerland. His correct affiliation is Department of Neurology, University Hospital Zurich, Zurich, Switzerland. These errors have been corrected in the HTML and PDF versions of the article.
References
Tafti, M., Dauvilliers, Y. & Overeem, S. Narcolepsy and familial advanced sleep-phase syndrome: molecular genetics of sleep disorders. Curr. Opin. Genet. Dev. 17, 222–227 (2007).
Mignot, E. et al. Complex HLA-DR and -DQ interactions confer risk of narcolepsy-cataplexy in three ethnic groups. Am. J. Hum. Genet. 68, 686–699 (2001).
Nishino, S., Ripley, B., Overeem, S., Lammers, G.J. & Mignot, E. Hypocretin (orexin) deficiency in human narcolepsy. Lancet 355, 39–40 (2000).
Peyron, C. et al. A mutation in a case of early onset narcolepsy and a generalized absence of hypocretin peptides in human narcoleptic brains. Nat. Med. 6, 991–997 (2000).
Thannickal, T.C. et al. Reduced number of hypocretin neurons in human narcolepsy. Neuron 27, 469–474 (2000).
Cvetkovic-Lopes, V. et al. Elevated Tribbles homolog 2 specific antibody levels in narcolepsy patients. J. Clin. Invest. 120, 713–719 (2010).
Miyagawa, T. et al. Variant between CPT1B and CHKB associated with susceptibility to narcolepsy. Nat. Genet. 40, 1324–1328 (2008).
Hallmayer, J. et al. Narcolepsy is strongly associated with the T-cell receptor alpha locus. Nat. Genet. 41, 708–711 (2009).
Lettre, G. & Rioux, J.D. Autoimmune diseases: insights from genome-wide association studies. Hum. Mol. Genet. 17, R116–R121 (2008).
Klein, J. & Sato, A. The HLA system. Second of two parts. N. Engl. J. Med. 343, 782–786 (2000).
Korbel, J.O. et al. Systematic prediction and validation of breakpoints associated with copy-number variants in the human genome. Proc. Natl. Acad. Sci. USA 104, 10110–10115 (2007).
McCarroll, S.A. et al. Integrated detection and population-genetic analysis of SNPs and copy number variation. Nat. Genet. 40, 1166–1174 (2008).
Perry, G.H. et al. Copy number variation and evolution in humans and chimpanzees. Genome Res. 18, 1698–1710 (2008).
Redon, R. et al. Global variation in copy number in the human genome. Nature 444, 444–454 (2006).
Wheeler, D.A. et al. The complete genome of an individual by massively parallel DNA sequencing. Nature 452, 872–876 (2008).
Wong, K.K. et al. A comprehensive analysis of common copy-number variations in the human genome. Am. J. Hum. Genet. 80, 91–104 (2007).
Wellcome Trust Case Control Consortium. Genome-wide association study of CNVs in 16,000 cases of eight common diseases and 3,000 shared controls. Nature 464, 713–720 (2010).
Mignot, E. et al. DQB1*0602 and DQA1*0102 (DQ1) are better markers than DR2 for narcolepsy in Caucasian and black Americans. Sleep 17, S60–S67 (1994).
Mignot, E. et al. Extensive HLA class II studies in 58 non-DRB1*15 (DR2) narcoleptic patients with cataplexy. Tissue Antigens 49, 329–341 (1997).
Firmann, M. et al. The CoLaus study: a population-based study to investigate the epidemiology and genetic determinants of cardiovascular risk factors and metabolic syndrome. BMC Cardiovasc. Disord. 8, 6 (2008).
Billiard, M. Diagnosis of narcolepsy and idiopathic hypersomnia. An update based on the International classification of sleep disorders, 2nd edition. Sleep Med. Rev. 11, 377–388 (2007).
Weir, B.S., Anderson, A.D. & Hepler, A.B. Genetic relatedness analysis: modern data and new challenges. Nat. Rev. Genet. 7, 771–780 (2006).
Novembre, J. et al. Genes mirror geography within Europe. Nature 456, 98–101 (2008).
Li, Y., Willer, C., Sanna, S. & Abecasis, G. Genotype imputation. Annu. Rev. Genomics Hum. Genet. 10, 387–406 (2009).
Benjamini, Y., Drai, D., Elmer, G., Kafkafi, N. & Golani, I. Controlling the false discovery rate in behavior genetics research. Behav. Brain Res. 125, 279–284 (2001).
Acknowledgements
We thank all the subjects for their generous participation in this study; our colleagues K. Harshman and O. Hagenbuchle and the staff of the Lausanne DNA array facility for their expert technical assistance; and F. Canellas from the University Hospital Son Dureta, Palma-Spain and P. Young from the Department of Neurology, University Hospital Muenster-Germany for providing additional data on narcolepsy patients. Support for this research was provided by the French Ministry of Research and Higher Education, Project Agence Nationale de la Recherche-07-MRAR (France), PHRC 2007-P070138 (France), European Narcolepsy Network, an unrestricted grant from UCB Pharma S.A. (Belgium), the University and the State of Vaud (Switzerland), the Swiss National Science Foundation (grants 3100AO-108478/2, 3100AO-116323/1 and 310000-112552/1), the European Framework Project 6 (EuroDia and the AnEuploidy projects) and GlaxoSmithKline (which sponsored in part the CoLaus study). S.B. is funded by the Giorgi-Cavaglieri Foundation and the Leenaards Foundation. Special thanks to T. Johnson for his valuable advice on data analysis. The computations were performed in part at the Vital-IT center for high-performance computing of the Swiss Institute of Bioinformatics. This study was conducted on behalf of the European Narcolepsy Network.
Author information
Authors and Affiliations
Contributions
Participant recruitment and clinical assessment: Y.D., G.J.L., C.E.H.M.D., A.I., J.S., R.P.A., J.L.V., S.O., I.A., I.T., P.J., S.K., C.B., J.M., M.L., G.M., P.G., A.B., M.B., G.E., F.H.J.C., P.V., G.W., D.M.W., V.M. and R.H. Sample processing and genotyping: H.H., J.L.V., I.T., B.P., C.P., J.V.B., G.D., G.E., W.V. and F.H.J.C. Data analysis: Z.K., H.H., A.V. and M.T. Study design and management: Y.D., G.J.L., J.S.B., S.B. and M.T. Manuscript preparation: H.H., Z.K., Y.D., J.S.B., S.B. and M.T. CoLaus control data: P.V., G.W., D.M.W. and V.M. All authors contributed to the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
Y.D., G.J.L., P.J., C.B., M.L., G.M., S.O., J.M. and M.T. declare having received honoraria as speakers and/or members of the advisory board of UCB Pharma S.A. (Belgium). P.G. has received honoraria as a speaker and member of the XYREM advisory board from UCB GmbH and an honorarium as a speaker from Cephalon GmbH (Germany). R.H. received unrestricted grants from ResMed P.V., and G.W. received an unrestricted grant from GSK for the CoLaus project. V.M. and D.M.W. are full-time employees of GlaxoSmithKline, a pharmaceutical company.
Supplementary information
Supplementary Text and Figures
Supplementary Note, Supplementary Tables 1–3 and Supplementary Figures 1 and 2 (PDF 1078 kb)
Rights and permissions
About this article
Cite this article
Hor, H., Kutalik, Z., Dauvilliers, Y. et al. Genome-wide association study identifies new HLA class II haplotypes strongly protective against narcolepsy. Nat Genet 42, 786–789 (2010). https://doi.org/10.1038/ng.647
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.647
This article is cited by
-
The immunopathogenesis of narcolepsy type 1
Nature Reviews Immunology (2024)
-
Narcolepsy type I-associated DNA methylation and gene expression changes in the human leukocyte antigen region
Scientific Reports (2023)
-
Genetics of circadian rhythms and sleep in human health and disease
Nature Reviews Genetics (2023)
-
Origins and immunopathogenesis of autoimmune central nervous system disorders
Nature Reviews Neurology (2023)
-
Narcolepsy risk loci outline role of T cell autoimmunity and infectious triggers in narcolepsy
Nature Communications (2023)