Abstract
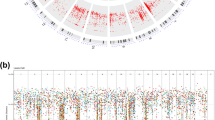
Genome-wide analyses of human lung adenocarcinoma have identified regions of consistent copy-number gain or loss, but in many cases the oncogenes and tumor suppressors presumed to reside in these loci remain to be determined. Here we identify the downstream of tyrosine kinase (Dok) family members Dok1, Dok2 and Dok3 as lung tumor suppressors. Single, double or triple compound loss of these genes in mice results in lung cancer, with penetrance and latency dependent on the number of lost Dok alleles. Cancer development is preceded by an aberrant expansion and signaling profile of alveolar type II cells and bronchioalveolar stem cells. In human lung adenocarcinoma, we identify DOK2 as a target of copy-number loss and mRNA downregulation and find that DOK2 suppresses lung cancer cell proliferation in vitro and in vivo. Given the genomic localization of DOK2, we propose it as an 8p21.3 haploinsufficient human lung tumor suppressor.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Carpino, N. et al. p62(dok): a constitutively tyrosine-phosphorylated, GAP-associated protein in chronic myelogenous leukemia progenitor cells. Cell 88, 197–204 (1997).
Yamanashi, Y. & Baltimore, D. Identification of the Abl- and rasGAP-associated 62 kDa protein as a docking protein, Dok. Cell 88, 205–211 (1997).
Di Cristofano, A. et al. Molecular cloning and characterization of p56dok-2 defines a new family of RasGAP-binding proteins. J. Biol. Chem. 273, 4827–4830 (1998).
Cong, F., Yuan, B. & Goff, S.P. Characterization of a novel member of the DOK family that binds and modulates Abl signaling. Mol. Cell. Biol. 19, 8314–8325 (1999).
Grimm, J. et al. Novel p62dok family members, dok-4 and dok-5, are substrates of the c-Ret receptor tyrosine kinase and mediate neuronal differentiation. J. Cell Biol. 154, 345–354 (2001).
Crowder, R.J., Enomoto, H., Yang, M., Johnson, E.M. Jr. & Milbrandt, J. Dok-6, a Novel p62 Dok family member, promotes Ret-mediated neurite outgrowth. J. Biol. Chem. 279, 42072–42081 (2004).
Okada, K. et al. The muscle protein Dok-7 is essential for neuromuscular synaptogenesis. Science 312, 1802–1805 (2006).
Yamanashi, Y. et al. Role of the rasGAP-associated docking protein p62(dok) in negative regulation of B cell receptor-mediated signaling. Genes Dev. 14, 11–16 (2000).
Di Cristofano, A. et al. p62(dok), a negative regulator of Ras and mitogen-activated protein kinase (MAPK) activity, opposes leukemogenesis by p210(bcr-abl). J. Exp. Med. 194, 275–284 (2001).
Jones, N. & Dumont, D.J. Recruitment of Dok-R to the EGF receptor through its PTB domain is required for attenuation of Erk MAP kinase activation. Curr. Biol. 9, 1057–1060 (1999).
Suzu, S. et al. p56(dok-2) as a cytokine-inducible inhibitor of cell proliferation and signal transduction. EMBO J. 19, 5114–5122 (2000).
Zhao, M., Janas, J.A., Niki, M., Pandolfi, P.P. & Van Aelst, L. Dok-1 independently attenuates Ras/mitogen-activated protein kinase and Src/c-myc pathways to inhibit platelet-derived growth factor-induced mitogenesis. Mol. Cell. Biol. 26, 2479–2489 (2006).
Yasuda, T. et al. Role of Dok-1 and Dok-2 in myeloid homeostasis and suppression of leukemia. J. Exp. Med. 200, 1681–1687 (2004).
Niki, M. et al. Role of Dok-1 and Dok-2 in leukemia suppression. J. Exp. Med. 200, 1689–1695 (2004).
Lau, S.K., Luthringer, D.J. & Eisen, R.N. Thyroid transcription factor-1: a review. Appl. Immunohistochem. Mol. Morphol. 10, 97–102 (2002).
Al-Hajj, M. & Clarke, M.F. Self-renewal and solid tumor stem cells. Oncogene 23, 7274–7282 (2004).
Dick, J.E. & Lapidot, T. Biology of normal and acute myeloid leukemia stem cells. Int. J. Hematol. 82, 389–396 (2005).
Kim, C.F. et al. Identification of bronchioalveolar stem cells in normal lung and lung cancer. Cell 121, 823–835 (2005).
Chitale, D. et al. An integrated genomic analysis of lung cancer reveals loss of DUSP4 in EGFR-mutant tumors. Oncogene 28, 2773–2783 (2009).
Weir, B.A. et al. Characterizing the cancer genome in lung adenocarcinoma. Nature 450, 893–898 (2007).
Wistuba, I.I. et al. Allelic losses at chromosome 8p21–23 are early and frequent events in the pathogenesis of lung cancer. Cancer Res. 59, 1973–1979 (1999).
Ohata, H. et al. Deletion mapping of the short arm of chromosome 8 in non-small cell lung carcinoma. Genes Chromosom. Cancer 7, 85–88 (1993).
Rhodes, D.R. et al. ONCOMINE: a cancer microarray database and integrated data-mining platform. Neoplasia 6, 1–6 (2004).
Su, L.J. et al. Selection of DDX5 as a novel internal control for Q-RT-PCR from microarray data using a block bootstrap re-sampling scheme. BMC Genomics 8, 140 (2007).
Stearman, R.S. et al. Analysis of orthologous gene expression between human pulmonary adenocarcinoma and a carcinogen-induced murine model. Am. J. Pathol. 167, 1763–1775 (2005).
Jackson, E.L. et al. Analysis of lung tumor initiation and progression using conditional expression of oncogenic K-ras. Genes Dev. 15, 3243–3248 (2001).
Pei, X.H., Bai, F., Smith, M.D. & Xiong, Y. p18Ink4c collaborates with Men1 to constrain lung stem cell expansion and suppress non-small-cell lung cancers. Cancer Res. 67, 3162–3170 (2007).
Yanagi, S. et al. Pten controls lung morphogenesis, bronchioalveolar stem cells, and onset of lung adenocarcinomas in mice. J. Clin. Invest. 117, 2929–2940 (2007).
Ventura, J.J. et al. p38alpha MAP kinase is essential in lung stem and progenitor cell proliferation and differentiation. Nat. Genet. 39, 750–758 (2007).
Yasuda, T. et al. Dok-1 and Dok-2 are negative regulators of T cell receptor signaling. Int. Immunol. 19, 487–495 (2007).
Shinohara, H. et al. Dok-1 and Dok-2 are negative regulators of lipopolysaccharide-induced signaling. J. Exp. Med. 201, 333–339 (2005).
Ng, C.H., Xu, S. & Lam, K.P. Dok-3 plays a nonredundant role in negative regulation of B-cell activation. Blood 110, 259–266 (2007).
Inoue, A., Yasuda, T., Yamamoto, T. & Yamanashi, Y. Dok-1 is a positive regulator of IL-4 signalling and IgE response. J. Biochem. 142, 257–263 (2007).
Kashiwada, M. et al. Downstream of tyrosine kinases-1 and Src homology 2-containing inositol 5′-phosphatase are required for regulation of CD4+CD25+ T cell development. J. Immunol. 176, 3958–3965 (2006).
Tonon, G. et al. High-resolution genomic profiles of human lung cancer. Proc. Natl. Acad. Sci. USA 102, 9625–9630 (2005).
Fujiwara, Y. et al. Evidence for the presence of two tumor suppressor genes on chromosome 8p for colorectal carcinoma. Cancer Res. 53, 1172–1174 (1993).
Emi, M. et al. Frequent loss of heterozygosity for loci on chromosome 8p in hepatocellular carcinoma, colorectal cancer, and lung cancer. Cancer Res. 52, 5368–5372 (1992).
Macoska, J.A. et al. Evidence for three tumor suppressor gene loci on chromosome 8p in human prostate cancer. Cancer Res. 55, 5390–5395 (1995).
Scholnick, S.B. et al. Chromosome 8 allelic loss and the outcome of patients with squamous cell carcinoma of the supraglottic larynx. J. Natl. Cancer Inst. 88, 1676–1682 (1996).
Dobbin, K.K. et al. Interlaboratory comparability study of cancer gene expression analysis using oligonucleotide microarrays. Clin. Cancer Res. 11, 565–572 (2005).
Ma, L., Chen, Z., Erdjument-Bromage, H., Tempst, P. & Pandolfi, P.P. Phosphorylation and functional inactivation of TSC2 by Erk implications for tuberous sclerosis and cancer pathogenesis. Cell 121, 179–193 (2005).
Acknowledgements
We thank K. Politi, H.E. Varmus, L.F. Cai, C.F. Kim, T. Motoi, R. Hobbs, J. Clohessy, T. Yung, A. Carracedo, K. Ito, Pandolfi lab members, members of the Memorial Sloan-Kettering Cancer Center (MSKCC) Lung Cancer Oncogenome Group and members of the Dana-Farber/Harvard Cancer Center Lung Cancer Research Program for advice and discussion; B. Clarkson (MSKCC) for reagents and discussion; M. Asher, T. Matos and A. Egia for histology services and immunohistochemistry; and the MSKCC, University of Iowa, and Dana-Farber Cancer Institute flow cytometry core facilities for technical assistance. This work was funded by National Cancer Institute grants and CA-129243 (to M.L.) CA-64593 (to P.P.P.) and by the Steps for Breath Fund from the Society of MSKCC and the Thomas G. Labrecque Foundation (to M.N.).
Author information
Authors and Affiliations
Contributions
A.H.B., M.N., A.M. and P.P.P. designed and analyzed the experiments. B.S.T., C.B., W.L.G. and M.L. conducted the human genetic studies. A.V. and N.D.S. analyzed the SNP array data. J.S., N.M., J.T.F., W.L.G. and M.L. coordinated the human pathological sample acquisition and distribution. J.T.-F. reviewed all mouse pathology. Some of the experiments were conducted in the laboratory of P.B.R. A.H.B., M.N. and P.P.P. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7 and Supplementary Tables 1 and 2 (PDF 11898 kb)
Rights and permissions
About this article
Cite this article
Berger, A., Niki, M., Morotti, A. et al. Identification of DOK genes as lung tumor suppressors. Nat Genet 42, 216–223 (2010). https://doi.org/10.1038/ng.527
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.527
This article is cited by
-
LncRNA-NEF suppressed oxaliplatin resistance and epithelial-mesenchymal transition in colorectal cancer through epigenetically inactivating MEK/ERK signaling
Cancer Gene Therapy (2023)
-
Co-expression network analysis identifies a gene signature as a predictive biomarker for energy metabolism in osteosarcoma
Cancer Cell International (2020)
-
Identification of candidate genes and miRNAs for sensitizing resistant colorectal cancer cells to oxaliplatin and irinotecan
Cancer Chemotherapy and Pharmacology (2020)
-
Analysis of the DOK1 gene in breast cancer
Molecular Biology Reports (2020)
-
Dok-3 deficient mice display different immune clustering and Tim-3 expression
European Journal of Medical Research (2019)